25.11: Cell-Surface Carbohydrates and Influenza Viruses
- Page ID
- 36463
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)After completing this section, you should be able to:
- Explain how carbohydrates are related to blood type.
- Explain how influenza works and how it is influenced by carbohydrates.
Oligosaccharides
An oligosaccharide is a saccharide polymer containing a small number (typically two to ten) of monosaccharides. Oligosaccharides can have many functions; for example, they are commonly found on the plasma membrane of animal cells where they can play a role in cell-cell recognition. In general, they are found attached to compatible amino acid side-chains in proteins or to lipids. Oligosaccharides are often found as a component of glycoproteins or glycolipids. They can be used as chemical markers on the outside of cells, often for cell recognition. Oligosaccharides are also responsible for determining blood type.
Glycoproteins
Carbohydrates are covalently attached to many different biomolecules, including lipids, to form glycolipids, and proteins, to form glycoproteins. Glycoproteins and glycolipids are often found in biological membranes, to which they are anchored by through nonpolar interactions. What is the function of these carbohydrates? Two are apparent. First, glycosylation of proteins helps protect the protein from degradation by enzyme catalysts within the body. However, their main functions arises from the fact that covalently attached carbohydrates that "decorate" the surface of glycoproteins or glycolipids provide new binding site interactions that allow interactions with other biomolecules. Hence glycosylation allows for cell:cell, cell:protein, or protein:protein interactions. Unfortunately, bacteria and viruses often recognize glycosylated molecules on cell membranes as well, allowing for their import into the cell.
Blood Type
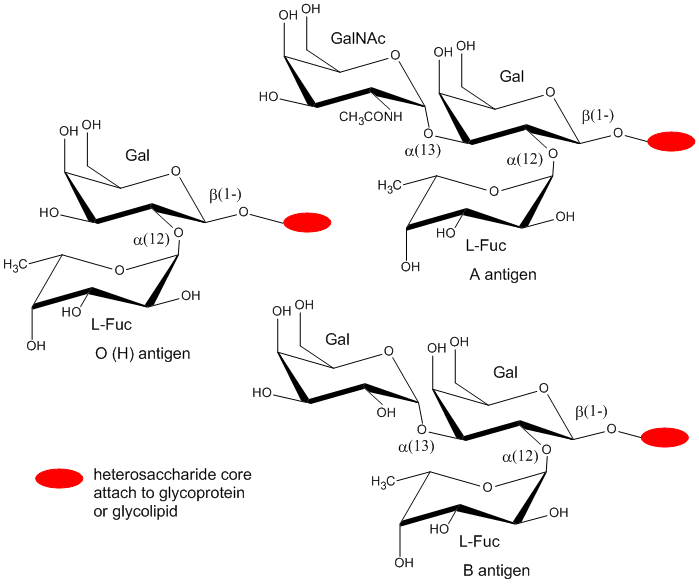
Cell markers are carbohydrate chains on the surface of cells where they act as “road signs” allowing molecules to distinguish one cell from another. Blood markers are exclusively made from four monosaccharides: D-galactose, L-fucose, N-acetylgalactosamine, and N-acetylglucosamine.

ABO blood markers found on red blood cells are made up of oligosaccharides that contain either three or four sugar units as shown below. Oligosaccharides attached to red blood cells determine the ABO blood type.
Of the four blood types, type O has the fewest types of saccharides attached to it while type AB has the most. As a result, type O blood is considered the universal donor because it doesn't have any saccharides present that will appear as foreign when transfused into blood of another type. The reverse is not true. For example:
- If type A blood is given to a patient with type O blood, it will be rejected by the body because there is an unknown species being introduced to the body. Type A blood cells contain N-acetylgalactosamine which is not present in type O blood.
- Since type AB blood has all possible saccharides, type AB blood is considered the universal acceptor.
The Rhesus factor (Rh) in blood also affects donor and acceptor properties but it does not depend on carbohydrates. The Rh factor is determined by the presence (Rh+) or absence (Rh-) of a specific protein on the surface of red blood cells.
Three pandemics of influenza have swept the world since the "Spanish" flu of 1918.
- The "Asian" flu pandemic of 1957;
- the "Hong Kong" flu pandemic of 1968;
- the "Swine" flu pandemic that began in April of 2009.
The influenza virus is a simple yet deadly virus. It interacts with human cells through a surface protein, hemagglutinin (HA). The virus binds to host cells through interaction of HA with cell surface carbohydrates. Once bound the virus internalizes, ultimately leading to release of the RNA genome of the virus into the host cell. The HA protein is the most abundant protein on the viral surface (as surmised by antibody formation).
The influenza virus is typically classified by two kinds of glycoproteins on the surface of the virus: in addition to HA is the enzyme neuraminidase. Two viral strains which have been often discussed are H5N1 and H1N1 which stands for hemagglutinin (H: type 5 or type 1) and neuraminidase (N: type 1). 15 avian and mammalian variants have been identified (based on antibody studies). Only 3 adapted to humans in the last 100 yr, giving pandemic strains H1 (1918), H2 (957) and H3 (1968). Three recent avian variants (H5, H7, and H9) have jumped directly to humans recently but have low human to human transmissibility.

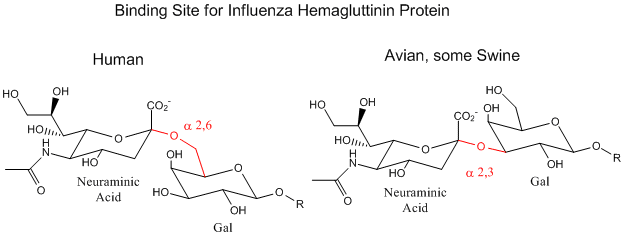
Influenza Virus binds to Cell Surface Glycoproteins with Neu5Ac - A protein on the surface of influenza virus. Hemagluttinin binds to sialic acid (Sia), which is covalently attached to many cell membrane glycoproteins. The sialic acid is usually connected through an alpha (2,3) or alpha (2,6) link to galactose on N-linked glycoproteins. The subtypes found in avian (and equine) influenza isolates bind preferentially to Sia (alpha 2,3) Gal which predominates in avian GI tract where viruses replicate. Human virus of H1, H2, and H3 subtype (cause of the 1918, 1957, and 1968 pandemics) recognize Sia (alpha 2,6) Gal, as the major form in the human respiratory tract. The swine influenza HA bind to Sia (alpha 2,6) Gal and some Sia (alpha 2,3) both of which are commonly found in swine.
The virus, before it leaves the cell, forms a bud on the intracellular side of the cell with the HA and NA in the cell membrane of the host cell. The virus in this state would not leave the cell since its HA molecules would interact with sialic acid residues in the host cell membrane, holding the virus in the membrane. Neuraminidase hydrolyzes sialic acid from cell surface glycoproteins, allowing the virus to complete the budding process and be released from the cell as new viruses. By mimicking the structure of sialic acid, the drugs Oseltamivir (Tamiflu) and zanamivir (Relenza) bind to and inhibit neuraminidase whose activity is necessary for viral release from infected cells. Tamiflu appears to work against N1 of the present H5N1 avian influenza viruses. Governments across the world are stockpiling this drug in case of a pandemic caused by the avian virus jumping directly to humans and becoming transmissible from human to human.

