12.8: Protein Misfolding and Denaturation
- Page ID
- 518933
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- Understand the misfolding of proteins and the medical problems associated with it.
- Understand the denaturation of proteins and the factors that cause it.
Protein Misfolding and Associated Diseases
Newly synthesized proteins fold in a specific way, i.e., into secondary, tertiary, and quaternary structures, to be able to perform their function, as illustrated in Figure \(\PageIndex{1}\). Some proteins can fold in only one way, but others can fold in multiple ways. There are proteins in the cell, called chaperones, that help newly formed proteins to fold in the way needed for their function.

Sometimes normal proteins misfold and become pathological. Often, these proteins are in soluble \(\alpha\) helix forms that re-assembles into \(\beta\)-pleated sheet forms that are sticky and aggregate into plaques or amyloid structures, as illustrated in Figure \(\PageIndex{2}\), with the example of plaque formation in Alzheimer's affected brain.

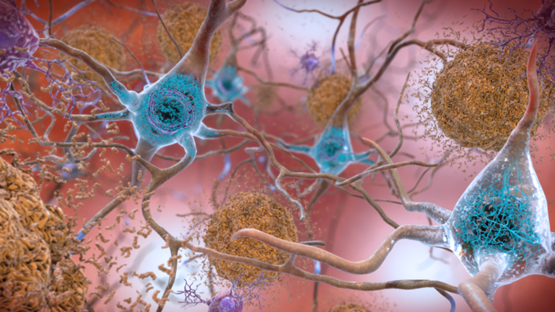
Prions are small proteins found in nerve tissue. Their exact functions are unknown, but when they misfold, they can cause more normal proteins to misfold. This protein misfolding is related to diseases such as mad cow disease in cows, Creutzfeldt–Jakob disease in humans, Alzheimer's disease, and familial amyloid cardiomyopathy or polyneuropathy, as well as intracellular aggregation diseases such as Huntington's and Parkinson's disease.
Denaturation of Proteins
The highly organized structures of proteins are truly masterworks of chemical architecture. But highly organized structures tend to have a certain delicacy, and this is true of proteins. Denaturation is the term used for any change in the three-dimensional structure of a protein that renders it incapable of performing its assigned function.
One of the most important ideas to understand regarding tertiary structure is that a protein, when properly folded, is polar on the surface and nonpolar in the interior. It is the protein's surface that is in contact with water, and therefore the surface must be hydrophilic in order for the whole structure to be soluble. If you examine a three dimensional protein structure you will see many charged side chains (e.g. lysine, arginine, aspartate, glutamate) and hydrogen-bonding side chains (e.g. serine, threonine, glutamine, asparagine) exposed on the surface, in direct contact with water. Inside the protein, out of contact with the surrounding water, there tend to be many more hydrophobic residues such as alanine, valine, phenylalanine, etc. If a protein chain is caused to come unfolded (through exposure to heat, for example, or extremes of pH), it will usually lose its solubility and form solid precipitates, as the hydrophobic residues from the interior come into contact with water. You can see this phenomenon for yourself if you pour a little bit of vinegar (acetic acid) into milk. The solid clumps that form in the milk are proteins that have come unfolded due to the sudden acidification, and precipitated out of solution.
A denatured protein cannot do its job. (Sometimes denaturation is equated with the precipitation or coagulation of a protein; our definition is a bit broader.) A wide variety of reagents and conditions, such as heat, organic compounds, pH changes, and heavy metal ions can cause protein denaturation (Figure \(\PageIndex{1}\)).
| Method | Effect on Protein Structure |
|---|---|
| Heating: Heat above 50°C or ultraviolet (UV) radiation |
Heat or UV radiation supplies kinetic energy to protein molecules, causing their atoms to vibrate more rapidly and disrupting relatively weak hydrogen bonding and dispersion forces (hydrophobic interactions), causing protein denaturation. Example: sterilizing surgical instruments by autoclave treatment. In laser surgery, laser, i.e., the light of a single wavelength, is focused on a spot, causing heat that denatures proteins. Heating by laser cauterizes incisions, i.e., burns the site or the wound. It helps prevent blood loss. |
| Organic compounds: Such as ethyl alcohol | These compounds are capable of engaging in intermolecular hydrogen bonding with protein molecules, disrupting intramolecular hydrogen bonding within the protein. |
| Heavy metal ions: Such as mercury, silver, and lead |
These ions form strong bonds with the carboxylate anions of the acidic amino acids or SH groups of cysteine, disrupting ionic bonds and disulfide linkages. \(\ce{Pb^{2+}}\) and \(\ce{Hg^{2+}}\) attack \(\ce{-SH}\) groups making salt bridges, like \(\ce{-S^{-}Pb^{2+} ^{-}S{-}}\). Egg white or milk is an antidote to heavy metal poisoning as they precipitate the metal ions in the stomach, based on the salt bridge forming reaction. Vomiting is induced to remove the metal before the precipitate is dissolved, releasing the metal in the later parts of the digestive system. |
| Acids and bases: Alkaloid reagents, such as tannic acid (used in tanning leather) | Disrupt hydrogen bonding and salt bridges, e.g., by neutralizing some ions involved in the salt bridges, when these reagents combine with positively charged amino groups in proteins to disrupt ionic bonds. |
| Agitation | physically disrupts hydrogen bonding and hydrophobic interactions. Example: Whipped cream and whipped egg whites are prepared based on the agitation process. |
Anyone who has fried an egg has observed denaturation. With the exposure to heat, the clear egg white turns opaque as the albumin denatures and coagulates. No one has yet reversed that process (Figure \(\PageIndex{3}\)).
However, it is hypothesized that given the proper circumstances and enough time, a protein that has unfolded under sufficiently gentle conditions can refold and MAY again exhibit biological activity (Figure \(\PageIndex{4}\)). Such evidence suggests that, at least for these proteins, the primary structure determines the secondary and tertiary structure. A given sequence of amino acids seems to adopt its particular three-dimensional arrangement naturally if conditions are right.

The primary structures of proteins are quite sturdy. Generally, fairly vigorous conditions are required to hydrolyze peptide bonds. At the secondary through quaternary levels, however, proteins are quite vulnerable to attack, though they vary in their vulnerability to denaturation. The delicately folded globular proteins are much easier to denature than are the tough, fibrous proteins of hair and skin.
Loss of the secondary, tertiary, and quaternary structures of proteins by a physical process or a chemical agent while maintaining the primary structure almost intact is called denaturation of proteins.
In recent years, scientists have become increasing interested in the proteins of so-called ‘thermophilic’ (heat-loving) microorganisms that thrive in hot water environments such as geothermal hot springs. While the proteins in most organisms (including humans) will rapidly unfold and precipitate out of solution when put in hot water, the proteins of thermophilic microbes remain completely stable, sometimes even in water that is just below the boiling point. In fact, these proteins typically only gain full biological activity when in appropriately hot water - at room-temperature they act is if they are ‘frozen’. Is the chemical structure of these thermostable proteins somehow unique and exotic? As it turns out, the answer to this question is ‘no’: the overall three-dimensional structures of thermostable proteins look very much like those of ‘normal’ proteins. The critical difference seems to be simply that thermostable proteins have more extensive networks of noncovalent interactions, particularly ion-ion interactions on their surface, that provides them with a greater stability to heat. Interestingly, the proteins of ‘psychrophilic’ (cold-loving) microbes isolated from pockets of water in arctic ice show the opposite characteristic: they have far fewer ion-ion interactions, which gives them greater flexibility in cold temperatures but leads to their rapid unfolding in room temperature water.


