Transferrin
- Page ID
- 213340
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)When systemic levels of iron are low within the body, the liver produces transferrin to help regulate iron levels. Transferrin is a blood plasma glycoprotein that helps with the uptake, possession, and transportation of iron(III) throughout the circulatory system of various vertebrates.1 Even though transferrin is found in different vertebrates including mammals, birds, reptiles, amphibians, and fish, this article will focus on the binding of iron(III) to human transferrin. Figure 1 shows that iron is stored in the liver and transferrin removes iron from the liver and transports it in a human’s circulatory system. Hemoglobin is an oxygen-transport metalloprotein founded in red blood cells. Without transferrin’s ability to transport iron from the liver, hemoglobin would not have access to iron. Thus, hemoglobin can not bind to oxygen for transport. As a result, the lack of oxygen transport throughout the body causes apoptosis, cell death.2 With this in mind, transferrin is vital to the overall upkeep and maintenance of the body.
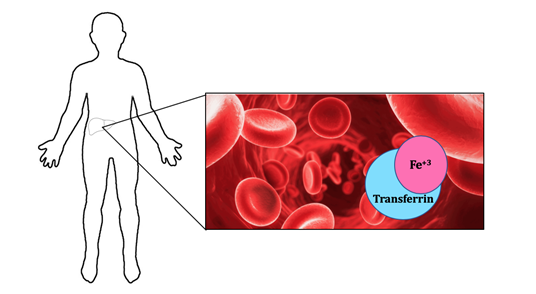
The purpose of this article is to help communicate the various bio-inorganic chemistry topics related to human transferrin and its metal centers. Some of these topics include diseases related to levels of transferrin, structure and function of transferrin, the reaction of iron(III) and transferrin, metal and ion selectivity for the protein, thermodynamics, kinetics, and redox of iron(III). These explanations will help the reader gain a general overview of transferrin through these important chemistry topics.
Structure and Function of Transferrin
Structure of Transferrin
Transferrin contains two identical subunits that contain three domains: N-terminal cytoplasmic tail of 62 amino acid residues, transmembrane segment of 26 amino acid residues, and large extracellular C-terminal region which makes up the remaining polypeptide (Figure 2).3 Apotransferrin, a transferrin molecule without iron(III) bound to it, binds to iron(III) to undergo a conformational change which is a change in the shape of the molecule.4 Iron(III) binds tightly and reversibly to transferrin with high affinity (~1022 M-1) at the two binding sites which are located at the N-lobe or C-lobe of transferrin.5,6 The binding sites are identical, but they are located near different lobes of the molecule. These binding sites have a non-synergistic effect on each other.
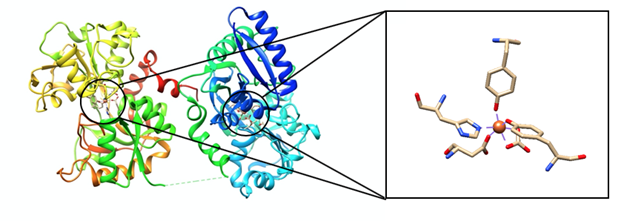
A carbonate anion is an integral part of transferrin binding to iron (III). Carbonate has a synergistic effect with the other ligands involved in transferrin’s coordination environment. Carbonate attaches to transferrin by making hydrogen bonds with amino acid residues not directly involved in the coordination of iron (III). When carbonate binds to iron, it brings with it a large part of the protein. This can be accredited to the placement of hydrogen bonds between amino acid residues and carbonate. In addition to changing the shape of the protein, the coordination environment can now offer iron (III) a tighter more secure binding pocket. This better sized pocket enhances the binding of the other ligands to iron (III) in addition to aiding in the specificity of this pocket for the the atomic radius of iron (III).
The Reaction of Transferrin Binding to Iron(III)
The reaction of iron(III) binding to transferrin is as follows (Figure 3). The hormone hepcidin regulates iron(II) leaving a cell through ferroportin which is a protein that transports iron into and out of a cell. Iron(II) is oxidized to iron(III) with the enzyme hephaestin.8 Iron(III) binds to two tyrosines, one histidine, one aspartic acid, and a carbonate ion in the apo-transferrin’s C-terminus binding site (Figure 2).9 One iron(III) binds to the transferrin and carbonate ion with the release of the three protons. Three protons are released due to ionization of two tyrosine residues and the release of the water molecule that was already bound to iron(III). Another iron(III) binds to transferrin’s N-terminus binding site and an additional carbonate ion.1 The apo-transferrin pKa values for tyrosine, histidine, and aspartic acid are 9.11, 6.00, and 9.60. When iron(III) binds to the amino acid side chains, the pKa values decrease because there is a shift of electron density from the ligands toward the iron(III) which allows the amino acid side chains to deprotonate more easily. When iron(III) is bound to transferrin, the pKa is lower than 7 for tyrosine.10 At physiological pH (7.4), there is a conformational change in transferrin due to iron and carbonate ion binding which causes tyrosine-92 and lysine-285 to be deprotonated. The conformational change also protonates aspartic acid-217 and lysine-277.11 When two transferrin molecules bind to a transferrin receptor 2 (TFR2), the molecules undergo endocytosis which is when substances are brought into the cell. As a result, a vesicle forms to help transport substances throughout the cell.12
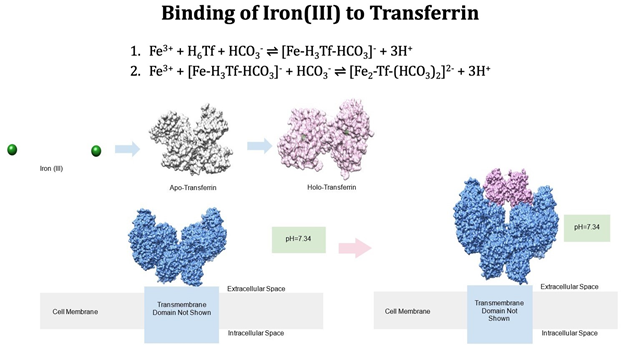
The release of iron is facilitated by the pH change that occurs in the formed endosome (Figure 4). The endosome utilizes a hydrogen pump that changes the pH of the endosome to about 5.5, significantly more acidic than the neutral pH of the blood. At this pH, the transferrin loses affinity for iron primarily due to the acidic pH’s effect on the carbonate anion and the histidine residue. At a pH of 6.0 or below, histidine is deprotonated which allows iron(III) to be released from transferrin.13,14 The increase in hydrogen ions within this vesicle reduces iron(III) reduces iron(II).1
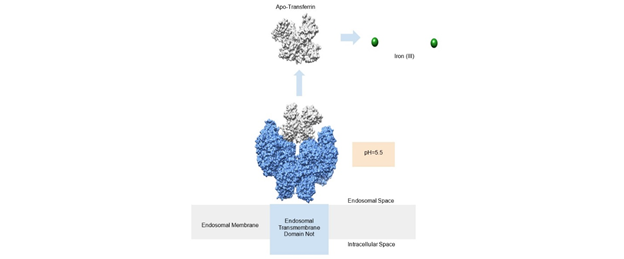
Another source of evidence for the pH dependence of iron (III) releasing from transferrin is Le Chatelier's principle. This principle describes how change in concentration can predict what products will be made in order to get back to equilibrium. Equations one and two (Figure 3) summarize the binding of iron(III) to transferrin. Equation one is the the first iron binding, and equation two is the second iron (III) binding. In both cases, a product of the reaction includes three hydrogen ions. These hydrogen ions are lost from the two tyrosine ligands and the histidine ligand. At an acidic pH, as is the case inside the endosome, there is an increase of hydrogen ions, and therefore an increase in the concentration of products. Le Chatelier’s principle says that the equilibrium will shift to reactants to compensate for this difference. The reactants in each case involve iron (III) no longer being bound to transferrin. This offers another explanation of what happens with transferrin in the acidic endosome.
Diseases Involving Iron
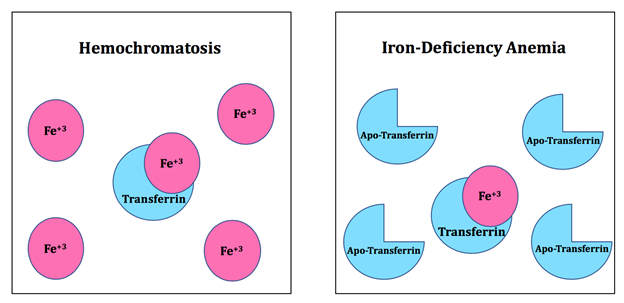
Some diseases may arise due to abnormal iron levels (Figure 5). Hemochromatosis is a chronic iron overload disease which can result from genetic mutations or environmental factors. Hereditary hemochromatosis arises from mutations in the transferrin receptor-2 (TFR2). TFR2 is required for adequate hepatic expression of hepcidin. When TFR2 is impaired, hepcidin production decreases which can lead to hereditary hemochromatosis.15 When hepcidin levels are low, iron overload occurs. There is an increase in ferroportin mediated iron efflux in the liver which causes the liver to store more iron. Ferroportin is a transmembrane protein which transports iron from inside a cell into the outside of a cell.16 In addition, there is an increase in iron absorption within the gastrointestinal tract.17 As a result, the body accumulates iron. Environmental hemochromatosis occurs due to excessive iron in one’s diet. The excess iron produces reactive oxygen species (ROSs) which can lead to cell death and is associated with cardiomyopathic and endocrine disorders.18 On the contrary, iron-deficiency anemia occurs when there is a high concentration of apo-transferrin in the blood. Apo-transferrin is a transferrin molecule that does not have iron(III) bound to it. When the blood does not have enough iron to produce hemoglobin, iron-deficiency anemia occurs.19
Metal Ion Selectivity
Hard Soft Acid Base Theory
Hard Soft Acid Base (HSAB) Theory is a way to help understand the interaction between acids and bases. Like prefers to bind with like, meaning that hard acids prefer to bond with hard bases while soft acids like to bond with soft bases. Three factors determine if an acid or base is hard or soft including charge density, polarizability, and covalent vs. ionic nature of the interactions.20 For this molecule, iron(III) binds to transferrin. Iron(III) is a hard acid and forms bonds with the oxygen of two tyrosine groups, the nitrogen in the imidazole of the one histidine group, the oxygen of the one aspartic acid group, and a carbonate.9 Tyrosine, aspartic acid, and carbonate bond to iron(III) using oxygen donor groups which are hard bases. Histidine is a borderline base. The histidine ring undergoes resonance which allows the nitrogen donor group to form a bond with the iron(III). This bond is not as stable as the bonds between iron(III) and tyrosine, iron(III) and aspartic acid, and iron(III) and carbonate.6 Transferrin’s binding site prefers hard acids which help prevent soft or borderline metals such as \(\ce{Cu^{+}}\) and \(\ce{Zn^{2+}}\) to bind to the molecule. On the contrary, a hard metal such as \({Ca^{2+}}\) could also interact with the binding site due to HSAB.
Ionic Size
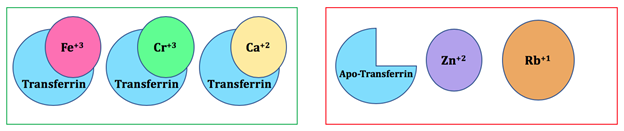
There are other reasons besides HSAB that why iron(III) binds to the active site of transferrin compared to other metal ions (Figure 6). One factor includes ionic size which means that the molecule selects the right ion that fits into the binding site. For example, other metals that might compete with the native iron(III) include chromium(III), zinc(II), and copper(II). These cations have a similar ionic size to iron(III). However, due to HSAB theory, zinc(II) and copper(II) do not bind to transferrin because they are borderline acids. Due to similarity in size, charge and hard character like ferric ion, chromium can be transported and stored in the bloodstream by transferrin.21
Some additional competitive ion examples include sodium and calcium which are both hard metal ions. However, these metal ions are larger than iron(III). Transferrin can bind sodium and calcium, but transferrin preferentially binds to iron(III) more due to its ionic size.22 Rubidium is a hard acid, but it is larger than iron(III). It does not bind to transferrin due to the large metal ion not fitting into the small binding site. This concept is similar to a lock and key. The key (metal) is too large to fit into the lock (binding site). It is reasonable to conclude that ionic size is a factor used to select the correct metal ion for this molecule. Metals with similar ionic size can fit into the active site of transferrin. If a metal is too small, it would not be able to form bonds with the ligands. The metal’s d-orbitals would not be able to overlap with the ligand’s orbitals. Thus, no bond could be formed and the ion would not “fit” into the binding site. For example: metal ions with ionic radii greater than that of europium can only bind to the larger site in the C-terminal domain of transferrin.1 If the size of the metal is different from iron(III), it would affect the shape and function of the molecule. Thus, the ionic size of a metal and HSAB theory help determine the correct metal ion for transferrin.
Electron Spin of Iron(III)
Iron(III) has an oxidation state of +3, five valence electrons, and is a 3d5 metal. There are six ligands including two tyrosines, one aspartic acid, and one histidine from transferrin, and a carbonate ion which binds to iron(III). The shape of this complex is octahedral (Figure 2).23
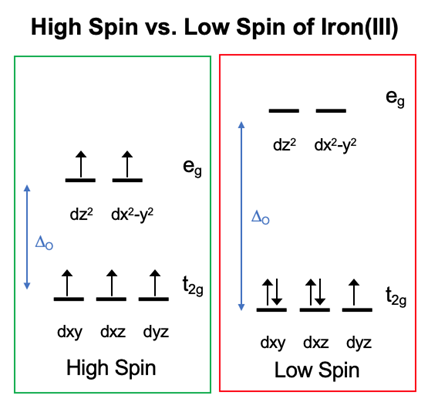
Metal-ligand octahedral geometry can display a range of different d-orbital splitting energies. Metals which are d4 through d7 can have different electron configurations depending on the strength of the d-orbital splitting. In an octahedron, the lower d energy level is t2g while the higher d energy level is eg. When the d-orbital splitting energy level is small, the t2g and eg d orbitals are close in energy (Figure 7). This situation is known as high-spin because electrons can easily enter into the higher orbital. If a d metal is high spin, it has a weak field interaction. When the d splitting energy level is high, the t2g and eg d orbitals are not close in energy. This situation is known as low-spin because electrons pair in the lower orbitals before entering into higher orbitals. If a d metal is low spin, it has a strong field interaction.24
These terms can be applied to the interaction between iron(III) and transferrin. The ligands of transferrin include two tyrosines, one aspartic acid, one histidine, and one carbonic acid ion. Tyrosine, aspartic acid, and carbonic acid are a sigma and pi donor which are weak field ligands and form a small delta o (\(\Delta_{o}\)) and a high spin complex. Histidine is a sigma donor and weak pi acceptor which is a strong field ligand and forms a large \(\Delta_{o}\). Each ligand helps contribute to the overall spin of the metal which is high spin. Overall, the d electrons are in a high-spin conformation because the ligands favor this formation due to the binding site having prearrangement (Figure 7).
Thermodynamics and Kinetics
Chelate Effect
Iron(III) and transferrin is a chelating complex. The chelate effect explains that multidentate ligands bind with higher affinity than a set of monodentate ligands. When the change in Gibbs Free energy is negative, the change in enthalpy is negative, and the change in entropy is positive, the reaction is favorable. The main driving factor for the chelate effect is entropy. When a bidentate ligand replaces two individual monodentate ligands, two ligands are released. The entropy increase is due to an increase in disorder. As a result, entropy increases when a chelating ligand replaces monodentate ligands.25 Enthalpy plays less of a role than entropy for the chelate effect. To remove a bidentate ligand, two coordinate bonds have to be broken which takes energy to break compared to forming a bond.26 For the iron(III) and transferrin complex, carbonic acid is a bidentate ligand and the two tyrosine, aspartic acid, and histidine are monodentate ligands. These ligands are pre-arranged due to protein folding which is why the entire transferrin molecule is a multidentate ligand that chelates iron(III) (Figure 8).
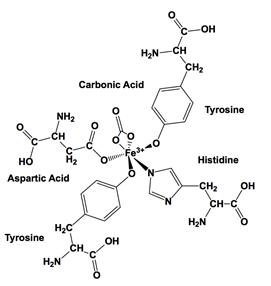
The chelate effect helps increase the entropy during metal association which favors the binding of iron(III) to transferrin. It is unfavorable to remove iron(III) from transferrin. There is a conformational change that occurs under physiological pH due to iron and carbonate ion binding which causes tyrosine-92 and lysine-285 to be deprotonated and aspartic acid-217 and lysine-277 to be protonated.11 At a pH of 6.0 or below, histidine is deprotonated, iron(III) is released from transferrin.13,14 At pH 5.4, the binding constant of apotransferrin to the transferrin receptor is Kd 13x10-9 M.27 Thus, at a pH lower than physiological pH, transferrin releases iron(III).
Macrocycle Effect
The macrocycle effect is similar to the chelate effect; however, it enforces a cyclic conformation of the ligand. When a multidentate ligand is in the cyclic form, the molecule is less likely to move around.26 There is thermodynamic stability and kinetic lability due to entropy and enthalpy being favorable. Entropy increases due to multidentate ligands bonding with the metal instead of monodentate ligands. The enthalpy is favorable because the breaking of bonds releases energy.28 Transferrin’s binding site is not a macrocyclic molecule; however, the binding site is pre-organized.
Lability of Iron(III) with Transferrin
The main function of transferrin is to transport iron(III) ions throughout the body. The iron(III) in the iron(III)-transferrin complex is high spin since the ligands have a set pre-arrangement due to the protein’s folding and the metal's 5 d-electrons. Lability is how easily metal-ligand bonds can break. The more labile a bond is, the easier it will break. On the contrary, the more inert a bond is, it will be harder to break the bond. Since the iron(III)-transferrin structure is high spin, the complex is labile due to eg orbitals (antibonding) being occupied. It is energetically favorable to put electrons into the t2g orbitals because they are lower in energy than eg orbitals. There are 3 electrons in the t2g orbitals which are -0.4\(\Delta_{o}\) each and 2 electrons in the eg orbitals which are +0.6\(\Delta_{o}\) each. This energy difference between the two sets of orbitals causes the Ligand Field Stabilization Energy (LFSE) to be zero. The iron(III) high spin LFSE is zero because the electrons in the eg orbitals have the same energy cost as the electrons in the t2g have the same energy benefit (Equation 1). Sterics influence the geometry and lability of the complex more than LFSE since it is 0\(\Delta_{o}\). Thus, iron(III) is labile.29 When the ligands are farthest apart, the complex is more stable which is why carbonate is sometimes shown as being distorted. Iron(III) is more inert than an iron(II) ion which is due to enthalpy being more negative. A high spin iron(II) has an LFSE of 0.6\(\Delta_{o}\), but a high spin iron(III) has an LFSE of 0\(\Delta_{o}\). Overall, the transferrin-iron(III) complex is labile because the kinetics play more of a factor in determining the rate of the reaction than sterics.30

Electron Interactions
UV-Vis Data of Iron(III) with Transferrin
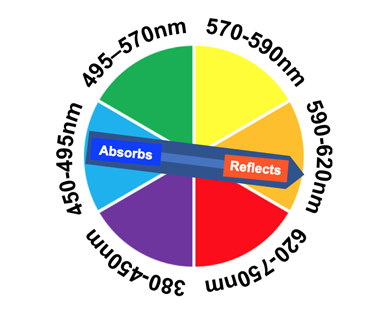
The iron(III) and transferrin complex has a wavelength maximum absorption at 465 nm with an ultraviolet-visible (UV-Vis) spectrometer. On the visible spectrum, 465 nm is in the blue region.31 Since the complex absorbs blue light, it reflects a red-orange color which is what the naked eye sees (Figure 9).32 The extinction coefficient for the iron(III) and transferrin complex at the C-terminus is 87,000 M-1cm-1 at a maximum wavelength of 465nm. The extinction coefficient for the iron(III) and transferrin complex at the N-terminus is 114,000 M-1cm-1 at a wavelength of 465nm.31 The large extinction coefficient suggests that the complex has charge transfer transitions occurring. Since the ligands are pi-donors, it is likely that the complex is experiencing a ligand to metal transfer (LMCT).33
Redox Potentials of Iron(III) with Transferrin
When iron is bound to transferrin, the oxidation state is +3. This oxidation state of iron does not change when transferrin is not bound to the metal due to physiological pH. Electron transfer is accomplished by electrons moving from one molecule to another molecule. A redox catalyst oxidizes or reduces atoms to allow biological reactions to occur due to either the donation or acceptance of electrons. For example, iron(III) is reduced to iron(II) when it is in the vesicle due a lower pH. Iron(III) can easily gain electrons to form iron(II) due to the excess electrons in the surroundings.32,33
Iron(III) is not used as an electron transfer or a redox catalyst in the transferrin system. As a result, there is little research done on the redox potential of iron(III) at physiological pH found throughout the system. Research has shown the reduction potential of iron(III) in free transferrin at endosomal pH (5.6) is below -500 mV. This reduction potential is too low for reduction by physiological agents such as pyridine nucleotides which have reduction potentials of -284 mV. Iron(II) can weakly bind to transferrin, but iron(III) binds stronger.34
Summary
Transferrin is a mammalian glycoprotein that helps with the uptake, possession, and transportation of iron(III) throughout the circulatory system in the body. Iron(III) binds to two binding sites which are located at the N-lobe or C-lobe of transferrin. The binding site contains two tyrosines, one histidine, one aspartic acid, and a carbonate ion. Due to Hard Soft Acid Base (HSAB) theory and ionic size, transferrin binds iron(III) at these binding sites. The five d electrons in iron(III) are in a high-spin conformation because the metal favors this spin conformation due to the binding site having prearrangement. The structure’s Ligand Field Stabilization Energy (LFSE) is zero because the electrons in the eg orbitals have the same energy cost as the electrons in the t2g have the same energy benefit. This structure is also entropically favorable because the protein’s ligands chelate the metal. The iron(III) and transferrin complex can be analyzed with an ultraviolet-visible (UV-Vis) spectrometer at the wavelength of 465 nm because the complex experiences a ligand to metal transfer (LMCT). Overall, transferrin is an important protein in biological systems because it helps transfer iron. Diseases can arise from low levels of iron which is anemia or high levels of iron which is hemochromatosis. Thus, there needs to be further research conducted on this protein to help find ways to cure these diseases. Overall, this article analyzed transferrin and iron(III) through various bio-inorganic chemistry concepts.
Sources
- Chung, M. C.-M. Structure and Function of Transferrin. Biochemical Education 1984, 12(4), 146–154. https://doi.org/10.1016/0307-4412(84)90118-3.
- Siimes, M. A.; Saarinen, U. M.; Dallman, P. R. Relationship between Hemoglobin Concentration and Transferrin Saturation in Iron-Sufficient Infants. Am. J. Clin. Nutr. 1979, 32 (11), 2295–2300. https://doi.org/10.1093/ajcn/32.11.2295.
- Thorstensen, K.; Romslo, I. The Role of Transferrin in the Mechanism of Cellular Iron Uptake. Biochem J 1990, 271(1), 1–9.
- Ogun, A. S.; Adeyinka, A. Biochemistry, Transferrin. In StatPearls; StatPearls Publishing: Treasure Island (FL), 2020.
- Steere, A. N.; Byrne, S. L.; Chasteen, N. D.; Mason, A. B. Kinetics of Iron Release from Transferrin Bound to the Transferrin Receptor at Endosomal PH. Biochim Biophys Acta 2012, 1820 (3), 326–333. https://doi.org/10.1016/j.bbagen.2011.06.003.
- Aisen, P.; Leibman, A.; Zweier, J. Stoichiometric and Site Characteristics of the Binding of Iron to Human Transferrin. J. Biol. Chem. 1978, 253(6), 1930–1937.
- Hall, D. R.; Hadden, J. M.; Leonard, G. A.; Bailey, S.; Neu, M.; Winn, M.; Lindley, P. F. The Crystal and Molecular Structures of Diferric Porcine and Rabbit Serum Transferrins at Resolutions of 2.15 and 2.60 Å, Respectively. Acta Crystallographica Section D Biological Crystallography 2001, 58 (1), 70–80.
- Drakesmith, H.; Nemeth, E.; Ganz, T. Ironing out Ferroportin. Cell Metab 2015, 22(5), 777–787. https://doi.org/10.1016/j.cmet.2015.09.006.
- Baker, E. N.; Lindley, P. F. New Perspectives on the Structure and Function of Transferrins. Journal of Inorganic Biochemistry 1992, 47(1), 147–160. https://doi.org/10.1016/0162-0134(92)84061-Q.
- Sun, X.; Sun, H.; Ge, R.; Richter, M.; Woodworth, R. C.; Mason, A. B.; He, Q.-Y. The Low PK a Value of Iron-Binding Ligand Tyr188 and Its Implication in Iron Release and Anion Binding of Human Transferrin. FEBS Letters 2004, 573(1–3), 181–185.
- Lee, D. A.; Goodfellow, J. M. The PH-Induced Release of Iron from Transferrin Investigated with a Continuum Electrostatic Model. Biophys J 1998, 74(6), 2747–2759.
- Mayle, K. M.; Le, A. M.; Kamei, D. T. The Intracellular Trafficking Pathway of Transferrin. Biochim Biophys Acta 2012, 1820(3), 264–281. https://doi.org/10.1016/j.bbagen.2011.09.009.
- Jon I Mujika, Elixabete Rezabal, Jose M Mercero, Fernando Ruipérez, Dominique Costa, Jesus M Ugalde and Xabier Lopez, ALUMINIUM IN BIOLOGICAL ENVIRONMENTS: A COMPUTATIONAL APPROACH, Computational and Structural Biotechnology Journal, 10.5936/csbj.201403002, 9, 15, (e201403002), (2014).
- Zak, O.; Aisen, P. Iron Release from Transferrin, Its C-Lobe, and Their Complexes with Transferrin Receptor: Presence of N-Lobe Accelerates Release from C-Lobe at Endosomal PH. Biochemistry 2003, 42(42), 12330–12334. https://doi.org/10.1021/bi034991f.
- Functional consequences of transferrin receptor-2 mutations causing hereditary hemochromatosis type 3 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4444164/ (accessed Feb 19, 2020).
- Ward, D.; Kaplan, J. Ferroportin-Mediated Iron Transport: Expression and Regulation. Biochim Biophys Acta 2012, 1823 (9), 1426–1433. https://doi.org/10.1016/j.bbamcr.2012.03.004.
- Papanikolaou, G.; Tzilianos, M.; Christakis, J. I.; Bogdanos, D.; Tsimirika, K.; MacFarlane, J.; Goldberg, Y. P.; Sakellaropoulos, N.; Ganz, T.; Nemeth, E. Hepcidin in Iron Overload Disorders. Blood 2005, 105 (10), 4103–4105. https://doi.org/10.1182/blood-2004-12-4844.
- Kleven, M. D.; Jue, S.; Enns, C. A. Transferrin Receptors TfR1 and TfR2 Bind Transferrin through Differing Mechanisms. Biochemistry 2018, 57(9), 1552–1559. https://doi.org/10.1021/acs.biochem.8b00006.
- Punnonen, K.; Irjala, K.; Rajamäki, A. Iron-Deficiency Anemia Is Associated with High Concentrations of Transferrin Receptor in Serum. Clinical Chemistry 1994, 40 (5), 774–776.
- Libretexts. 3.2.1: Hard and Soft Acid and Base Theory. https://chem.libretexts.org/Courses/Saint_Mary's_College,_Notre_Dame,_IN/CHEM_342:_Bio-inorganic_Chemistry/Readings/Week_3:_Metal-Ligand_Interactions_continued/3.2:_The_identity_of_metal_ion_and_the_ligand_donor_atom(s)_affects_affinity/3.2.1:_Hard_and_Soft_Acid_and_Base_Theory (accessed Jan 28, 2020).
- Harris, D. C. Different Metal-Binding Properties of the Two Sites of Human Transferrin. Biochemistry 1977, 16 (3), 560–564.
- Hemmaplardh, D.; Morgan, E. H. The Role of Calcium in Transferrin and Iron Uptake by Reticulocytes. Biochimica et Biophysica Acta (BBA) - Biomembranes 1977, 468 (3), 423–436. https://doi.org/10.1016/0005-2736(77)90292-9.
- Abdizadeh, H.; Atilgan, A. R.; Atilgan, C. Mechanisms by Which Salt Concentration Moderates the Dynamics of Human Serum Transferrin. J. Phys. Chem. B2017, 121(18), 4778–4789. https://doi.org/10.1021/acs.jpcb.7b02380.
- Libretexts. 4.1.2: Introduction to Ligand Field Theory (Octahedral complexes). https://chem.libretexts.org/Courses/Saint_Mary's_College,_Notre_Dame,_IN/CHEM_342:_Bio-inorganic_Chemistry/Readings/Week_4:_Ligand_Field_Theory_(Octahedral_Complexes)/4.1:_Ligand_Field_Theory_(LFT)_and_Crystal_Field_Theory_(CFT)_of_Octahedral_Complexes/4.1.2:_Introduction_to_Ligand_Field_Theory_(Octahedral_complexes)#Reasons_for_Low-spin_vs._High-spin:_The_Effect_of_the_Metal_Ion (accessed Feb 8, 2020).
- Chelate Effect. Encyclopedia of Inorganic Chemistry 2006.
- Schaller, C. 3.1.2: The Chelate Effect (and Macrocycle Effect). https://chem.libretexts.org/Courses/Saint_Mary's_College,_Notre_Dame,_IN/CHEM_342:_Bio-inorganic_Chemistry/Readings/Week_3:_Metal-Ligand_Interactions_continued./3.1_Ligands_with_more_than_one_donor_atom_(Chelating_Ligands)_have_enhanced_metal_ion_affinity/3.1.2:_The_Chelate_Effect_(and_Macrocycle_Effect)(accessed Jan 28, 2020).
- Dautry-Varsat, A.; Ciechanover, A.; Lodish, H. F. PH and the Recycling of Transferrin during Receptor-Mediated Endocytosis. Proceedings of the National Academy of Sciences 1983, 80 (8), 2258–2262
- Weller, M.; Overton, T.; Rourke, J.; Armstrong, F. A. Inorganic chemistry; Oxford University Press: Oxford, 2014.
- Libretexts. 7.2.2: Trends in M-L lability (Kinetics). https://chem.libretexts.org/Courses/Saint_Mary's_College,_Notre_Dame,_IN/CHEM_342:_Bio-inorganic_Chemistry/Readings/Week_7:_LFT_for_Other_Geometries_//_Applying_LFT_to_predict_Kinetics_of_M-L_Exchange/7.2:_Trends_in_Stability_and_Lability/7.2.2:_Trends_in_M-L_lability_(Kinetics) (accessed Mar 2, 2020).
- Cabantchik, Z. I. Labile Iron in Cells and Body Fluids: Physiology, Pathology, and Pharmacology. Front Pharmacol 2014, 5. https://doi.org/10.3389/fphar.2014.00045.
- Ross, D. C.; Egan, T. J.; Purves, L. R. Periodate Modification of Human Serum Transferrin Fe(III)Binding Sites. INHIBITION OF CARBONATE INSERTION INTO Fe(III) AND Cu(II)-CHELATOR-TRANSFERRIN TERNARY COMPLEXES. J. Biol. Chem. 1995, 270 (21), 12404–12410. https://doi.org/10.1074/jbc.270.21.12404.
- Libretexts. 8.1.1: Colors of Coordination Compounds (Electronic Absorption Spectra). https://chem.libretexts.org/Courses/Saint_Mary's_College,_Notre_Dame,_IN/CHEM_342:_Bio-inorganic_Chemistry/Readings/Week_8:_LFT_continued.Electronic_Properties/8.1:_Absorption_Spectra_and_Magnetic_Properties/8.1.1:_Colors_of_Coordination_Compounds_(Electronic_Absorption_Spectra) (accessed Mar 2, 2020).
- Libretexts. 8.1.2: Predicting UV-visible Absorption. https://chem.libretexts.org/Courses/Saint_Mary's_College,_Notre_Dame,_IN/CHEM_342:_Bio-inorganic_Chemistry/Readings/Week_8:_LFT_continued.Electronic_Properties/8.1:_Absorption_Spectra_and_Magnetic_Properties/8.1.2:_Predicting_UV-visible_Absorption (accessed Mar 2, 2020).
- Redox Properties of Human Transferrin Bound to Its Receptor. https://pubs.acs.org/doi/10.1021/bi0353631 (accessed Mar 5, 2020).
Contributed By:
Whitney Lewis, a chemistry major at Saint Mary's College, class of 2020 and Mary Green, a biology major at Saint Mary's College, class of 2018.


