26.5: Peptides and Proteins
- Page ID
- 36470
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)After completing this section, you should be able to
-
- show, by means of a diagram, how two different amino acid residues can be combined to give two different dipeptides.
- draw the structure of a relatively simple peptide, given its full or abbreviated name and the structures of the appropriate amino acids.
- draw, or name, the six possible isomeric tripeptides that can be formed by combining three different amino acid residues (amino acid units) of given structure.
- account for the fact that there is restricted rotation about the C$\ce{-}$N bonds in peptides.
- illustrate the formation of a disulfide linkage between two cysteine residues, and show how such bonds can link together two separate peptide chains or can provide a bridge between two cysteine residues present in a single peptide molecule.
Make certain that you can define, and use in context, the key terms below.
- C‑terminal amino acid
- N‑terminal amino acid
- peptides
- residues
If necessary, review the discussion of the delocalization of the nitrogen lone‑pair electrons in amides that was presented in Section 24.3. Similarly, you may wish to refer back to Section 18.8 to review the interconversion of thiols and disulfides.
Peptide Bond Formation or Amide Synthesis
The formation of peptides is nothing more than the application of the amide synthesis reaction. By convention, the amide bond in the peptides should be made in the order that the amino acids are written. The amine end (N terminal) of an amino acid is always on the left, while the acid end (C terminal) is on the right. The reaction of glycine with alanine to form the dipeptide glyclalanine is written as shown in the graphic on the left. Oxygen (red) from the acid and hydrogens (red) on the amine form a water molecule. The carboxyl oxygen (green) and the amine nitrogen (green) join to form the amide bond.
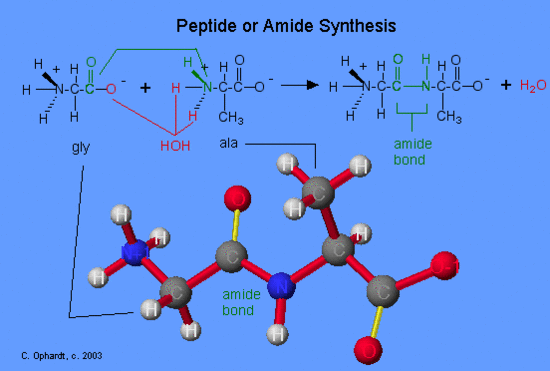
If the order of listing the amino acids is reversed, a different dipeptide is formed such as alaninylglycine.
Resonance contributors for the peptide bonds
A consideration of resonance contributors is crucial to any discussion of the amide functional group. One of the most important examples of amide groups in nature is the ‘peptide bond’ that links amino acids to form polypeptides and proteins.
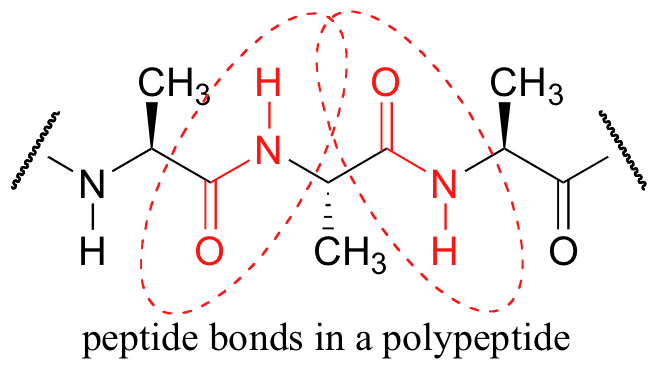
Critical to the structure of proteins is the fact that, although it is conventionally drawn as a single bond, the C-N bond in a peptide linkage has a significant barrier to rotation, almost as if it were a double bond.
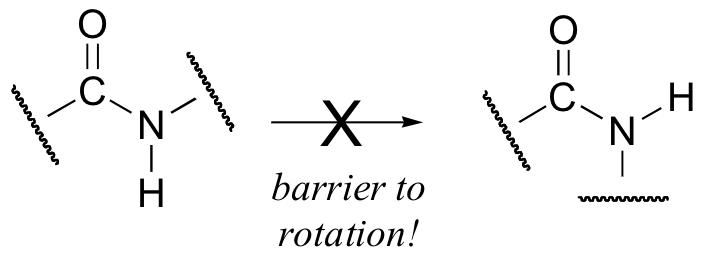
This, along with the observation that the bonding around the peptide nitrogen has trigonal planar geometry, strongly suggests that the nitrogen is sp2-hybridized. An important resonance contributor has a C=N double bond and a C-O single bond, with a separation of charge between the oxygen and the nitrogen.
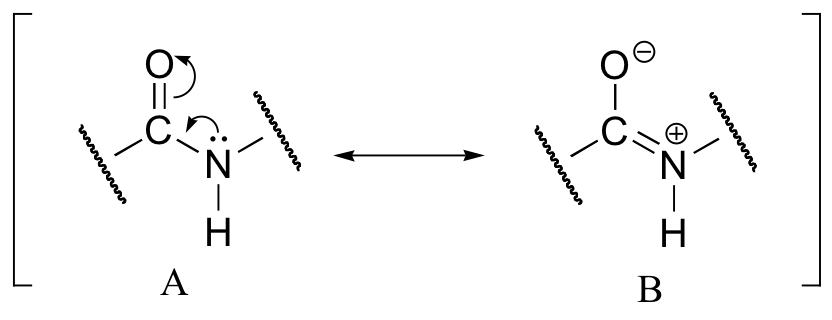
Although B is a minor contributor due to the separation of charges, it is still very relevant in terms of peptide and protein structure – our proteins would simply not fold up properly if there was free rotation about the peptide C-N bond.
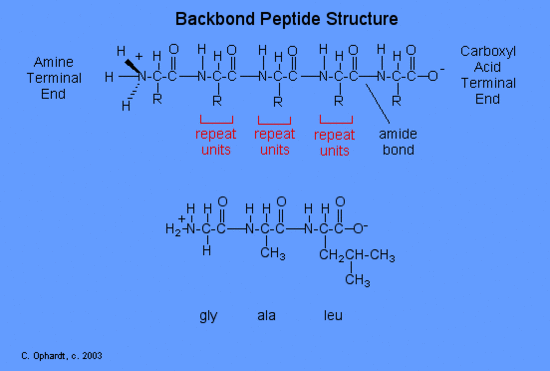
Backbone Peptide or Protein Structure
The structure of a peptide can be written fairly easily without showing the complete amide synthesis reaction by learning the structure of the "backbone" for peptides and proteins.
The peptide backbone consists of repeating units of "N-H 2, CH, C double bond O; N-H 2, CH, C double bond O; etc. See the graphic on the left .
After the backbone is written, go back and write the specific structure for the side chains as represented by the "R" as gly-ala-leu for this example. The amine end (N terminal) of an amino acid is always on the left (gly), while the acid end (C terminal) is on the right (leu).
Write the tripeptide structure for val-ser-cys. First write the "backbone" and then add the specific side chains.
SolutionQUES. Write the structure for the tripeptide:
2 a ) glu-cys-gly ---> Answer graphic
2 b) phe-tyr-asn ---> Answer graphic
Disulfide Bridges and Oxidation-Reduction
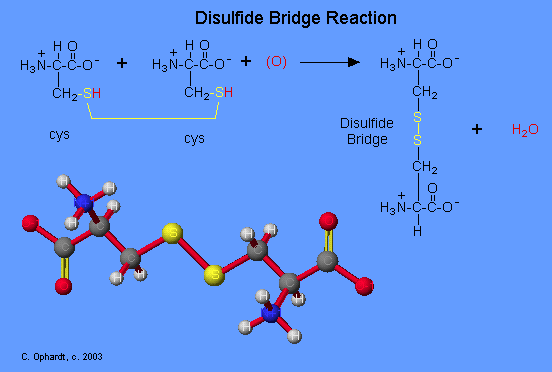
The amino acid cysteine undergoes oxidation and reduction reactions involving the -SH (sulfhydryl group). The oxidation of two sulfhydryl groups results in the formation of a disulfide bond by the removal of two hydrogens. The oxidation of two cysteine amino acids is shown in the graphic. An unspecified oxidizing agent (O) provides an oxygen which reacts with the hydrogen (red) on the -SH group to form water. The sulfurs (yellow) join to make the disulfide bridge. This is an important bond to recognize in protein tertiary structure. The reduction of a disulfide bond is the opposite reaction which again leads to two separate cysteine molecules. Remember that reduction is the addition of hydrogen.
Cysteine residues in the the peptide chain can form a loop buy forming the disulfide bond (—S—S—), while cysteine residues in different peptide chains can actually link what were otherwise separate chains. Insulin was the first protein whose amino acid sequence was determined. This pioneering work, completed in 1953 after some 10 years of effort, earned a Nobel Prize for British biochemist Frederick Sanger (born 1918). He found the primary structure to comprise of two chains linked by two cysteine disulfide bridges. Also note the first peptide chain possesses an internal loop.
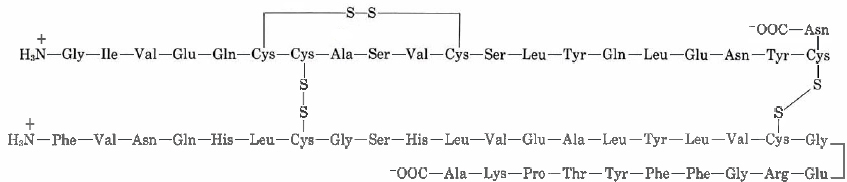
Insulin

