3.6: Intermolecular Forces
- Page ID
- 494841
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Describe the types and relative strengths of intermolecular forces possible between atoms or molecules (dispersion forces, dipole-dipole attractions, and hydrogen bonding)
- Identify the types of intermolecular forces experienced by specific molecules based on their structures
- Relate the strength of the intermolecular forces to physical properties, such as boiling point.
So far, we have focused on the forces within molecules, or intramolecular forces, which include covalent and ionic bonds. In this section, we will explore intermolecular forces (IMFs)—the forces of attraction between molecules. While IMFs are also electrostatic in nature, they are much weaker than chemical bonds but play a critical role in determining the physical properties of substances.
Intermolecular forces and the State of Matter
Intermolecular forces strongly influence a substance’s physical state. Consider the three states of matter:
- In solids, particles are closely packed and vibrate in place.
- In liquids, particles are still close together but can move past each other.
- In gases, particles are far apart and move freely.
The phase of a substance depends on the balance between intermolecular forces and kinetic energy (KE). IMFs hold molecules together, while kinetic energy provides the energy required to overcome the attractive forces and thus increase the distance between molecules. The kinetic energy of a system is defined by its temperature and Figure \(\PageIndex{2}\) illustrates how increasing temperature and therefore increasing kinetic energy can cause phase changes.
To illustrate the processes shown in this figure, consider water vapor. When gaseous water is cooled sufficiently, the attractions between H2O molecules can hold the molecules together when they come into contact with each other. This causes the water vapor to condense, forming liquid H2O. For example, liquid water forms on the outside of a cold glass as the water vapor in the air is cooled by the cold glass, as seen in Figure \(\PageIndex{3}\).
We can also liquefy many gases by compressing them, if the temperature is not too high. The increased pressure brings the molecules of a gas closer together, so that the attractions between the molecules are strong relative to their kinetic energy. Consequently, they form liquids. Butane, C4H10, is the fuel used in disposable lighters and is a gas at standard temperature and pressure. Inside the lighter’s fuel compartment, the butane is compressed to a pressure that results in its condensation to the liquid state (Figure \(\PageIndex{4}\)).
Finally, if the temperature of a liquid becomes sufficiently low, or the pressure on the liquid becomes sufficiently high, the molecules of the liquid no longer have enough kinetic energy to overcome the intermolecular forces between them and move past each other, and a solid forms.
There are different types of intermolecular forces between molecules, including dispersion forces, dipole-dipole interactions, and hydrogen bonding. Ion-dipole forces also exist between molecules and ions. The strengths of these attractive forces vary widely, although the intermolecular forces between small molecules are usually weak compared to the intramolecular forces that bond atoms together within a molecule. For example, to overcome the intermolecular forces in one mole of liquid HCl and convert it into gaseous HCl requires only about 17 kilojoules. However, to break the covalent bonds between the hydrogen and chlorine atoms in one mole of HCl requires about 25 times more energy—430 kilojoules (Figure \(\PageIndex{1}\).
Dispersion Forces
All atoms and molecules experience dispersion forces, which arise due to temporary shifts in electron distribution. Because electrons are constantly moving, an atom or molecule can develop a temporary dipole when its electrons become unevenly distributed. This dipole can then induce a dipole in a nearby atom or molecule, leading to a weak attraction between them (Figure \(\PageIndex{5}\)).
Dispersion forces increase with atomic and molecular size. For example, among the halogens, F2 and Cl2 are gases at room temperature (reflecting weaker attractive forces); Br2 is a liquid, and I2 is a solid (reflecting stronger attractive forces). Melting and boiling points increase as atomic size and dispersion forces increase., as seen in Table \(\PageIndex{1}\).
| Halogen | Molar Mass | Atomic Radius | Melting Point | Boiling Point |
|---|---|---|---|---|
| fluorine, F2 | 38 g/mol | 72 pm | 53 K | 85 K |
| chlorine, Cl2 | 71 g/mol | 99 pm | 172 K | 238 K |
| bromine, Br2 | 160 g/mol | 114 pm | 266 K | 332 K |
| iodine, I2 | 254 g/mol | 133 pm | 387 K | 457 K |
| astatine, At2 | 420 g/mol | 150 pm | 575 K | 610 K |
Why do larger atoms Have stronger dispersion forces? Larger atoms have more electrons, and their valence electrons are farther from the nucleus. Because they are less tightly held, their electron clouds are easier to distort, meaning they can form temporary dipoles more easily.
The ease with which a molecule’s electron cloud is distorted is called polarizability.
- Highly polarizable molecules have strong dispersion forces because their electron clouds can shift easily.
- Less polarizable molecules have weaker dispersion forces because their electron clouds are more rigid.
Order the following compounds from lowest to highest boiling point: CH4, SiH4, GeH4, and SnH4. Explain your reasoning.
Solution
C, Si, Ge, and Sn are in the same column of the periodic table and thus have the same number of valence electrons. All four compounds have a tetrahedral geometry and are therefore nonpolar. The only intermolecular forces between the molecules are dispersion forces. Smaller molecules are less polarizable and have weaker dispersion forces. Weaker attractions between molecules require less kinetic energy to overcome the intermolecular forces and therefore the boiling point is lower. The molar masses of CH4, SiH4, GeH4, and SnH4 are approximately 16 g/mol, 32 g/mol, 77 g/mol, and 123 g/mol, respectively. Therefore, CH4 is expected to have the lowest boiling point and SnH4 the highest boiling point. The ordering from lowest to highest boiling point is expected to be CH4 < SiH4 < GeH4 < SnH4.
A graph of the actual boiling points of these compounds versus the period of the group 14 element shows this prediction to be correct:

Order the following hydrocarbons from lowest to highest boiling point: C2H6, C3H8, and C4H10.
- Answer
-
C2H6 < C3H8 < C4H10. All of these compounds are nonpolar and only have London dispersion forces: the larger the molecule, the larger the dispersion forces and the higher the boiling point. The ordering from lowest to highest boiling point is therefore C2H6 < C3H8 < C4H10.
The shape of a molecule affects the strength of its dispersion forces by influencing how much surface area is available for molecular interactions. Molecules with larger surface areas experience stronger dispersion forces because they have more contact points with neighbouring molecules.
For example, boiling points for n-pentane, isopentane, and neopentane (shown in Figure \(\PageIndex{6}\)) are 36 °C, 27 °C, and 9.5 °C, respectively. Even though all three molecules have the same chemical formula, their boiling points are different because their shapes affect the strength of dispersion forces. n-Pentane is elongated, providing more surface area for intermolecular contact. This allows for stronger dispersion forces and a higher boiling point. Isopentane has a more compact shape, reducing the available surface area for interactions, leading to weaker dispersion forces and a lower boiling point. Neopentane is the most compact, offering the least surface area for molecular interactions. As a result, it has the weakest dispersion forces and the lowest boiling point.
This behavior is similar to VELCRO brand fasteners. A long strip of Velcro creates stronger connections because it has more surface area in contact. A small patch of Velcro forms weaker connections because less of it is available for attachment. Likewise, larger molecules with greater surface contact experience stronger dispersion forces, just as longer Velcro strips create stronger connections.
Geckos have an amazing ability to adhere to most surfaces. They can quickly run up smooth walls and across ceilings that have no toe-holds, and they do this without having suction cups or a sticky substance on their toes. And while a gecko can lift its feet easily as it walks along a surface, if you attempt to pick it up, it sticks to the surface. How are geckos (as well as spiders and some other insects) able to do this? Although this phenomenon has been investigated for hundreds of years, scientists only recently uncovered the details of the process that allows geckos’ feet to behave this way.
Geckos’ toes are covered with hundreds of thousands of tiny hairs known as setae, with each seta, in turn, branching into hundreds of tiny, flat, triangular tips called spatulae. The huge numbers of spatulae on its setae provide a gecko, shown in Figure \(\PageIndex{7}\), with a large surface area for sticking to a surface. In 2000, Kellar Autumn, who leads a multi-institutional gecko research team, found that geckos adhered equally well to both polar silicon dioxide and nonpolar gallium arsenide. This proved that geckos stick to surfaces because of dispersion forces. Although dispersion forces are very weak, the total attraction over millions of spatulae is large enough to support many times the gecko’s weight.
In 2014, two scientists developed a model to explain how geckos can rapidly transition from “sticky” to “non-sticky.” Alex Greaney and Congcong Hu at Oregon State University described how geckos can achieve this by changing the angle between their spatulae and the surface. Geckos’ feet, which are normally non-sticky, become sticky when a small shear force is applied. By curling and uncurling their toes, geckos can alternate between sticking and unsticking from a surface, and thus easily move across it. Later research led by Alyssa Stark at University of Akron showed that geckos can maintain their hold on hydrophobic surfaces (similar to the leaves in their habitats) equally well whether the surfaces were wet or dry. Stark's experiment used a ribbon to gently pull the geckos until they slipped, so that the researchers could determine the geckos' ability to hold various surfaces under wet and dry conditions. Further investigations may eventually lead to the development of better adhesives and other applications.

Dipole-Dipole Forces
As discussed in Section 3.6, polar molecules have a separation of charge due to differences in electronegativity. This results in a partially positive end (δ⁺) and a partially negative end (δ⁻), forming a dipole.
Because opposite charges attract, polar molecules experience dipole-dipole attractions—the electrostatic force between the δ⁺ end of one molecule and the δ⁻ end of another. A good example is hydrogen chloride (HCl). In HCl, chlorine (Cl) is more electronegative, so it carries the partial negative charge (δ⁻), while hydrogen (H) carries the partial positive charge (δ⁺). The attraction between opposite dipoles of neighboring HCl molecules creates dipole-dipole forces (Figure \(\PageIndex{8}\)).
The effect of dipole-dipole forces can be seen when comparing the properties of HCl (polar) and F₂ (nonpolar). Both HCl and F₂ have similar molecular masses. At 150 K, the molecules of both substances have the same average kinetic energy. However, HCl remains a liquid, while F₂ stays a gas.
Why? The dipole-dipole attractions in HCl are strong enough to keep molecules close together, preventing them from escaping into the gas phase. In contrast, F₂ molecules only experience weak dispersion forces, which are not strong enough to hold them together as a liquid at this temperature. The higher normal boiling point of HCl (188 K) compared to F2 (85 K) is also a reflection of the greater strength of dipole-dipole attractions between HCl molecules, compared to the attractions between nonpolar F2 molecules.
Since stronger intermolecular forces require more energy to overcome, boiling and freezing points are useful indicators of IMF strength.
Predict which will have the higher boiling point: N2 or CO. Explain your reasoning.
Solution
CO and N2 are both diatomic molecules with masses of about 28 amu, so they experience similar dispersion forces. Because CO is a polar molecule, it also experiences dipole-dipole attractions. Because N2 is nonpolar, its molecules cannot exhibit dipole-dipole attractions. The dipole-dipole attractions between CO molecules are comparably stronger than the dispersion forces between nonpolar N2 molecules, so CO is expected to have the higher boiling point.
Predict which will have the higher boiling point: ICl or Br2. Explain your reasoning.
- Answer
-
ICl. ICl and Br2 have similar masses (~160 amu) and therefore experience similar dispersion forces. ICl is polar and thus also exhibits dipole-dipole attractions; Br2 is nonpolar and does not. The relatively stronger dipole-dipole attractions require more energy to overcome, so ICl will have the higher boiling point.
Hydrogen Bonding
Hydrogen bonding is a particularly strong type of dipole-dipole attraction that occurs when a hydrogen atom is covalently bonded to fluorine (F), oxygen (O), or nitrogen (N)—the three most electronegative elements.
This leads to a large electronegativity difference, creating a strong partial positive charge (δ⁺) on hydrogen and a strong partial negative charge (δ⁻) on the electronegative atom. Additionally, because hydrogen, fluorine, oxygen, and nitrogen are all small atoms, the charged regions can get very close together, further strengthening the electrostatic attraction between molecules. Figure \(\PageIndex{9}\) illustrates hydrogen bonding with the dotted lines between water molecules.
Despite the name "hydrogen bond," these interactions are not actual chemical bonds. They are intermolecular forces—weaker than covalent bonds (only 5–10% as strong) but stronger than regular dipole-dipole attractions and much stronger than dispersion forces.
Hydrogen bonds significantly affect the properties of liquids and solids. For example, consider the boiling points of binary hydrides in groups 15 (NH3, PH3, AsH3, SbH3), 16 (H2O, H2S, H2Se, H2Te), and 17 (HF, HCl, HBr, HI). Figure \(\PageIndex{10}\) shows the boiling points of the heaviest three hydrides in each group. As we move down these groups, the molecule polarities decrease slightly and the molecular sizes increase substantially. Stronger dispersion forces outweigh weaker dipole-dipole attractions, causing boiling points to rise steadily.
If we use this trend to predict the boiling points for the lightest hydride for each group, we would expect NH3 to boil at about −120 °C, H2O to boil at about −80 °C, and HF to boil at about −110 °C. However, when we measure the boiling points for these compounds, we find that they are dramatically higher than the trends would predict, as shown in Figure \(\PageIndex{11}\). The stark contrast between those predictions and reality provides compelling evidence for the strength of hydrogen bonding.
Consider the compounds dimethylether (CH3OCH3), ethanol (CH3CH2OH), and propane (CH3CH2CH3). Their boiling points, not necessarily in order, are −42.1 °C, −24.8 °C, and 78.4 °C. Match each compound with its boiling point. Explain your reasoning.
Solution
The VSEPR-predicted shapes of CH3OCH3, CH3CH2OH, and CH3CH2CH3 are similar, as are their molar masses (46 g/mol, 46 g/mol, and 44 g/mol, respectively), so they will exhibit similar dispersion forces. Since CH3CH2CH3 is nonpolar, it may exhibit only dispersion forces. Because CH3OCH3 is polar, it will also experience dipole-dipole attractions. Finally, CH3CH2OH has an −OH group, and so it will experience the uniquely strong dipole-dipole attraction known as hydrogen bonding. So the ordering in terms of strength of IMFs, and thus boiling points, is CH3CH2CH3 < CH3OCH3 < CH3CH2OH. The boiling point of propane is −42.1 °C, the boiling point of dimethylether is −24.8 °C, and the boiling point of ethanol is 78.5 °C.
Ethane (CH3CH3) has a melting point of −183 °C and a boiling point of −89 °C. Would you expect the melting and boiling points of methylamine (CH3NH2) to be higher or lower than for ethane? Explain your reasoning.
- Answer
-
The melting point and boiling point for methylamine are predicted to be significantly greater than those of ethane. CH3CH3 and CH3NH2 are similar in size and mass, but methylamine possesses an −NH group and can have hydrogen bonding between molecules . This greatly increases its IMFs, and therefore its melting and boiling points. It is difficult to predict values, but the known values are a melting point of −93 °C and a boiling point of −6 °C.
Deoxyribonucleic acid (DNA) is found in every living organism and contains the genetic information that determines the organism’s characteristics, provides the blueprint for making the proteins necessary for life, and serves as a template to pass this information on to the organism’s offspring. A DNA molecule consists of two chains of repeating nucleotides, which form its well-known double helical structure, as shown in Figure \(\PageIndex{12}\).
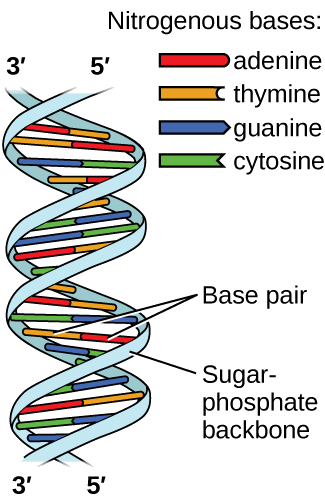
Each nucleotide contains a (deoxyribose) sugar bound to a phosphate group on one side, and one of four nitrogenous bases on the other. Two of the bases, cytosine (C) and thymine (T), are single-ringed structures known as pyrimidines. The other two, adenine (A) and guanine (G), are double-ringed structures called purines. These bases form complementary base pairs consisting of one purine and one pyrimidine, with adenine pairing with thymine, and cytosine with guanine. Each base pair is held together by hydrogen bonding. A and T share two hydrogen bonds, C and G share three, and both pairings have a similar shape and structure Figure \(\PageIndex{13}\).
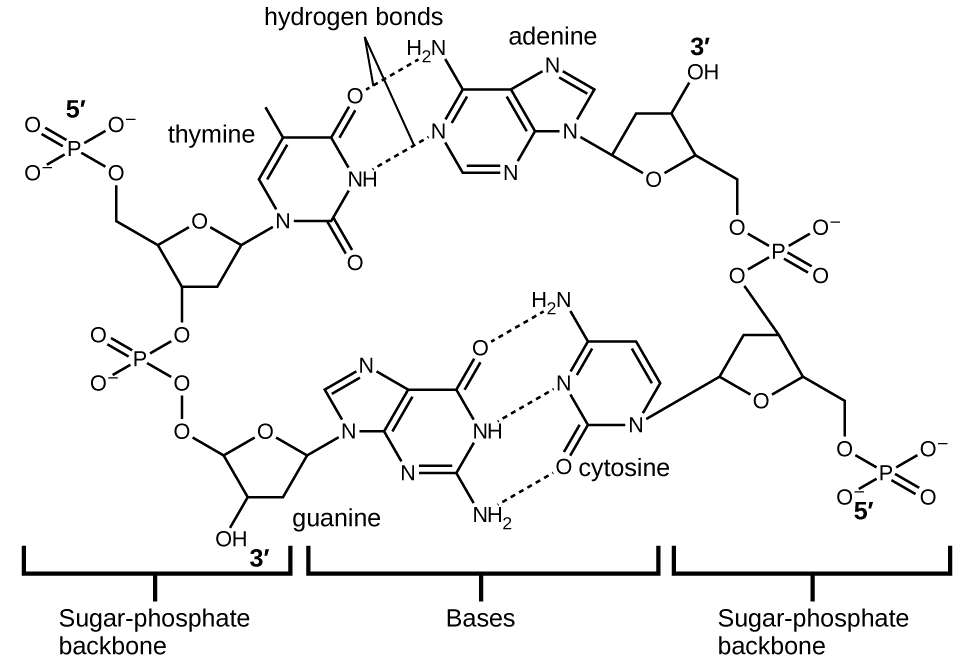
The cumulative effect of millions of hydrogen bonds effectively holds the two strands of DNA together. Importantly, the two strands of DNA can relatively easily “unzip” down the middle since hydrogen bonds are relatively weak compared to the covalent bonds that hold the atoms of the individual DNA molecules together. This allows both strands to function as a template for replication.
Intermolecular Forces Involving Ions
So far, we’ve focused on intermolecular forces between neutral molecules (or atoms), and how the strength of these interactions depends on factors such as size, shape, polarity, and polarizability. But how do ions fit into this discussion?
All intermolecular forces arise from electrostatic attractions between regions of positive and negative charge. The strength of these forces depends on the size and permanence of those charges. Ions, which carry full, permanent charges, interact more strongly than any neutral molecule can. As a result, ion–ion interactions (also known as ionic bonds) are much stronger than any intermolecular force between neutral particles.
These strong interactions are responsible for the structure and stability of ionic solids like sodium chloride. Unlike a covalent bond, which is directional and involves shared electrons between two atoms, an ionic bond is a nondirectional attraction between oppositely charged ions. The ions arrange themselves so that each cation is surrounded by anions and vice versa, forming a highly stable lattice held together by electrostatic forces.
Although ion–ion interactions in ionic solids are very strong, ions can also interact with polar molecules, which have partial charges. For example, when a salt dissolves in water, water molecules surround each ion and form ion–dipole interactions. These are weaker than the original ion–ion attractions in the solid, but many ion–dipole interactions form for each ion pair. This can compensate for the energy required to separate the ions and allow the salt to dissolve.
Because the same physical principle (opposite charges attract) underlie both ionic bonding and intermolecular forces, you can think of ionic bonding as a very strong form of electrostatic interaction, in some ways like an extremely strong intermolecular force between full permanent charges. Depending on the context, it can be helpful to think of ionic bonding either as a chemical bond (when describing salt structures) or as a strong intermolecular force (when considering solubility and ion–dipole interactions). Whether we call it an ionic bond or an ion–ion interaction, the important thing is recognizing that both describe the same physical force, the attraction between full, opposite charges. This attraction explains why ionic solids are stable, why salts dissolve in polar solvents, and why ions interact so strongly with other charged or partially charged species.
Summary
Intermolecular forces (IMFs) are electrostatic forces of attraction between molecules that influence physical properties such as boiling points, melting points, and phase changes.
There are three main types of intermolecular forces:
- Dipole-dipole forces occur between polar molecules, where the partially negative end of one molecule is attracted to the partially positive end of another.
- Hydrogen bonds are a particularly strong type of dipole-dipole force that occurs when hydrogen is bonded to fluorine (F), oxygen (O), or nitrogen (N)—the three most electronegative elements.
- Dispersion forces (also called London forces) arise from temporary shifts in electron distribution that create momentary dipoles, inducing weak attractions between molecules. These forces increase in strength as molecular size and electron cloud polarizability increase.


