Copper Chaperone CCS Delivers Cu(I) to Cu,Zn-SOD (SOD1)
- Page ID
- 98115
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Why Does Cu(I) Need a Chaperone?
Copper is an essential element in neurotransmission, metabolic homeostasis, and for superoxide dismutase enzymes in converting reactive oxygen species.1,2,3 Superoxides are one type of reactive oxidative species (ROS). Although a natural byproduct of normal respiration, these species can induce oxidative stress in cells, which can lead to cell death.4 Neurodegenerative diseases like Alzheimer’s exhibit oxidative stress, which is why copper is an important metal ion in superoxide dismutase proteins in combating the formation of oxidative stress.4 Cu/Zn superoxide dismutase 1 (SOD1) uses copper to dismutate superoxides (O2●-) into hydrogen peroxide (H2O2) and oxygen (O2).
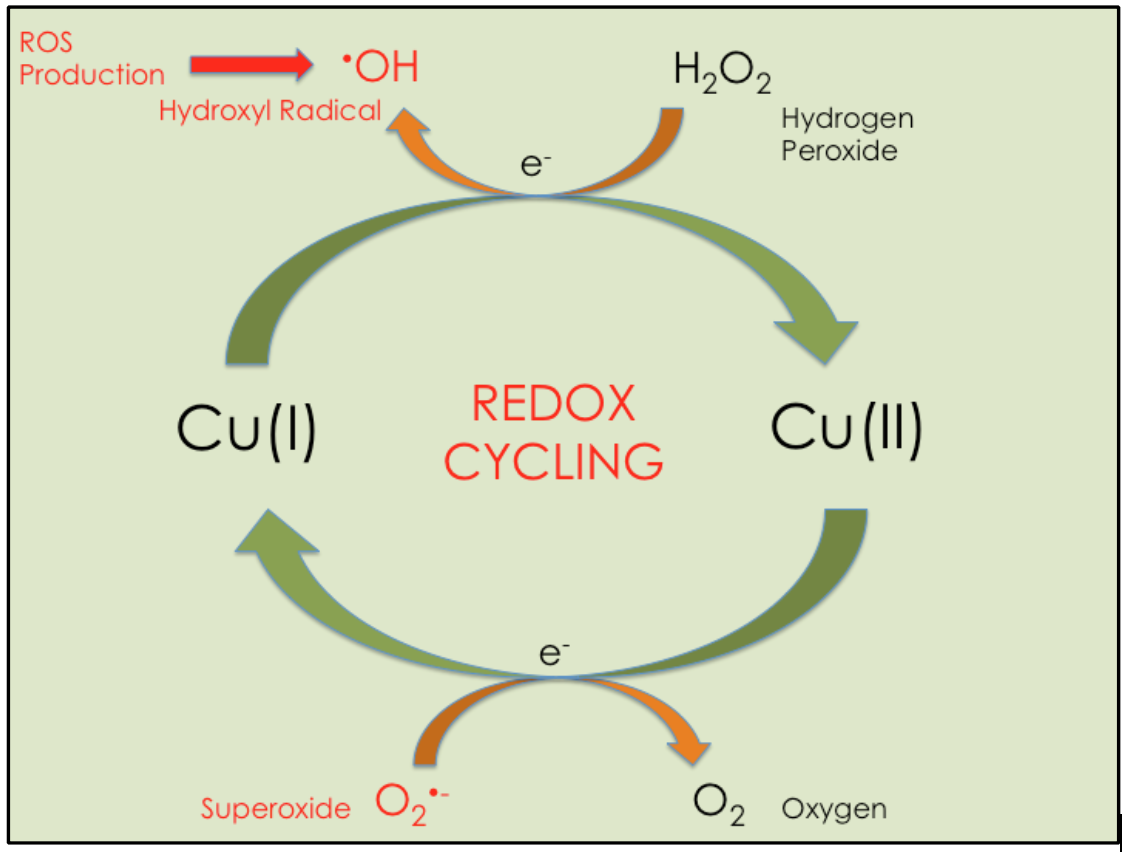
The paradox of copper is its essentiality in cellular redox reactions and its toxicity when unregulated in redox reactions. Unbound Cu(I) can participate in redox cycling reactions because it can donate an electron to oxygen containing molecules, which produces ROS (see Figure 1). The cycle part of the reaction stems from copper’s ability to reverse oxidation states. Cu(I) oxidizes to Cu(II) by donating an electron to an oxygen species, like hydrogen peroxide (H2O2), resulting in ROS production, and Cu(II) reduces to Cu(I) by accepting an electron from an electron donating species, like oxygen radicals. Thus, copper has the ability to produce the species that it is supposed to eliminate if not regulated.
Copper is not incorporated into SOD1 during protein synthesis; it must be delivered. The mechanism by which copper is delivered to proteins like SOD1 is through a copper chaperone (see Figure 2). Copper chaperones transport Cu(I) safely in intracellular regions to other proteins that require Cu(I) for their functions. Without copper chaperones, unbound Cu(I) would rarely get to the target proteins.
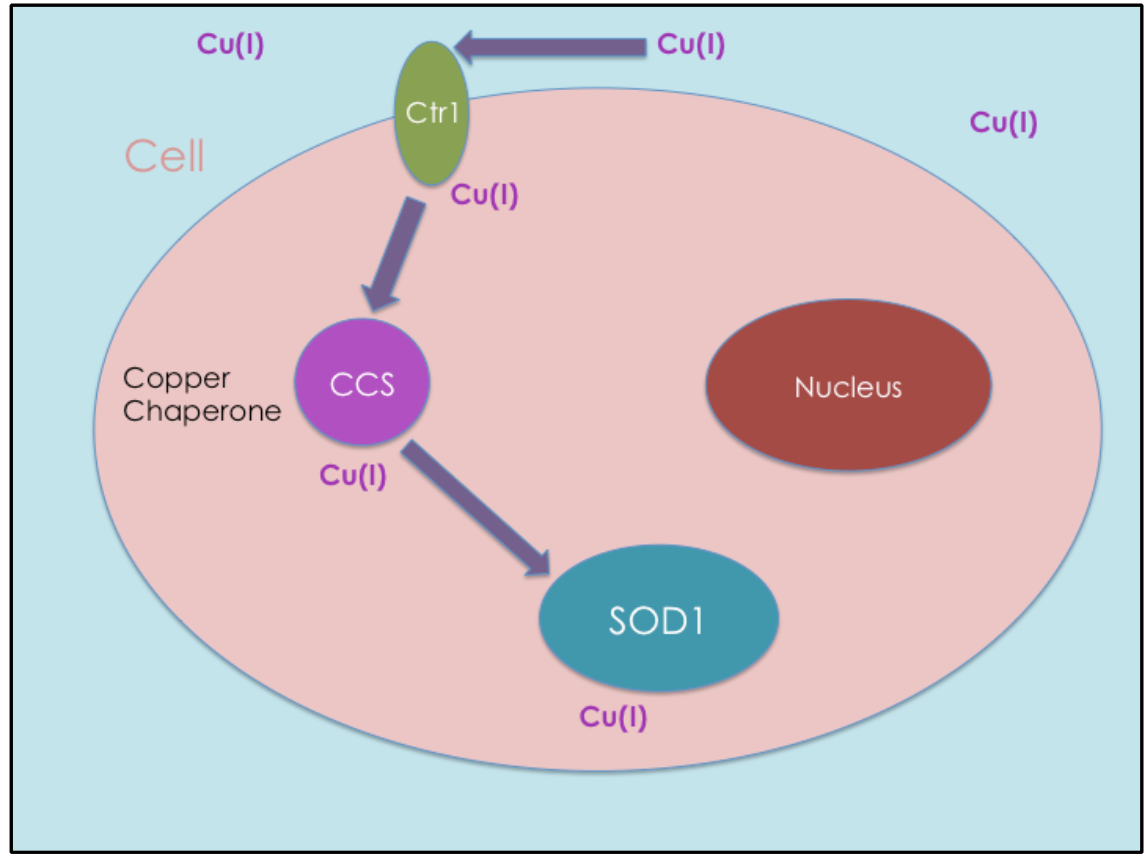
Copper chaperone proteins also aid in regulating Cu(I) from participating in redox cycling reactions. Copper chaperones are required inside of a cell in order to prevent oxidative damage by regulating Cu(I). One chaperone is copper chaperone for superoxide dismutase 1 (CCS).
Human copper chaperone for superoxide dismutase (hCCS) is a metalloprotein that delivers copper(I) to SOD1 inside human cells (see Figure 3). It is made up of 274 amino acids and weighs 54 kDa.5 Inside cells, copper is primarily found in its +1 oxidation state since the intracellular regions of cells are reducing environments.6 Thus, hCCS binds Cu(I) over Cu(II) and other transition metals due to the cytoplasm environment and other chemical principles that will be addressed later.
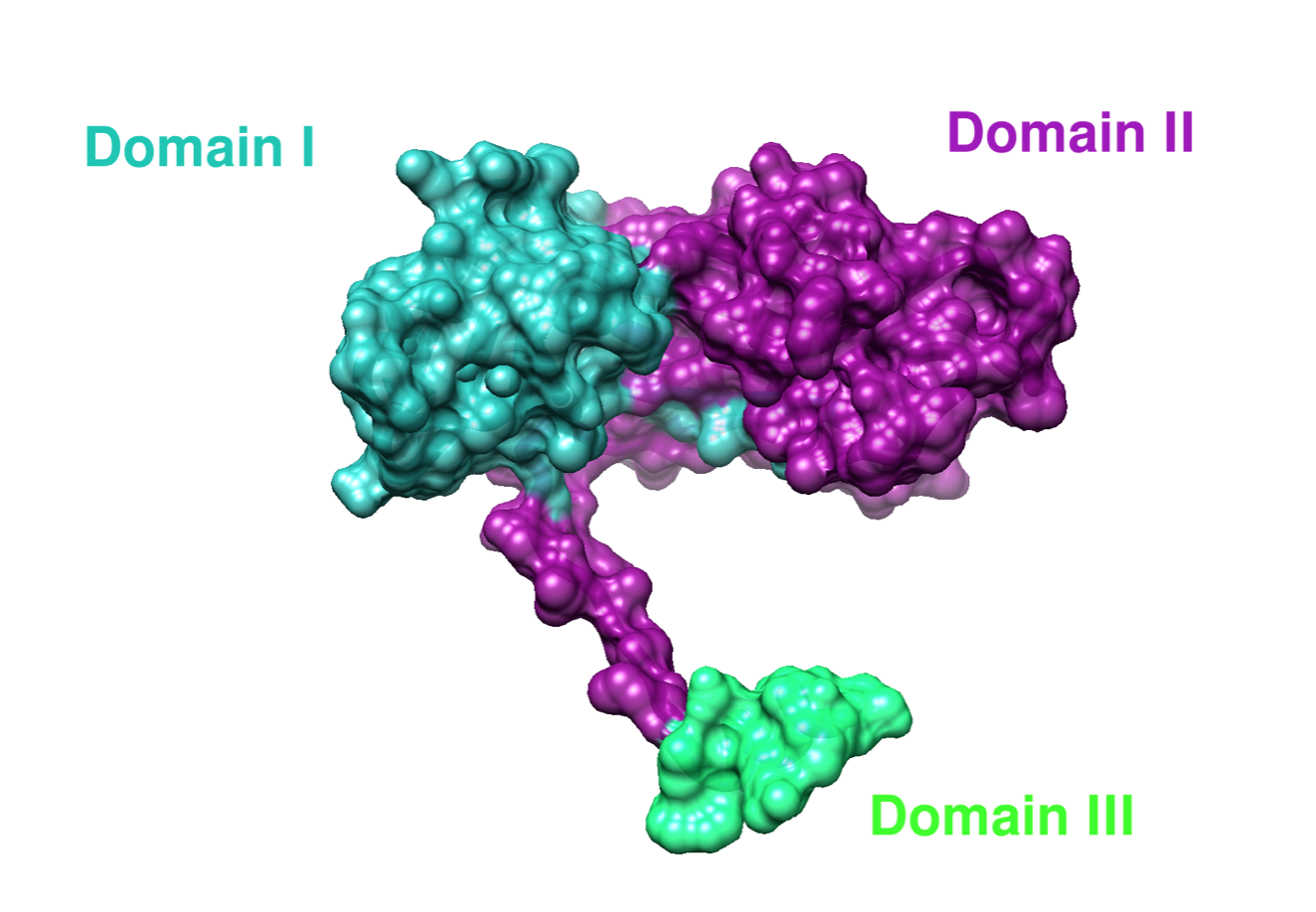
Cu(I) can bind to two out of the three domains of CCS. Zn(II) binds to the second domain of CCS, but Zn(II) function in domain II is unclear.7 Each domain is designed for a specific transport or delivery purpose.8 Domain I has a MXCXXC motif for copper binding, which would include the act of first acquiring Cu(I).8 The role of domain II is to recognize and bind to SOD1.8 Domain III contains a CXC motif, which is involved in the transferring of Cu(I) to SOD1.8 A more specific summary of the 3 domains of hCCS can be found below.
Cu(I) Binding in Domains I and III
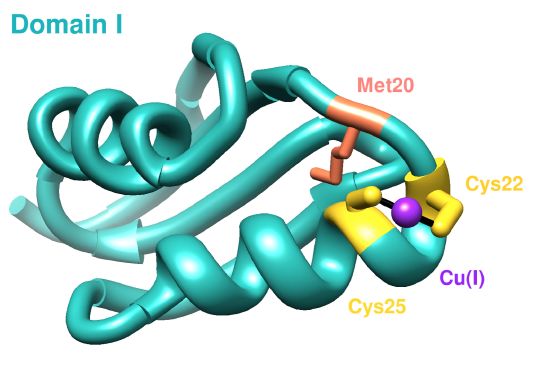
The first domain of hCCS has an NMR structure (see Figure 4). Residues 1-85 compose domain I, and it includes a MTCXXC metal binding motif which is conserved in other copper chaperones for acquiring Cu(I).9 Domain I is important in both searching for copper within cells and accepting copper from membrane-bound copper transporters, like hCtr1.9 The sulfur atoms from Cys20 and Cys22 coordinate Cu(I) in a linear geometry according to the NMR structure (present here) below, but other literature sources have suggested a (third sulfur ligand coordinated with Cu(I)) distorted trigonal planar geometry for Cu(I) in domain I.10 The third ligand to coordinate with Cu(I) would be the sulfur from the Met20 residue.10
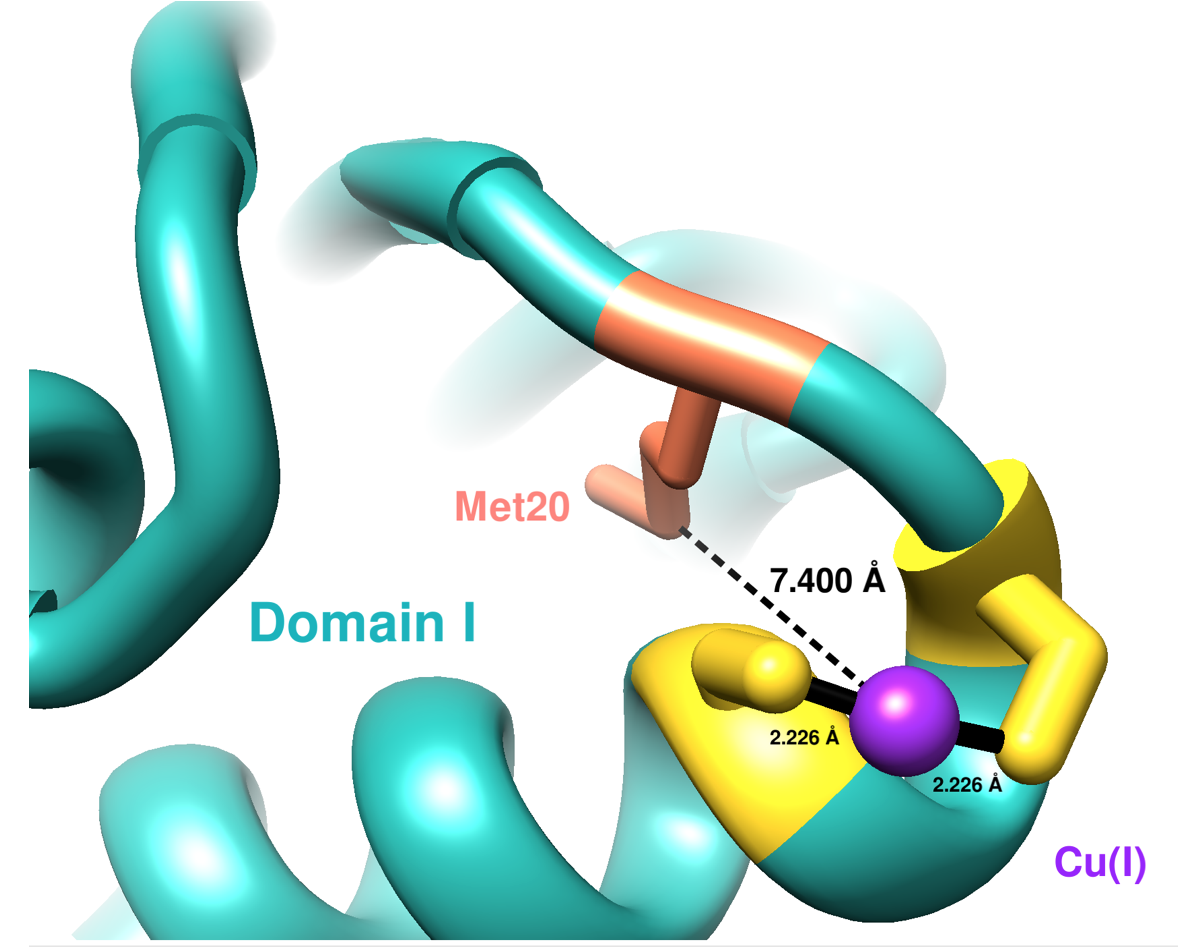
Sterics slightly affect how Cu(I) binds to Domain I of CCS because of both hydrophobic and hydrophilic amino acids surrounding the binding site. The major dictator of the Domain I binding site is geometric constraints, specifically protein folding (see Figure 5). The binding site lies on an alpha helix coil, so the two cysteine residues are positioned perfectly to coordinate Cu(I). Other residues do not participate in binding Cu(I) because of the unfavorable positions on the alpha helix coil.
Domain III
The third domain of hCCS is disordered, so there are few full structures available. However, there is one partial apo yeast x-ray crystal structure from PDB: 1JK9. The remaining residues are not shown due to their disordered nature, which is not picked up by x-ray crystallography. Domain III includes residues 235-274 on hCCS as well as a CXC motif that is proposed to function in metal translocation.9 One experiment showed that the copper binding site in domain III was essential in the CCS-SOD1 transmetalation reaction.9 The candidate ligands involved in binding Cu(I) in domain III on hCCS are Cys244 and Cys246.9 The CXC motif is shown on Cys229 and Cys231 in domain II on yCCS below in Figure 6.
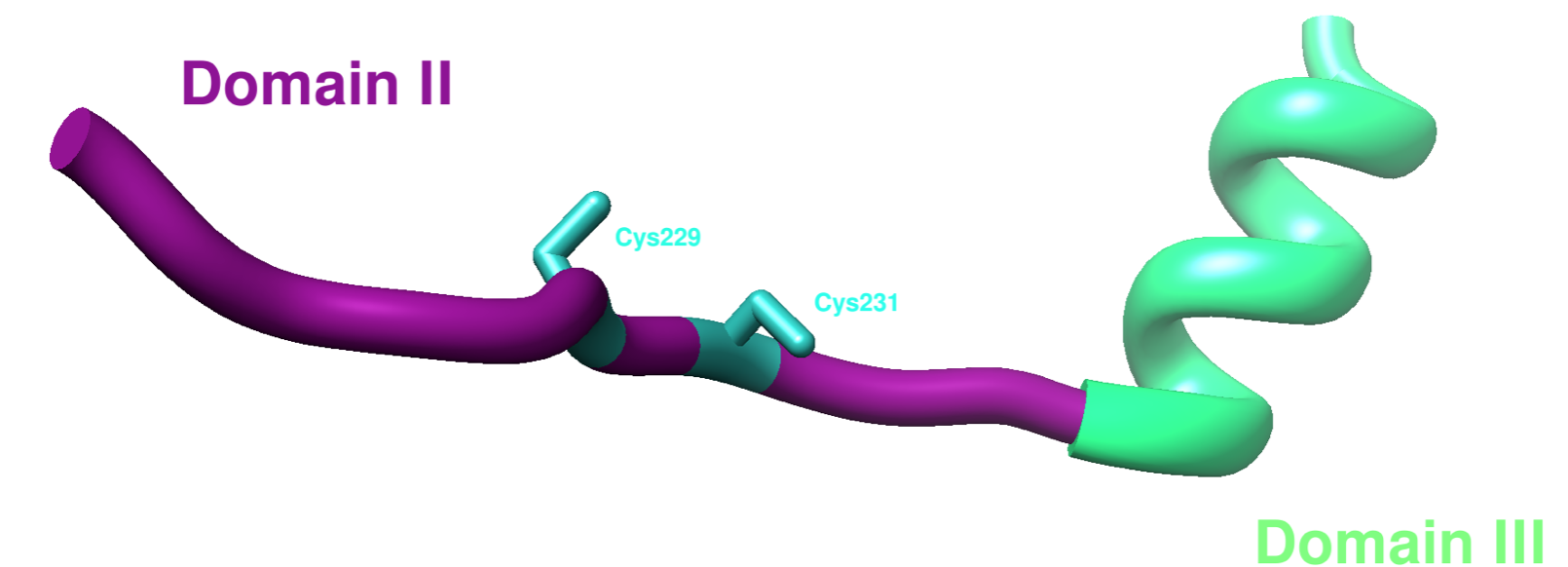
Cu(I) Selectivity
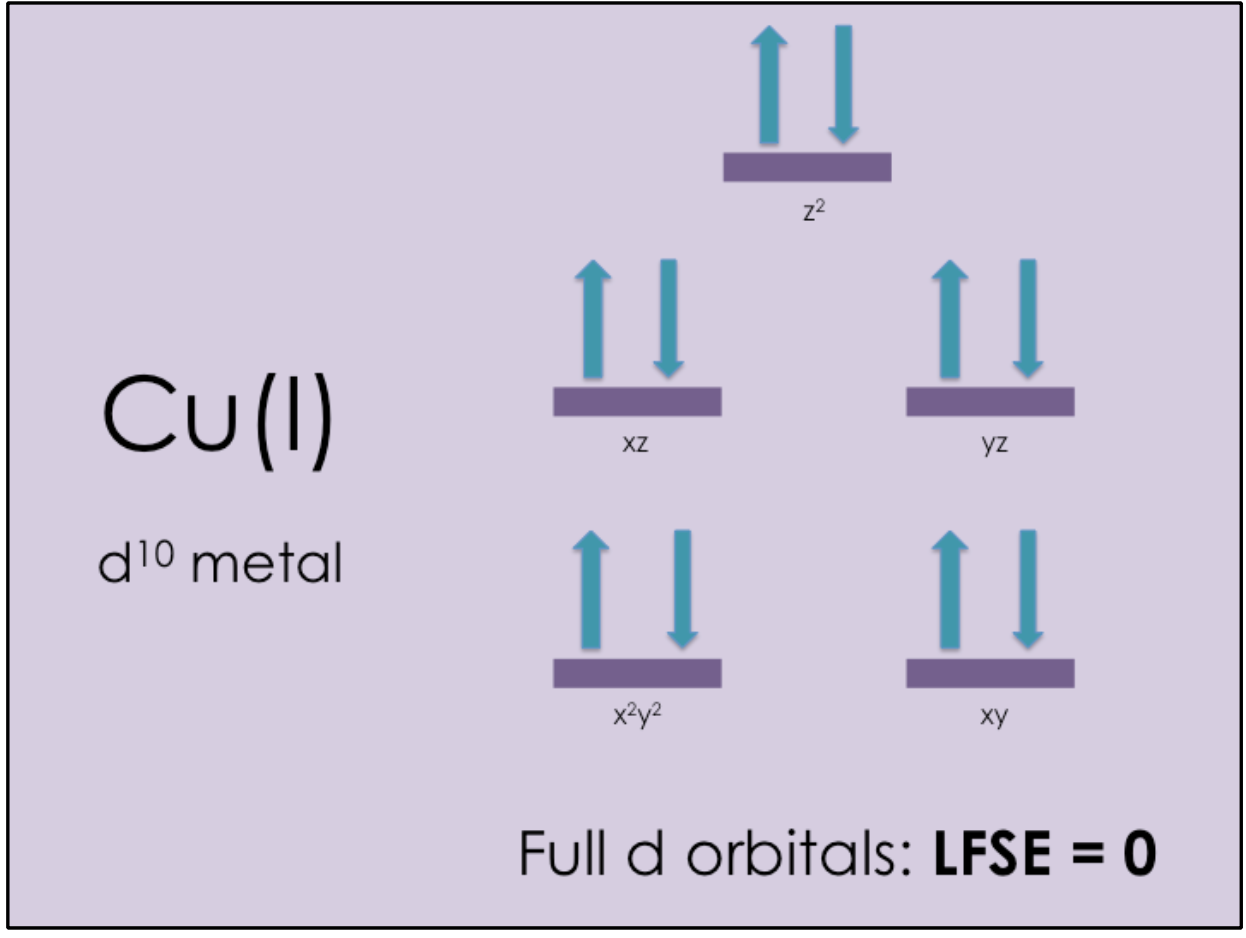
Cu(I) is a d10 transition metal, which means that all of its d electron orbitals are filled. According to the Ligand Field Theory (LFT), any transition metal with full d orbitals has an Ligand Field Stabilization Energy (LFSE) of zero (see Figure 7).
Both domain I and III bind Cu(I) with cysteine residues, which are soft bases according to Hard-Soft Acid-Base theory. Cu(I) is also a soft acid under this theory, so it readily binds to soft sulfur bases.
An LFSE of zero means that no specific geometry is preferred by the metal. That is one reason why Cu(I) can coordinate with two or three ligands and remain stable in a linear or trigonal planar geometry. A trigonal planar geometry has been proposed by Rosenweig and Boal. They propose a third sulfur coordinating with Cu(I).10 Three sulfurs at 2.26 Å are reported to coordinate Cu(I) on hCCS, but which domain this occurs on was not specified.10 The third possible sulfur on domain I could come from Met20, but it is 7.40 Å away from Cu(I) according to Chimera. Thus, sterics and geometric constraints like protein folding dictate which geometry Cu(I) takes on.
An LFSE of zero also means that Cu(I) is labile. Lability refers to the rate at which metal-ligand bonds are broken. When Cu(I) is in excess, domains I and III can quickly acquire and transfer Cu(I) because of Cu(I)’s lability.
Domains I and III select Cu(I) over other transition metals because of many chemical theories and principles. One theory that helps to explain why Cu(I) is selected is the Hard-Soft Acid-Base Theory (HSAB). Based on this theory, hard acids form bonds stronger and faster with hard bases, while soft acids form bonds stronger and faster with soft bases. Metal ions and ligands are differentiated between “hard” and “soft” acids and bases partially due to the charge density of the metal ion. Since Cu(I) has a relatively small charge density, it is considered a soft acid. The types of interactions occurring between the metal ion and ligand also determines whether a metal ion or ligand is “hard” or “soft”. The sulfurs on the cysteine residues form covalent bonds with Cu(I), which makes sulfurs soft bases. Thus, the soft sulfur cysteine side chains preferentially select soft Cu(I) over other transition metals.
According to LFT, Zn(II) also has an LFSE of zero, but neither domain I or III bind Zn(II) because of HSAB. Zn(II) is a borderline acid, so the soft sulfur bases prefer soft Cu(I) over a borderline acid like Zn(II). Likewise, Cu(II) is a d9 transition metal, so its LFSE is not zero. A linear geometry is not favorable for Cu(II), so Cu(I) is preferred.
Zn(II) Binding in Domain II
The second domain of hCCS has one x-ray crystal structure currently in the literature (see Figure 8). Domain II is made up of residues 86-236 on hCCS. It contains a metal binding site for Zn(II) including ligands His147, His 155, His164, and Asp167.7 This Zn(II) binding site is conserved in SOD1.7 Loop 6 in hCCS is proposed to be important in metal delivery and aiding in the recognition of and docking with SOD1.7 Since hCCS domain II does not easily bind copper, the function of domain II is proposed to recognize the target enzyme SOD1 and bind to SOD1 to prepare for Cu(I) transfer.7
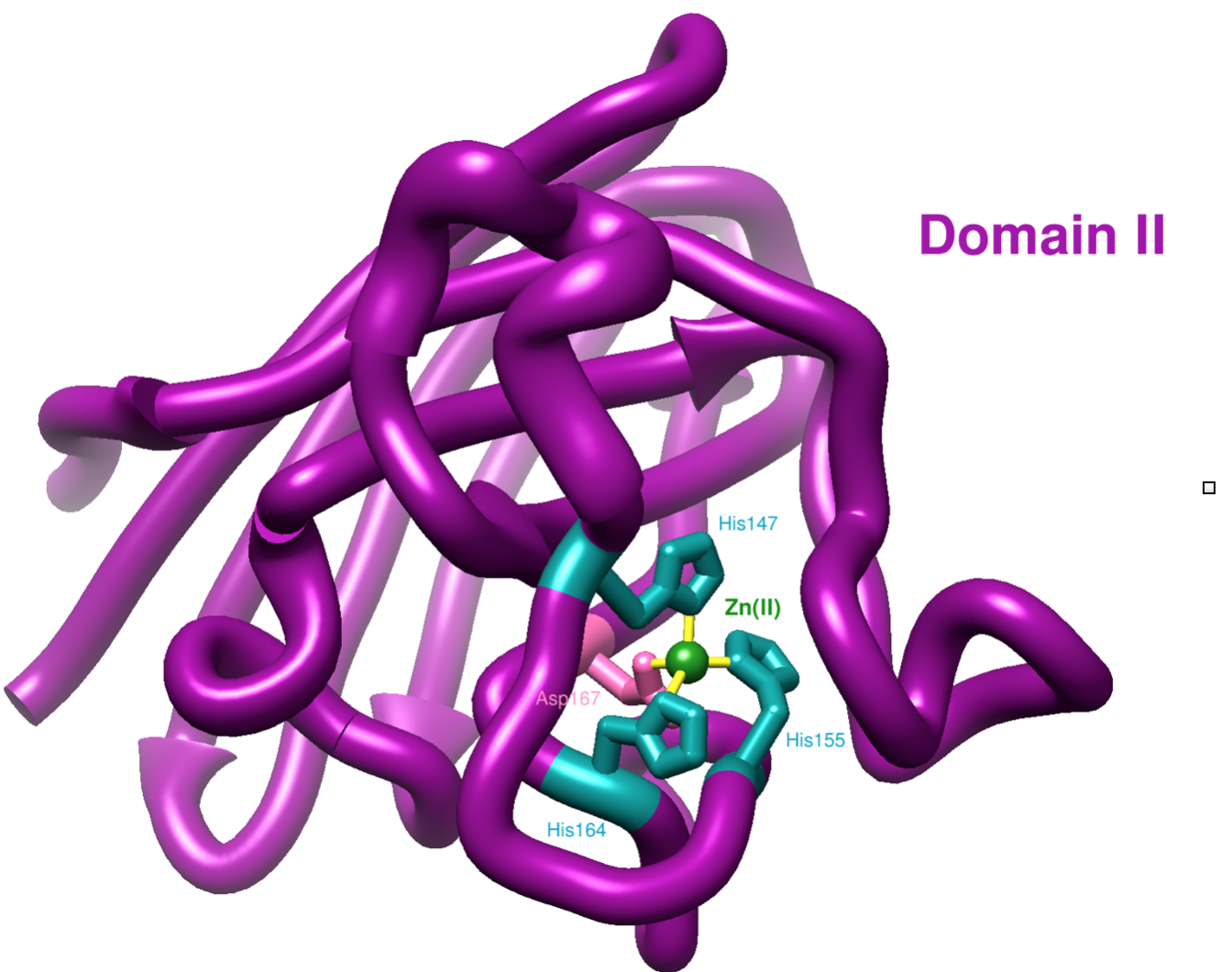
Zn(II) Selectivity
Based on LFT, Zn(II) is a d10 transition metal, which means that it has an LFSE of zero, just like Cu(I). This means that Zn(II) can take on any geometry with no specific preference. The geometry seen above in Figure 8 is a tetrahedral, which is a stable coordination for Zn(II).
However, the three imidazole rings from three histidine residues are borderline bases. Zn(II) is a borderline acid, which is why Zn(II) is preferentially selected over Cu(I) in domain II. Aspartic acid also coordinates Zn(II) even though it is characterized as a hard acid. The exact function of Zn(II) in domain II has yet to be found, but it may play a role in docking CCS to SOD1 during Cu(I) transfer.11
CCS Interactions with Ctr1 and SOD1
CCS functions in several intracellular regions, including the nucleus, cytosol, and mitochondrial intermembrane space.8 The way CCS obtains Cu(I) is still debated. One proposal is that a molecule called GSH (��-glutamyl-cysteinyl-glycine) directly interacts with human copper transporter 1 (hCtr1), then transports Cu(I) to chaperone proteins like CCS.12
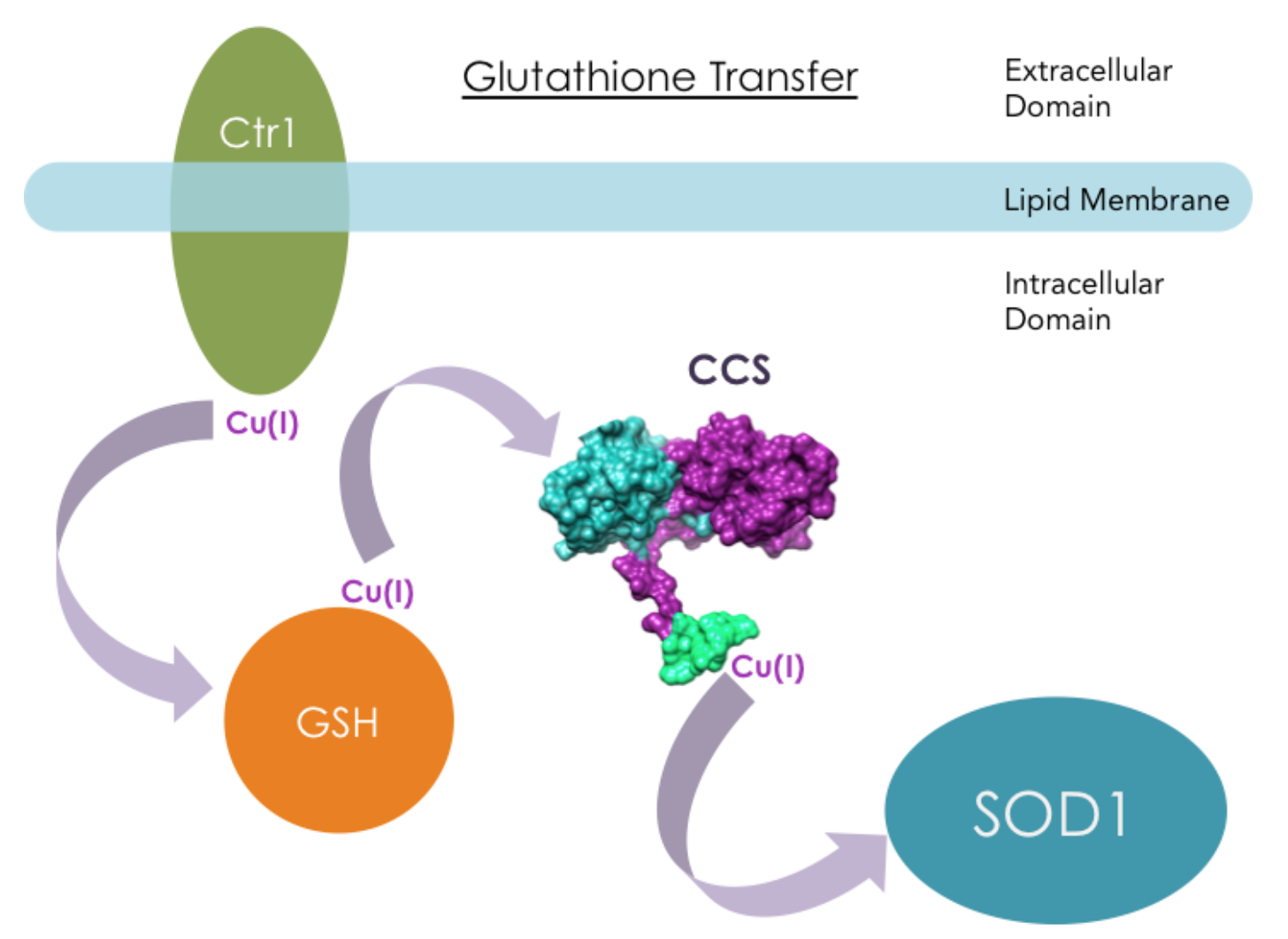
However, it has also been suggested that CCS directly interacts with hCtr1 in Cu(I) transfer. Ctr1 transports extracellular copper across cell membranes into the intracellular region where CCS resides. Experiments from literature have shown that yCCS can bind to lipid membranes, and hCCS has direct interactions with hCtr1 in a copper dependent manner in vitro.13 When Cu(I) was bound to hCtr1, apo-hCCS was able to recognize the presence of Cu(I) on hCtr1 and acquire Cu(I).13 Figure 10 shows another hypothetical Cu(I) transport mechanisms: Direct Transfer.
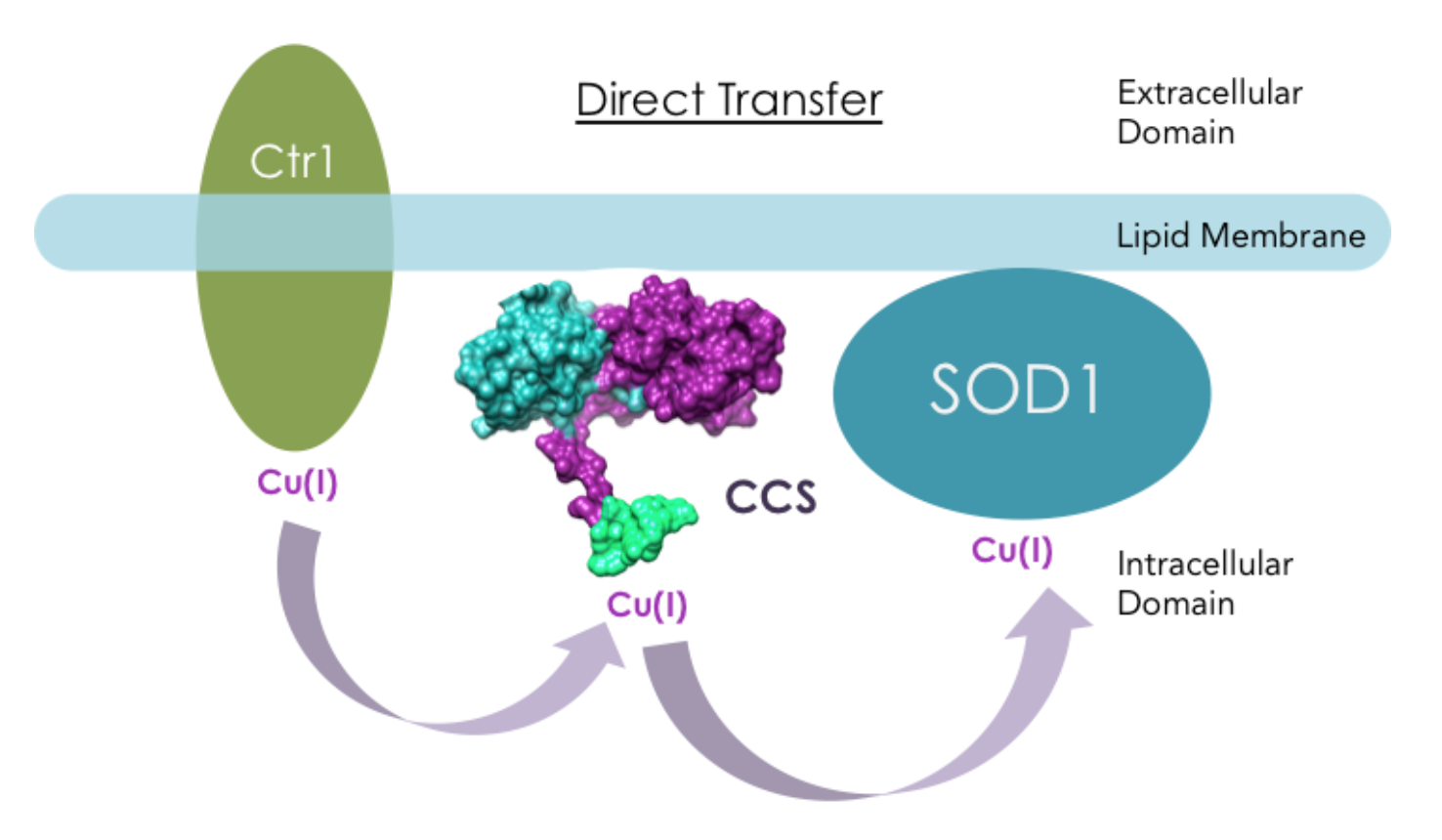
The mechanism by which Cu(I) transfers from domain III on hCCS to SOD1 is unclear. The mechanism has been hypothesized to include three-coordinate intermediates according to Cu(I)-Atx1 coordination chemistry, which is a yeast analog of another copper chaperone.9,14 The formation of three-coordinate intermediates would stabilize Cu(I) from interactions with ROS and ensure safe transfer from CCS to SOD1.
Without proper transportation and transfer, Cu(I) could produce more ROS molecules that cause oxidative stress in cells. CCS plays an important role in protecting cells against oxidative stress by chaperoning Cu(I). Copper chaperones play an important role in both the defense against ROS production. Newly proposed mechanisms of Cu(I) transfer between CCS and SOD1 are important in understanding how neurodegenerative diseases like Alzheimer’s disease develop under oxidative stress. Research on CCS will provide more knowledge on when and how Cu(I) redox chemistry goes wrong.
Sources
[1] Fukai, T., and Ushio-Fukai, M. (2011) Superoxide Dismutases: Role in Redox Signaling, Vascular Function, and Diseases. Antioxidants & Redox Signaling 15, 1583–1606.
[2] Dodani, S. C., Firl, A., Chan, J., Nam, C. I., Aron, A. T., Onak, C. S., Ramos-Torres, K. M., Paek, J., Webster, C. M., Feller, M. B., and Chang, C. J. (2014) Copper is an endogenous modulator of neural circuit spontaneous activity. Proceedings of the National Academy of Sciences 111, 16280–16285.
[3] Balamurugan, K., and Schaffner, W. (2006) Copper homeostasis in eukaryotes: Teetering on a tightrope. Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1763, 737–746.
[4] Ahsan, H., Ali, A., and Ali, R. (2003) Oxygen free radicals and systemic autoimmunity. Clinical and Experimental Immunology 131, 398–404.
[5] CCS (gene). Wikipedia. Wikimedia Foundation. (2007, December 20)
[6] Turski, M. L., and Thiele, D. J. (2009) New Roles for Copper Metabolism in Cell Proliferation, Signaling, and Disease. Journal of Biological Chemistry 284, 717–721.
[7] Lamb, A., Wernimont, A., Pufahl, R., O'Halloran, T., and Rosenzweig, A. (2000) Crystal Structure of the Second Domain of the Human Copper Chaperone for Superoxide Dismutase. Biochemistry 39, 1589–1595.
[8] Inesi, G. IUBMB Life 2016, 69 (4), 211–217.
[9] Eisses, J. F., Stasser, J. P., Ralle, M., Kaplan, J. H., and Blackburn, N. J. (2000) Domains I and III of the Human Copper Chaperone for Superoxide Dismutase Interact via a Cysteine-Bridged Dicopper(I) Cluster†. Biochemistry 39, 7337–7342.
[10] Boal, Amie K., and Amy C. Rosenzweig. “ChemInform Abstract: Structural Biology of Copper Trafficking.” ChemInform, vol. 41, no. 6, Sept. 2010.
[11] Bertini, Gray, H. B., Stiefel, E. I., and Valentine. (2007) Biological inorganic chemistry structure and reactivity. University Science Books Sausalito, California.
[12] Kaplan, J. C. A. H.; Maryon, E. B. Biophysical Journal 2016, 110 (1), 7–13.
[13] Pope, C. R., Feo, C. J. D., and Unger, V. M. (2013) Cellular distribution of copper to superoxide dismutase involves scaffolding by membranes. Proceedings of the National Academy of Sciences 110, 20491–20496.
[14] Pufahl, R. A. (1997) Metal Ion Chaperone Function of the Soluble Cu(I) Receptor Atx1. Science 278, 853–856.
Contributed By:
This work was originally written by Erica Slogar, Spring 2018: Erica is currently (as of 2018) a junior chemistry major at Saint Mary's College in Notre Dame, IN.
This work was originally edited by Dr. Kathryn Haas, Assistant Professor and Madison Sendzik, Teaching and Research Assistant of Saint Mary's College.

