6.2: Heteronuclear 3D NMR- Resonance Assignment in Proteins
- Page ID
- 398288
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)In the previous Chapter we described 2D NMR spectroscopy, which offers significantly greater spectral resolution than basic 1D spectra. In this Chapter we will show how the well-resolved 2D 15N-HSQC resonances can be assigned to specific residues and chemical groups within protein samples. As an example, we will consider a couple of complementary types of 3D NMR data: HNCACB and CBCA(CO)NH and their joint application for making heteronuclear NMR resonance assignment in proteins. Such an assignment opens a number of ways to probe structure and function (e.g. ligand binding) for the target protein samples.
- Grasp why the resonance assignment of 2D 15N-HSQC can be beneficial : the case of ligand (drug) binding by a protein (therapeutic target)
- Familiarize with 3D heteronuclear through-bond (J-coupling) NMR : introduction and case of HNCACB and CBCA(CO)NH pair of 3D experiments
- Follow an example of assignment of heteronuclear NMR resonances (1HN, 15NH, 13Cα, 13Cβ) from a combination of 2D 15N-HSQC and 3D HNCACB/CBCA(CO)NH
15N-HSQC as an assay for probing protein – ligand interactions: the need for the NMR resonance assignment
During the process of rational drug design, it is often necessary to characterize the interactions between the therapeutic target (protein) and candidate drug (ligand) beyond determination of the binding affinity (Kd). Heteronuclear solution NMR experiments 15N-HSQC can provide significant insight for such interactions. Let’s recall that most of the signals in this 2D NMR spectra originate from backbone H-N amide groups and some (minority) from the side chain NH and NH2 groups. The position of 15N-HSQC resonances are defined by the 1HN and 15NH chemical shift values, which in tern depend on the local electronic environment. Ligand binding changes such an environment for the residues forming the binding site even if the tertiary structure of the rest of the protein does not get perturbed. In such a case, the 15N-HSQC resonance pattern undergoes local changes: only the resonances representing NH groups involved in the binding site change their position significantly (>0.05 ppm in 1H and/or >0.2 ppm in 15N dimension) or signal intensity (including peak disappearance). Figure VI.2.A illustrates such a change.
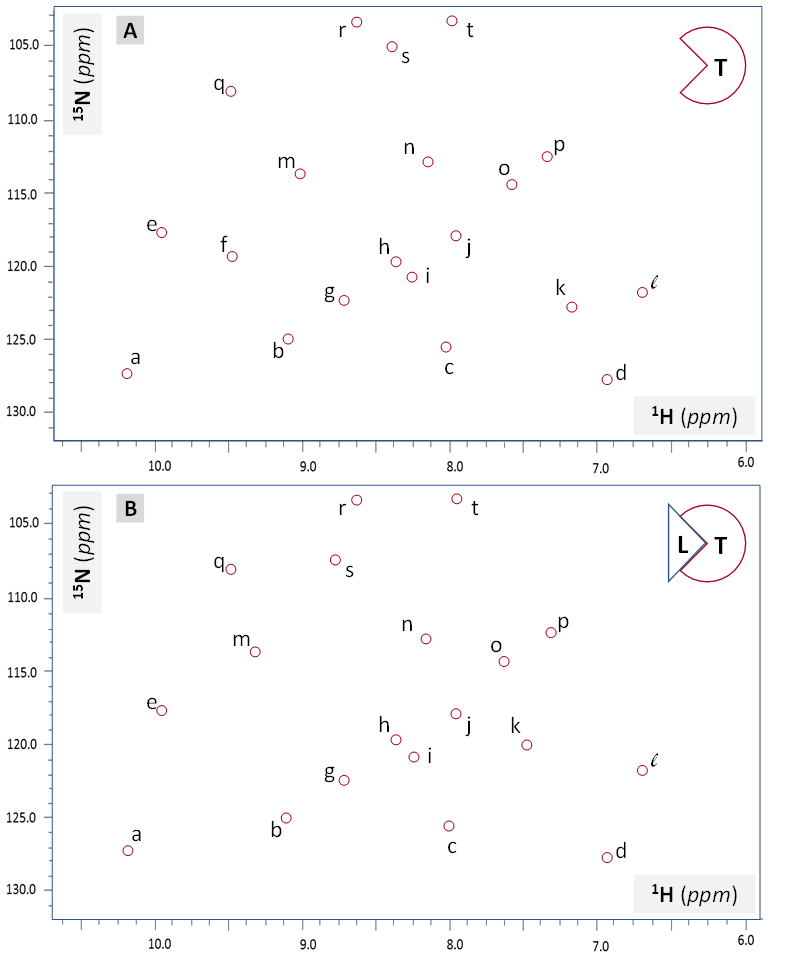
Importantly, every 15N-HSQC resonance in Figure VI.2.A is labeled with a single letter to help identify specific peaks which undergo spectral changes upon ligand binding. This data could have much greater impact if the peaks which underwent the most pronounced changes in position and/or intensity were assigned to specific amino acid residues within the polypeptide and chemical groups within those residues (backbone vs. side chain). The rest of this Chapter demonstrates some of the fundamentals of the heteronuclear NMR resonance assignment methodology.
Heteronuclear 3D NMR introduction: CBCA(CO)NH spectrum as an example
Just like every 2D 15N-HSQC resonance reports a J-coupling via a covalent bond between an 15N and 1H spin-½ nuclei, there are 3D NMR experiments which report resonances originating from J-coupling (through-bond) of three types of spin-½ nuclei (1H, 13C, 15N). In this section we will introduce two such types of 3D NMR data: HNCAB and CBCA(CO)NH. In order to produce a protein sample with nearly complete uniform labeling with 13C and 15N isotopes, bacterial recombinant protein expression can be performed in a minimal media supplemented with 13C-labeled glucose and 15N-labeled ammonium chloride as the sole sources of carbon and nitrogen respectively. Figure VI.2.B introduces a general concept of a 3D NMR data and shows an element of 3D CBCA(CO)NH spectrum.
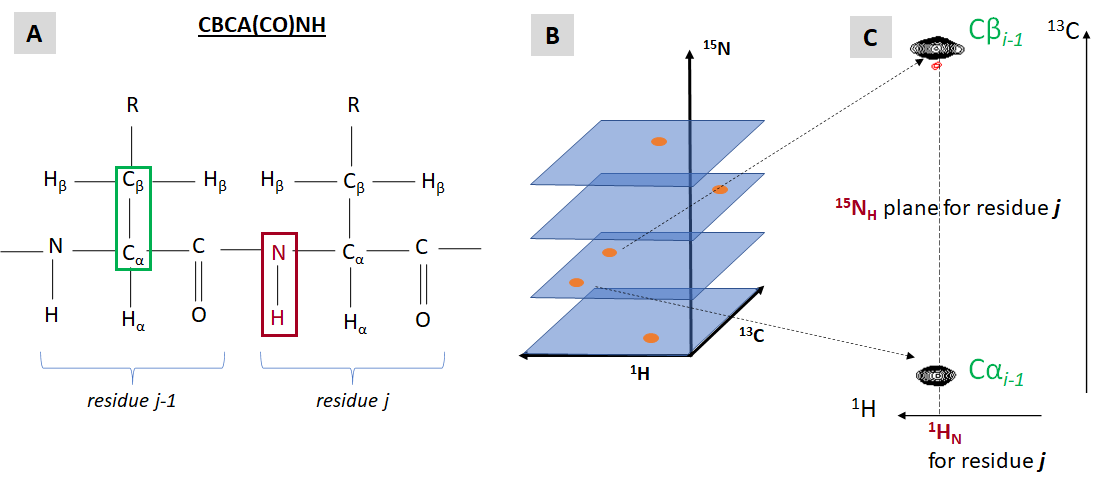
Each resonance (“cross-peak”) of a 3D CBCA(CO)NH spectrum indicates a through-bond (J-coupling scalar) interaction between two atoms of the backbone amide group (1HN and 15NH) or residue j and Cα and Cβ nuclei (13C) of preceding residue j-1. The name of the experiment, CBCA(CO)NH refers to the specific spin-½ nuclei involved (and not involved) in relevant J-coupling interactions: Cβ and Cα are J-coupled to NH while the connecting carbonyl carbon is not reporting any NMR signal (although its magnetization state is affected during the experiment). Two types of residues generate special CBCA(CO)NH peak pattern: prolines have no amide proton, so they do not have CBCA(CO)NH peaks linked with their amide groups. Glycine residues have no Cβ, therefore for any residue following a glycine only a single CBCA(CO)NH resonance will be observed (from glycine NH to previous Cα).
The NMR resonance assignment: combined use of two complementary datasets HNCACB and CBCA(CO)NH
By itself, CBCA(CO)NH does not convey much of sequential information. Another heteronuclear 3D NMR dataset, HNCACB, affords a powerful complement here. Just like CBCA(CO)NH, HNCACB reports resonances originating from J-coupling between backbone amide group and Cα / Cβ nuclei. The difference is that HNCACN reports two additional peaks, all intra-residual: between HN and Cα a Cβ spins (Figure VI.2.C).
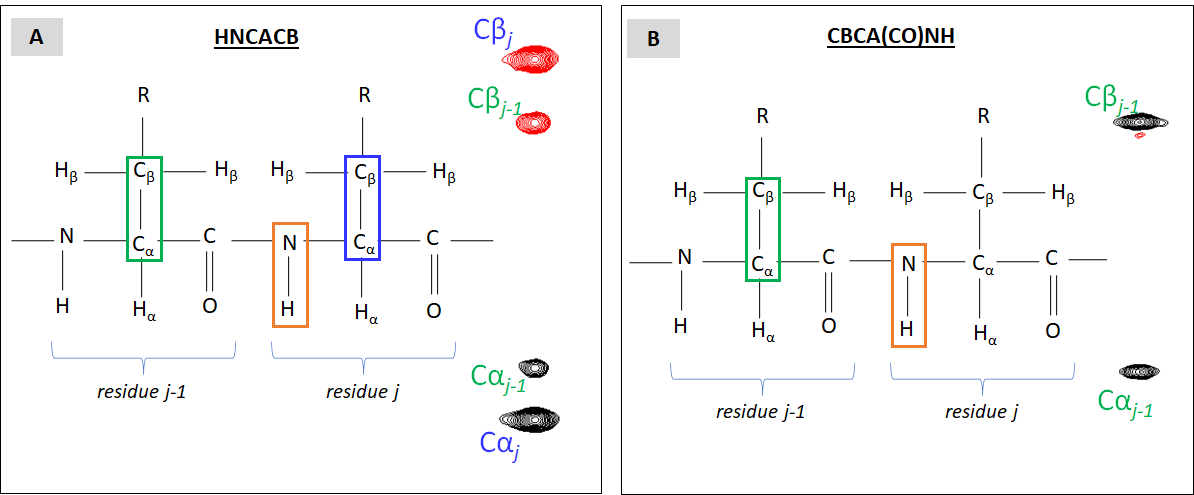
Typically, HNCACB and CBCA(CO)NH are acquired with identical parameters including spectral width in all three dimensions and the same number of data points in the 15N dimension (or 15N planes as on panel B of Figure VI.2.B) Now, let’s imagine that we go through every 15N plane and build the pairs of “residue j / residue j-1″ HNCAB/CBCA(CO)NH peaks. This does not give us the sequence-specific NMR resonance assignments yet but already creates such pairs of 3D cross-peaks linked to di-peptides within the sequence. Now, let’s take into account that for some types of residues their 13Cα and 13Cβ chemical shift values differs remarkably from those from other residue types. For details, take a look at BMRB chemical shift statistics for amino acid residues with emphasis on Gly, Ala, Ser, Thr. Knowing where such residues are positioned within the polypeptide sequence, we can start “connecting the dots” by mapping HNCACB/CBCA(CO)NH planes and di-peptides on actual amino acid sequence.
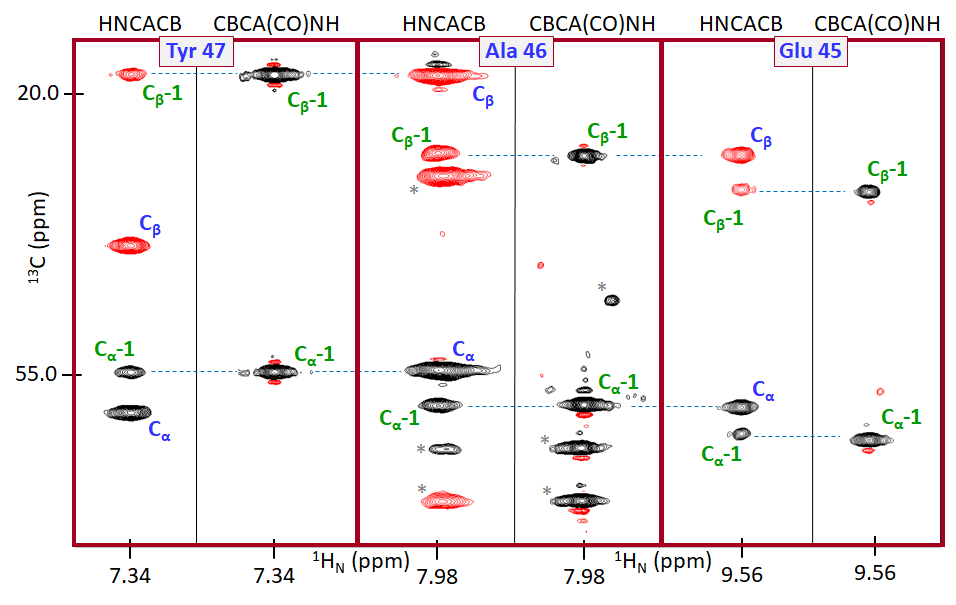
Figure VI.2.D provides a general idea of how the two 3D NMR experiments HNCACB and CBCA(CO)NH can be utilized together to map the signals on the amino acid sequence of a protein sample. The C of Ala residues typically has chemical shift values below 20.0 ppm, which is unique. This allows identification of Ala patterns HNCACB/CBCA(CO)NH spectral patters. Starting from this starting points (as well from other distinct values, e.g. Cα for Gly and Cβ for Ser/Thr), one can continue “connecting the dots” process outlined in Figure VI.2.D to cover the entire sequence. If these two 3D NMR datasets encounter resonance overlaps, which are impossible to resolve, more 3D NMR dataset pairs are utilized in a similar way, e.g. HNCO/HN(CA)CO and others. This process allows assignment to specific residues and chemical groups of nearly all backbone and some side-chain resonances (1HN, 15NH, 13Cα, 13Cβ). Methods for assigning side-chain chemical shift values are not discussed in this chapter but conceptually they are similar to the ones described here.
With the general process of the protein NMR resonance assignment described, let’s assume that this method was successfully applied to the protein target (T) sample presented in Figure VI.2.A. The resonance assignment completion allows one to replace letter labels with residue-number labels (similar to the ones used in Figure VI.2.D). This in turn allows one to determine the specific residues affected directly or allosterically by binding of the ligand (L) to the target. In many cases, such information together with other data leads to the determination of the ligand binding residues within the target. If the ligand is a candidate therapeutic agent, identification of the ligand binding residues greatly advances ensuing efforts to optimize the drug.
Analyze Figure VI.2.A and list at least two resonances which undergo major spectral changes upon binding of the unlabeled ligand (L) to the 15N-labeled target protein (T). Major spectral changes for this model spectrum include resonances moving by >0.05 ppm in 1H or >0.2 ppm in 15N dimensions as well as peak disappearance (peak intensity going down to zero).
Solution
Upon ligand L binding target protein (T), resonance f disappears and resonance s moves by >0.05 ppm in 1H dimension.
Inspect BMRB entry 50205 and list all the heteronuclear NMR datasets utilized for the NMR resonance assignment.
Solution
BMRB entry 50205 contains the chemical shift assignment data for the target sample and offers several ways to look at its underlying NMR data including the list of experiments used to perform the NMR resonance assignment and the chemical shift values. E.g., the NMR-STAR v3 text file has a section titled _Experiment_list, which sums up the heteronuclear NMR data types used for making the assignments: 2D 1H-15N HSQC and 3D HNCACB, CBCA(CO)NH, HNCO and HN(CA)CO.
How many 3D HNCACB resonances would you expect to originate from a Lys residue which is preceded by a Met?
Solution
four as both Lys and Met have backbone amide (HN) groups and both have Cα and Cβ atoms.
Practice Problems
Problem 1. Analyze Figure VI.2.A and list all the resonances which undergo major spectral changes upon binding of the unlabeled ligand (L) to the 15N-labeled target protein (T). Example 1 above will help you start the analysis.
Problem 2. From BMRB entry linked to PDB 5VNT, list all the heteronuclear NMR datasets utilized for the NMR resonance assignment for the target sample.
Problem 3. Let’s consider panel B of Figure VI.2.B. Imagine that the 13C dimension is taken out of the spectrum (all 13C planes are collapsed together). What type of 2D spectrum will remain after such a dimension reduction?
Problem 4. How many 3D HNCACB resonances would you expect to originate from a Gly residue which is preceded by a Pro?
Problem 5. How many 3D HNCACB resonances would you expect to originate from a Pro residue which is preceded by a Gly?
Problem 6*. Look up the amino acid NMR chemical shift values statistics table presented with BMRB repository and list the average values for the following resonances: 15N, 13Cα and 13Cβ for Gly, Ala, Tyr, Glu, Arg, Ser, Thr, Pro. From this analysis, suggest what types of residues tend to report unusually low or high chemical shift values in comparison with the rest of the amino acids?


