INTERCHAPTER: Retrosynthetic analysis and metabolic pathway prediction
- Page ID
- 395046
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)
Imagine that you are a biological chemist doing research on bacterial metabolism. You and your colleagues isolate an interesting biomolecule from a bacterial culture, then use mass spectrometry, NMR, and other analytical techniques to determine its structure. Using your 'toolbox' of known organic reaction types - nucleophilic substitution, phosphorylation, aldol additions, and so forth - can you figure out a chemically reasonable pathway by which your compound might be enzymatically synthesized from simple metabolic precursors? In other words, can you fill in the missing biochemical steps (or at least some of them) to come up with a potential new metabolic pathway, which can then be used a hypothesis for future experimental work to try to find and study the actual enzymes involved?
An actual example approximating this scenario is shown below. A complete biosynthetic pathway for isopentenyl diphosphate (IPP), the building block molecule for all isoprenoid compounds, has been known since the 1960's. This pathway, which begins with acetyl-CoA, was shown to be active in yeast, plants, and many other species including humans. However, researchers in the late 1980s uncovered evidence indicating that the known pathway is not present in bacteria, although they clearly use IPP as a building block molecule just as other forms of life do.
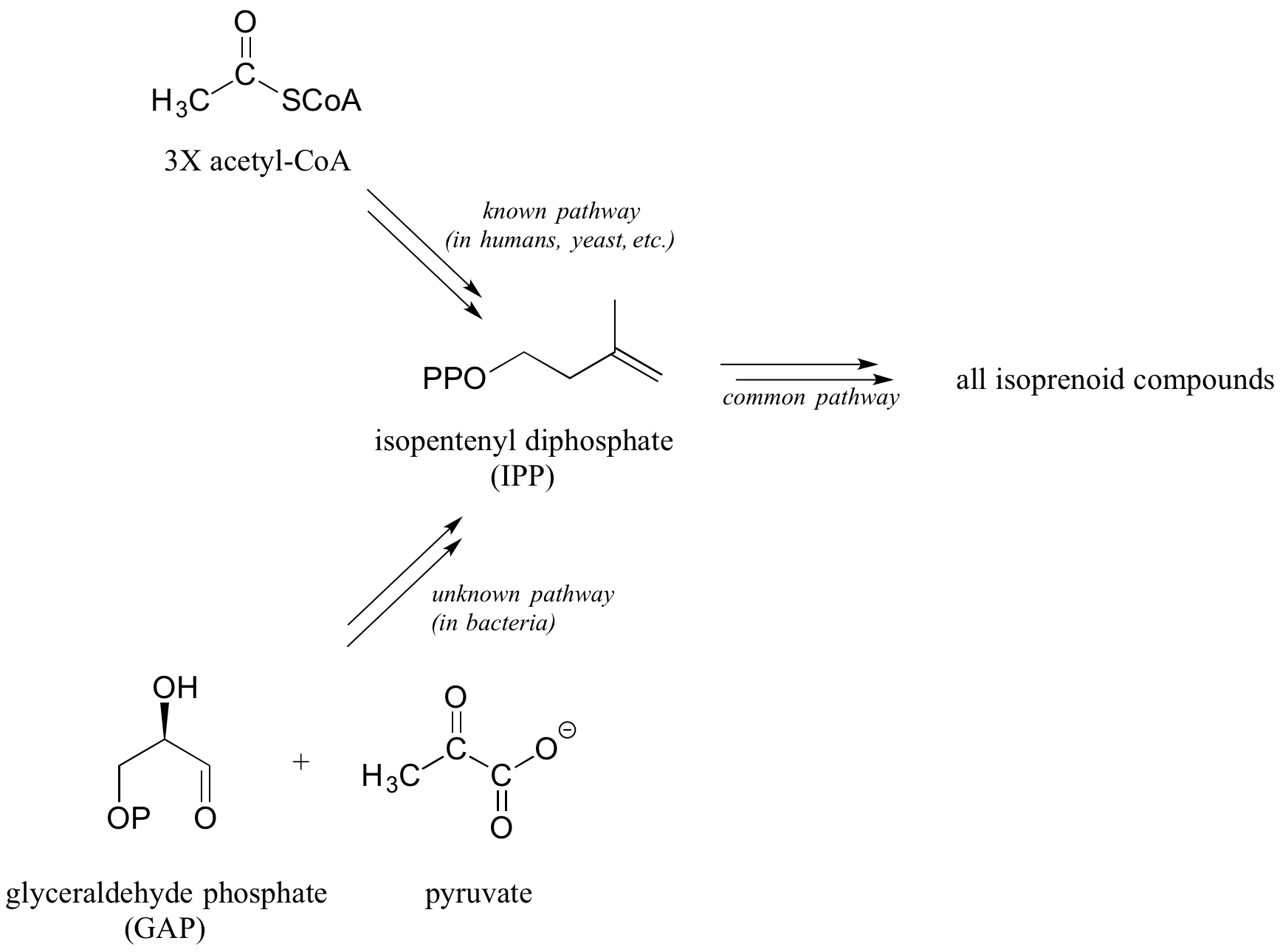
Over the next several years, the researchers conducted a number of experiments in which bacteria were grown on a medium containing glucose 'labeled' with the 13C isotope. With the results from these experiments, combined with their knowledge of common biological organic reaction types, the researchers were able to correctly predict that the bacterial pathway starts with two precursor molecules (pyruvate and glyceraldehyde phosphate instead of acetyl CoA) and they also correctly predicted the first two enzymatic steps of the newly discovered bacterial pathway. This accomplishment eventually led to elucidation of every step in the pathway, and isolation of the enzymes catalyzing them. (Biochem J. 1993, 295, 517; J. Am. Chem. Soc. 1996, 118, 2564; Lipids 2008, 43, 1095)
Why weren't they able to predict the whole pathway? It turns out that several of the later steps were somewhat unusual, unfamiliar reaction types - but discovery of these reactions hinged upon the correct prediction of the more familiar first two steps.
Multi-step transformation problems of this type offer an unparalleled opportunity to use our knowledge of biological organic chemistry combined with creative reasoning to solve challenging, relevant scientific puzzles. At this point in your organic chemistry career, you have not yet accumulated quite enough tools in your reaction toolbox to tackle most real-life biochemical pathway problems such as the one addressed above - but by the time we finish with oxidation and reduction chemistry in chapter 15, you will be able to recognize most of the reaction types that you will encounter in real metabolism, and will be challenged to predict some real pathways in the end-of-chapter problems.
You do, however, have right now enough of a bioorganic repertoire to begin to learn how multi-step pathway problems can be approached, using for practice some generalized, hypothetical examples in which the reaction types involved are limited to those with which you are already familiar.
Imagine that you want to figure out how an old-fashioned mechanical clock is put together. One way to do this is to start with a working clock, and take it apart piece-by-piece. Alternatively, one could start with all of the disassembled pieces, plus a lot of other small parts from different clocks, and try to figure out how to put together the specific clock you are interested in. Which approach is easier? The answer is intuitively obvious - it's usually easier to take things apart than to put them back together.
The same holds true for molecules. If we want to figure out the biosynthetic pathway by which a large, complex biomolecule might be made in a cell, it makes sense to start with the finished product and then mentally work backwards, taking it apart step-by-step using known, familiar reactions, until we get to simpler precursor molecules. Starting with a large collection of potential precursor molecules and trying to put the right ones together to make the target product would be a formidable task.
Retrosynthetic analysis - the concept of mentally dismantling a molecule step by step all the way back to smaller, simpler precursors using known reactions - is a powerful and widely-used intellectual tool first developed by synthetic organic chemists. The approach has also been adapted for use by biological chemists in efforts to predict pathways by which known biomolecules could be synthesized (or degraded) in living things.
In retrosynthesis, we think about a series of reactions in reverse. A backwards (retro) chemical step is symbolized by a 'thick' arrow, commonly referred to as a retrosynthetic arrow, and visually conveys the phrase 'can be formed from'.
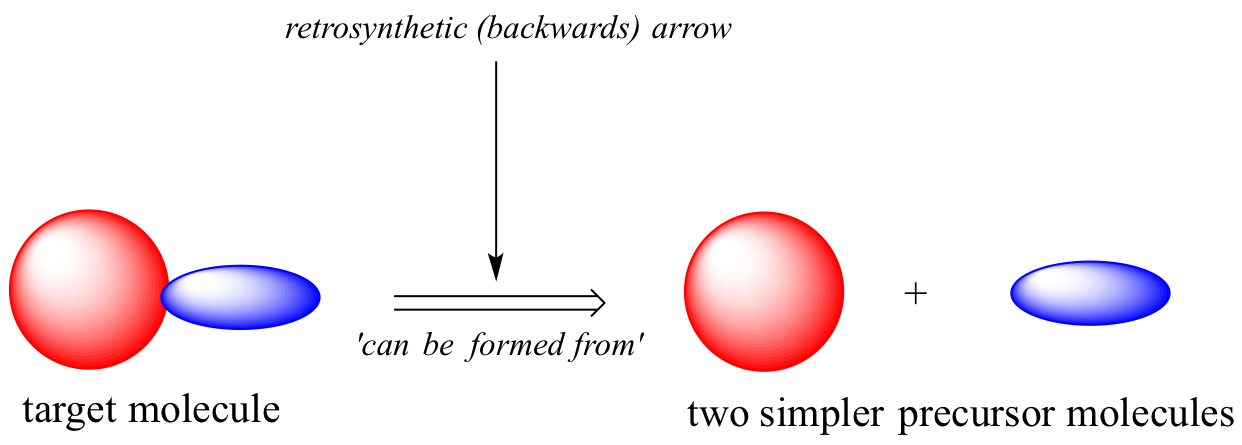

Consider a simple, hypothetical example: starting with the target molecule below, can we come up with a chemically reasonable pathway starting from the precursors indicated?
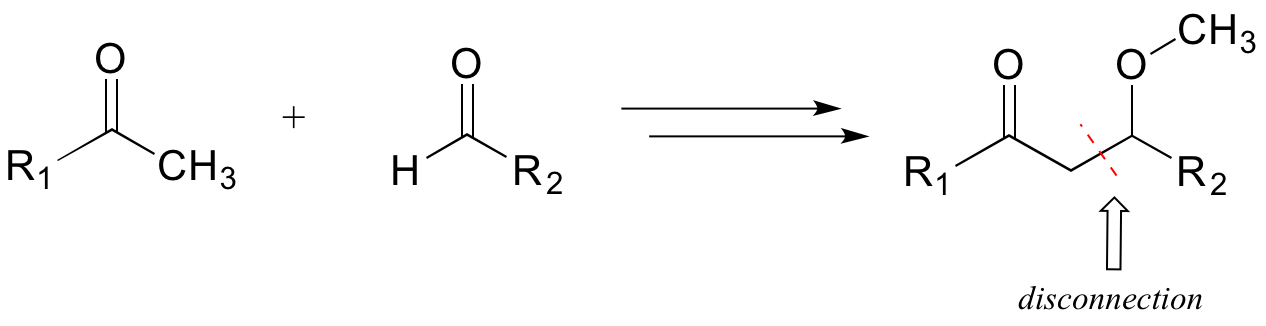
A first step is to identify the relevant disconnection: a key bond (usually a carbon-carbon bond) that must be formed to make the target product from smaller precursors. We search our mental 'toolbox' of common biochemical reaction types, and remember that the only way we know of (so far!) to make a new carbon-carbon bond is through an aldol addition reaction, which takes place at an alpha-carbon. Therefore, we can make a likely disconnection next to the alpha-carbon in the target molecule.
Next, we need to recognize that the aldol addition reaction results in a beta-hydroxy ketone. But our target molecule is beta-methoxy ketone! Working backwards, we realize that the beta-methoxy group could be formed from beta-hydroxy group by a SAM methylation reaction. This is our first retrosynthetic (backwards) step.
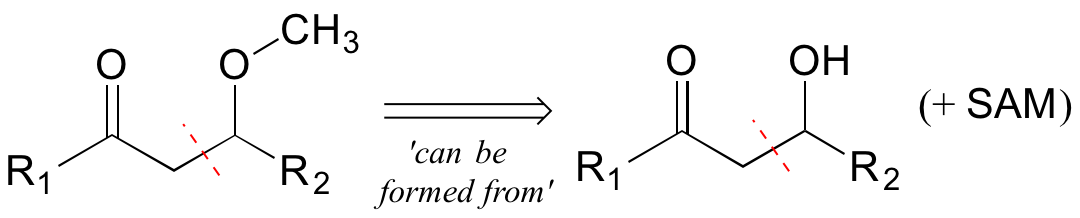
The second retro step (aldol) accounts for the disconnection we recognized earlier, and leads to the two precursor molecules.
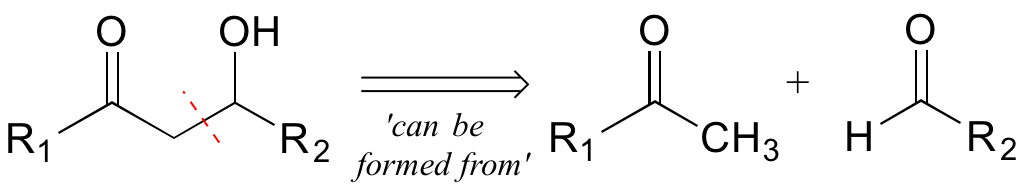
Now, consider the more involved (but still hypothetical) biochemical transformation below:
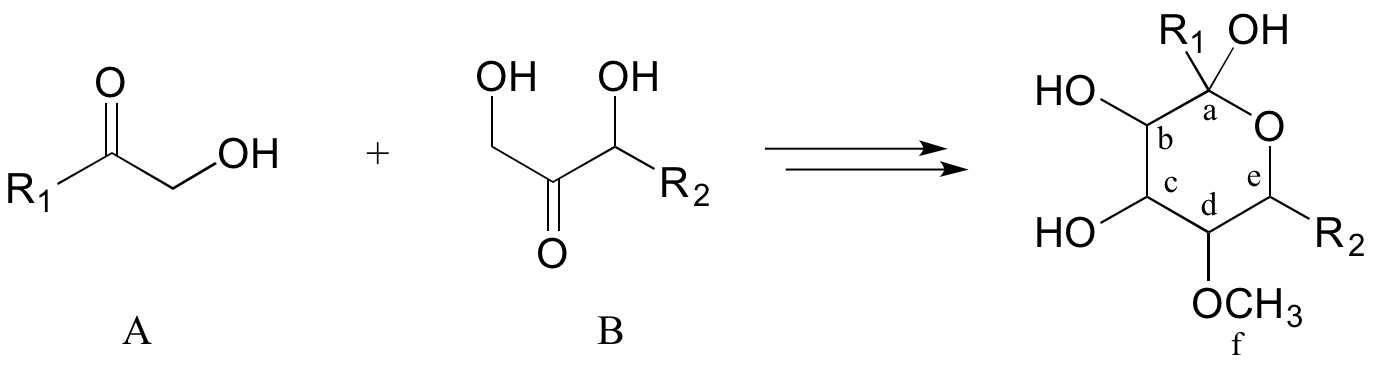
Often the best thing to do first in this type of problem is to count the carbons in the precursor compounds and product - this allows us to recognize when extra carbons on either side must at some point be accounted for in our solution. In this case, one carbon (labeled 'f'') has been gained in the form of a methyl ether in the product. This is easy to account for: we know that the coenzyme S-adenosyl methionine (SAM) often serves as the methyl group donor in enzymatic O- or N-methylation reactions. So, we can propose our first backwards (retro) step: the product as shown could be derived from SAM-dependent methylation of an alcohol group on a proposed intermediate I.
Retrosynthetic step 1:
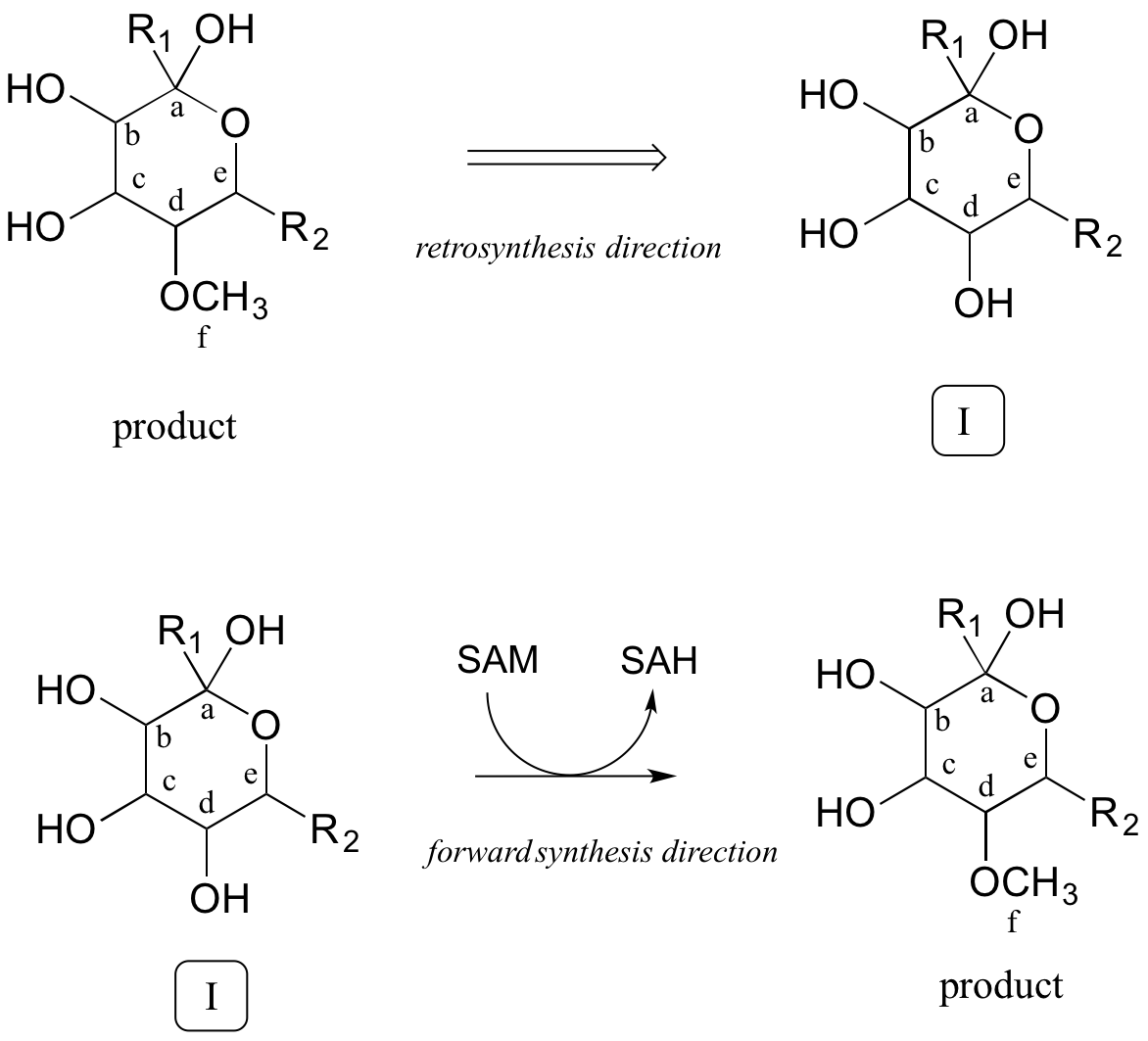
How do we know that the methylation step occurs last? We don't - remember, we are proposing a potential pathway, so the best we can do is propose steps that make chemical sense, and which hopefully can be confirmed or invalidated later through actual experimentation. For now, we'll stick with our initial choice to make the methylation step the last one.
Now that we have accounted for the extra carbon, a key thing to recognize regarding the transformation in question is that two linear molecules are combining to form a cyclic product. Thus, two connections need to be made between reactants A and B, one to join the two, the other to close the circle. Our primary job in the retro direction, then, is to establish in the product the two points of disconnection: in other words, to find the two bonds in the product that need to be taken apart in our retrosynthetic analysis. Look closely at the product: what functional groups do you see? Hopefully, you can identify two alcohol groups, a methyl ether, and (critically) a cyclic hemiketal. We've already accounted for the methyl ether. Identifying the cyclic hemiketal is important because it allows us to make our next 'disconnection': we know how a hemiketal forms from a ketone and an alcohol, so we can mentally work backwards and predict the open-chain intermediate II that could cyclize to form our product.
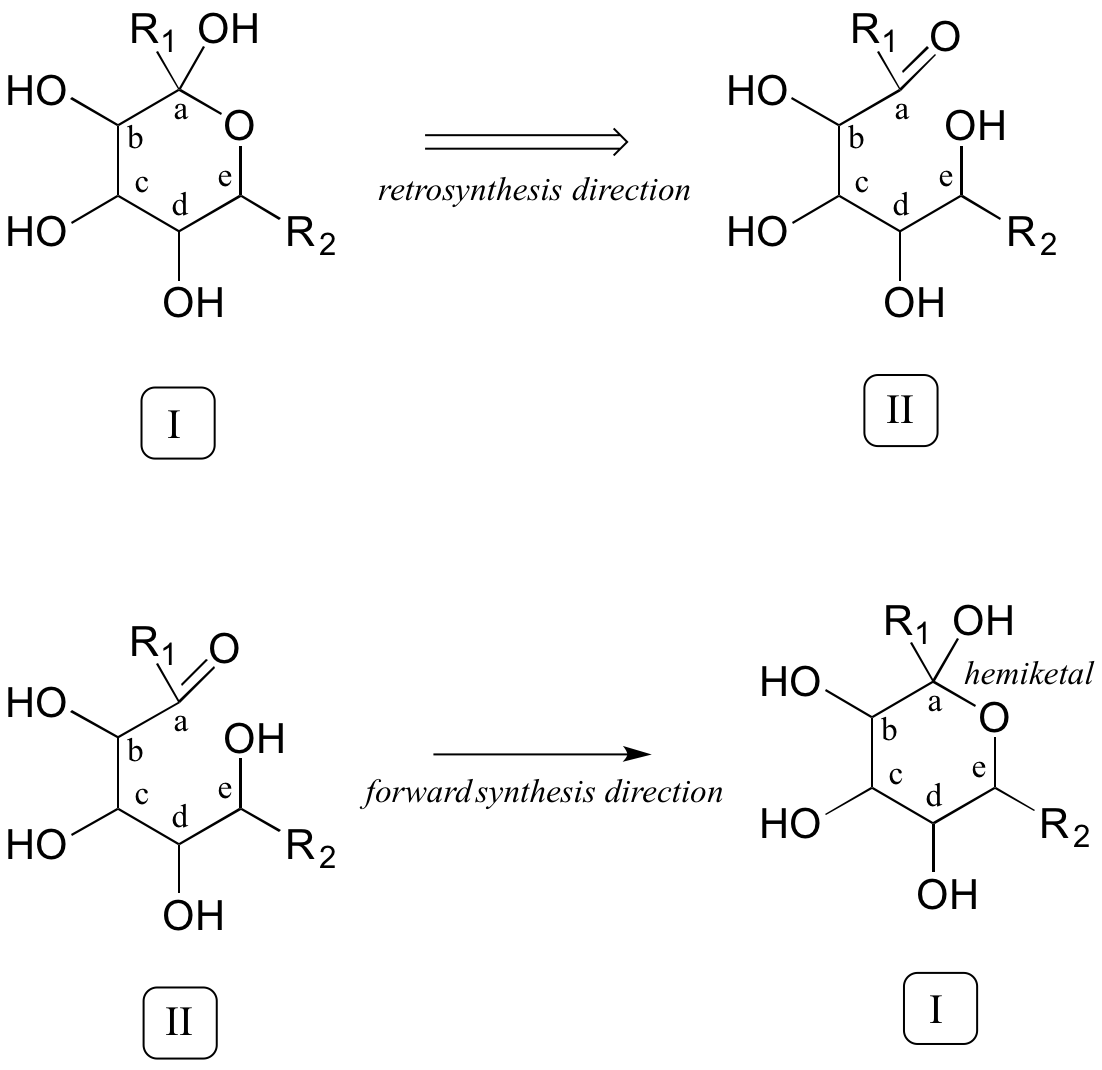
Now, starting with the R1 group and working along the carbon chain, we can account for carbons a-e on the two precursors.
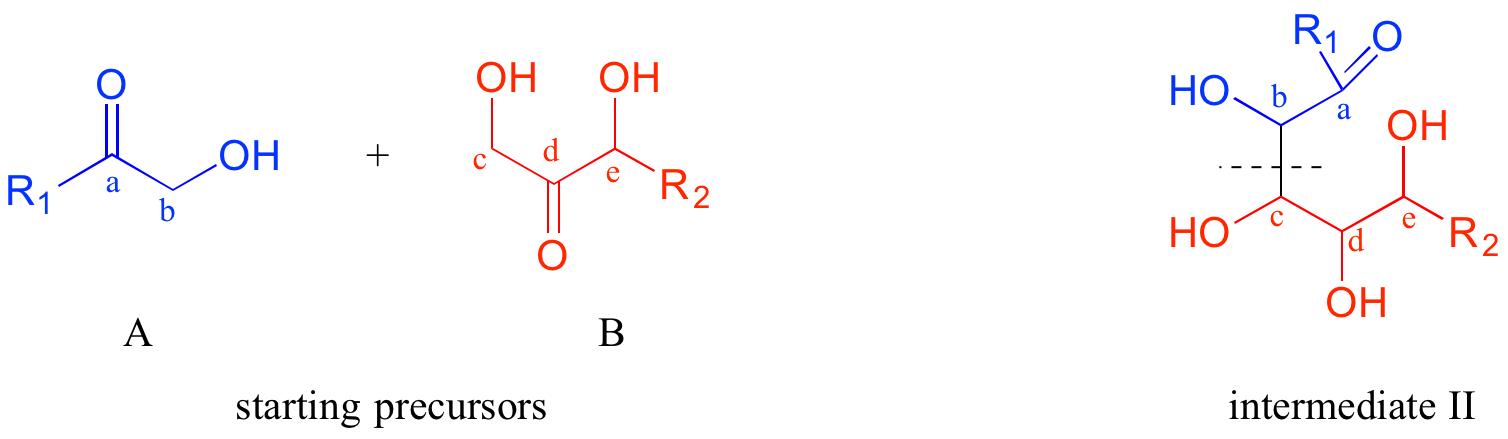
Thus, the next disconnection is between carbons b and c. Here's where our mastery of biological organic reactivity really comes into play: the OH at carbon c of intermediate II is in the beta position relative to carbonyl carbon a. Aldol addition reactions result in beta-hydroxy ketones or aldehydes. Therefore, we can work backward one more step and predict that our intermediate II was formed from an aldol addition reaction between intermediate III (as the nucleophile) and precursor molecule A (as the electrophile).
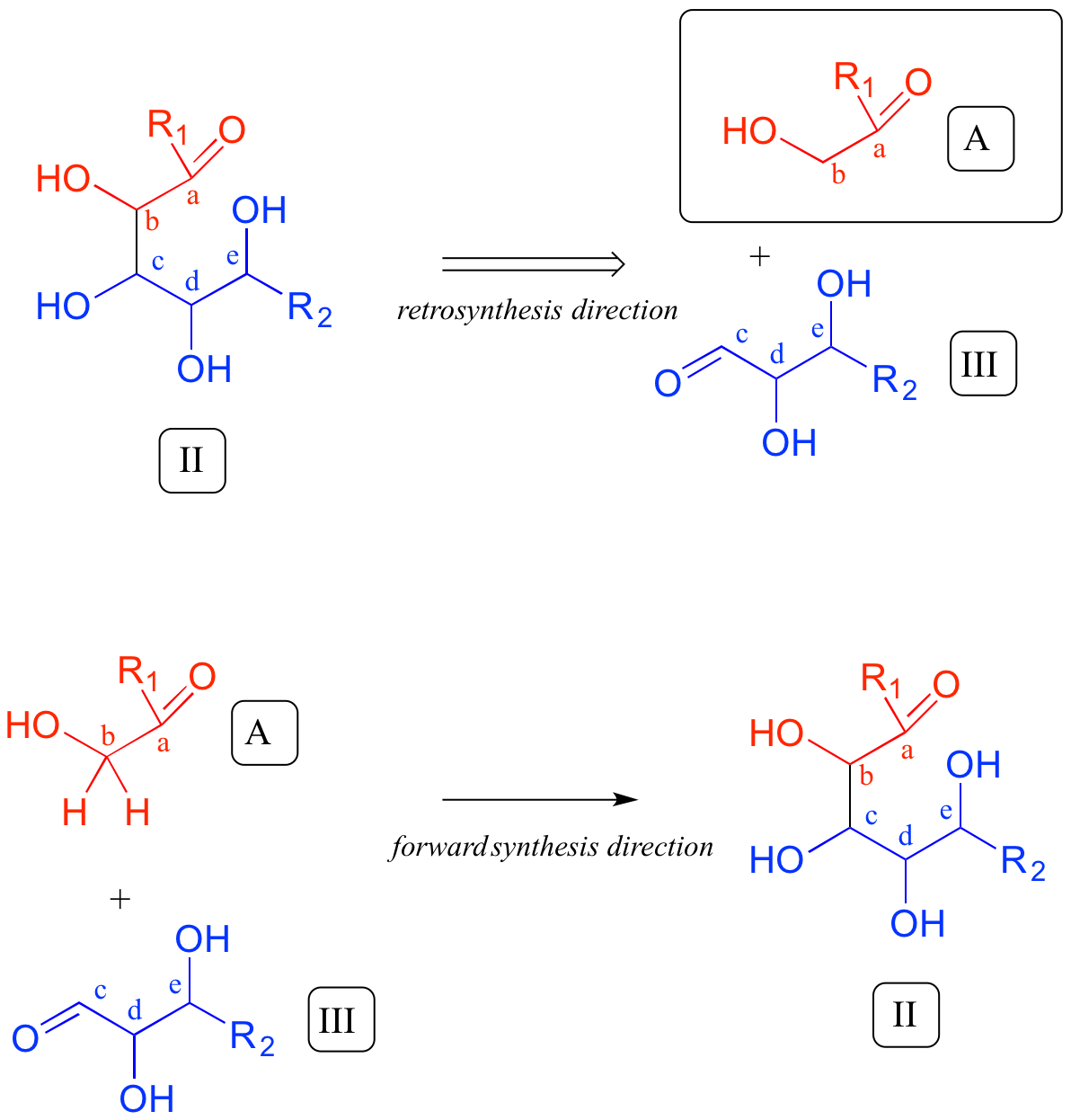
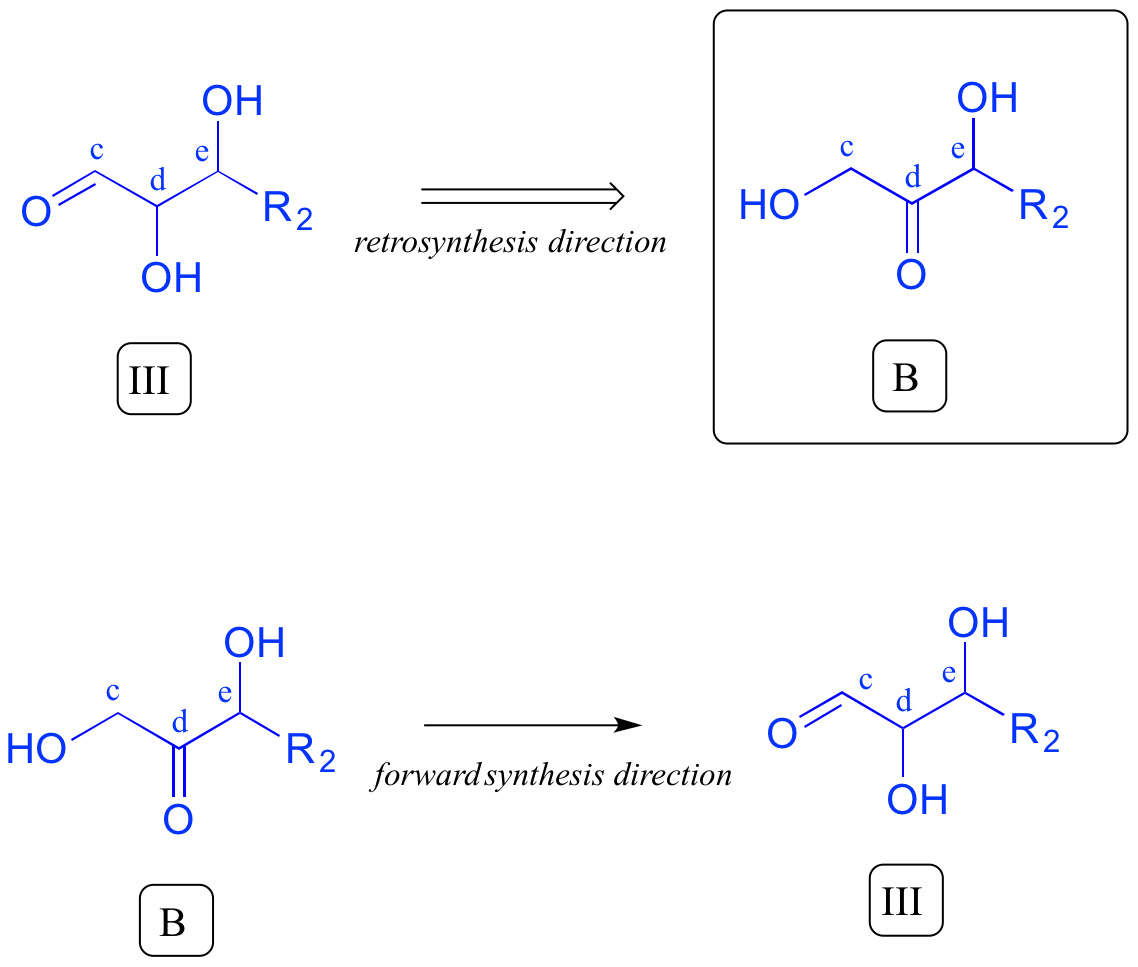
We are most of the way home - we have successfully accounted for given precursor A. Intermediate III, however, is not precursor B. What is different? Both III and B have a carbonyl and two alcohol groups, but the positioning is different: III is an aldehyde, while B is a ketone. Think back to earlier in this chapter: intermediate III could form from isomerization of the carbonyl group in compound B. We have now accounted for our second precursor - we are done!
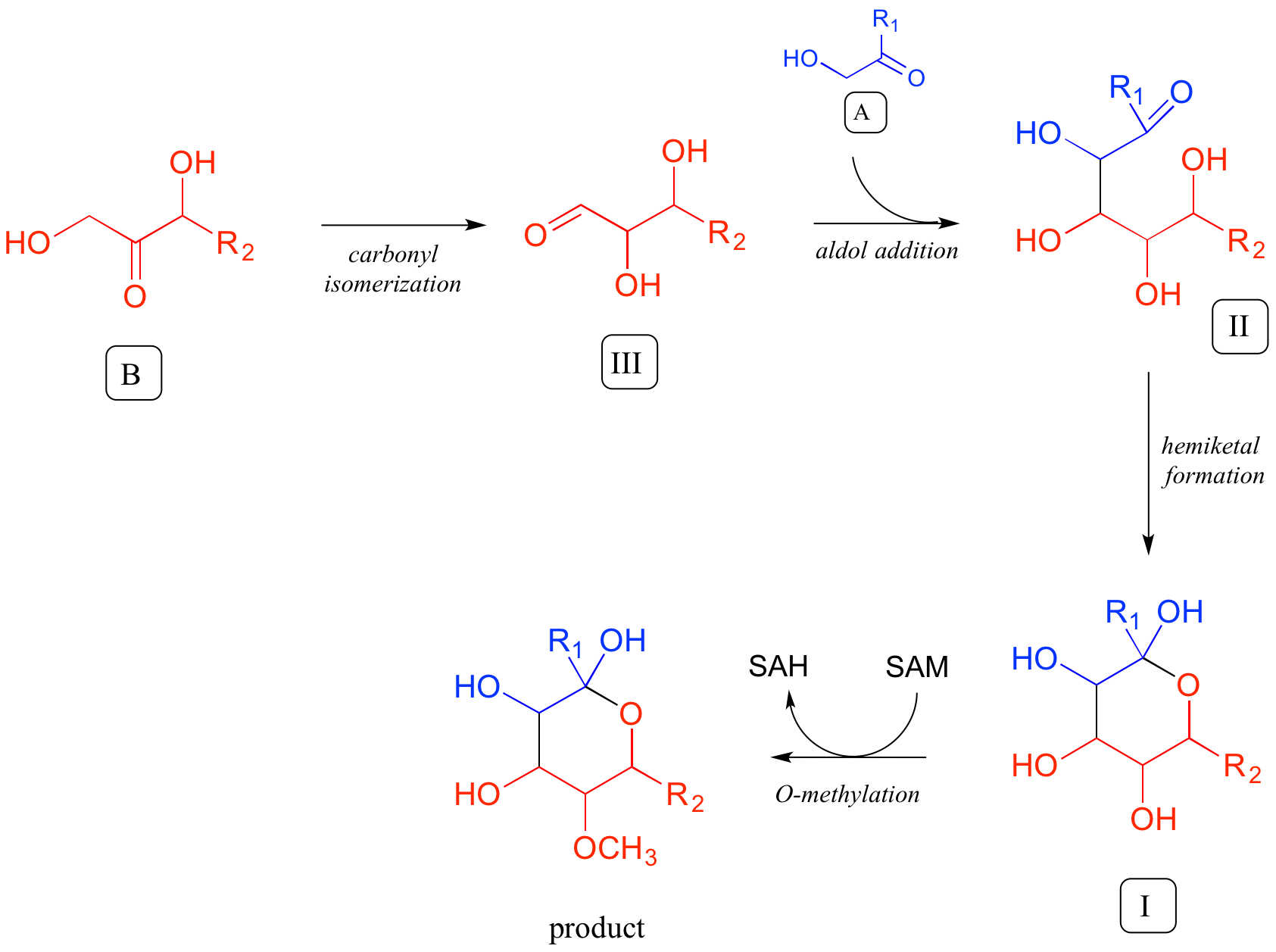
In the forward direction, a complete pathway diagram can be written as follows:
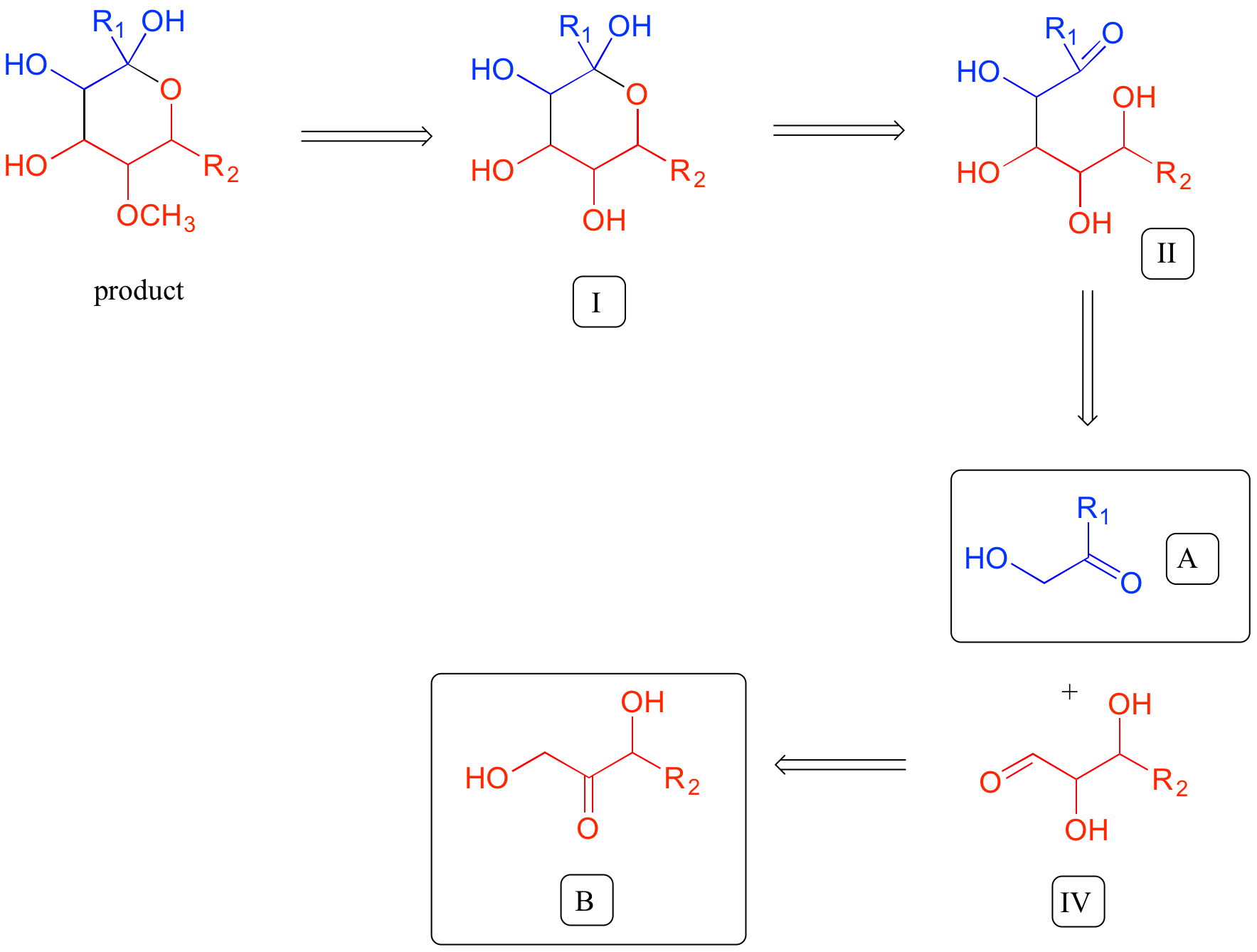
A full 'retrosynthesis' diagram for this problem looks like this:
Practice problems for retrosynthesis/pathway prediction


