Zinc Transporter Protein: ZntB
- Page ID
- 97958
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Antimicrobial resistance is a problem that is felt world-wide. Antimicrobial resistance occurs when microorganisms (bacteria, viruses, etc) adapt and change to resist the effects of antibiotic treatment. Microorganisms that are able to do this are often referred to as “superbugs”.8 When harmful bacteria become resistant to the antibiotics used to treat them, health care measures become increasingly more difficult and expensive.8 This is a global health care concern because the more resistant the microorganisms are, the more likely it is for disease to spread while prolonging illness and increasing the chances of disability and death.8 Further, antimicrobial resistance is a naturally occurring process over time, but it is the misuse and overuse of current antibiotics that is accelerating the process. Therefore, it is imperative that new drugs are developed in order to target the new “superbugs”. New potential drug targets could include any processes which bacteria and other harmful pathogens harvest resources from the host. Recently, essential micronutrients have been of interest due to their role in maintaining cellular homeostasis. Specifically, the role of zinc in host:pathogen interactions has become of significant interest because both systems rely on its presence for survival. More interestingly, zinc and it’s relative concentrations within the cell has been found to be a helpful tool used to help suppress pathogens and reduce virulence.
Zinc is necessary for life, as many cell processes are Zn-dependent. Zinc is classified as an essential micronutrient as it is important cofactor in many enzyme and transcription factors, a messenger in immune cells, among others.9,10 Additionally, zinc plays an important role in managing host cell integrity and the efficiency of adaptive defense systems.8 Depleted or excess concentrations of zinc can cause toxicity to biological systems. Depletion of zinc can affect many of the Zn-dependent cellular process (proper protein folding, transcription, etc) and cause disarray. Similarly, excess concentrations of zinc can affect cellular processes by allowing unwanted pathogens to thrive. Changes in cellular zinc concentrations are used to reduce the viability of bacteria and other viral pathogens by the hosts either upscaling or downscaling the concentrations of zinc within cellular systems.2 Pathogens require zinc for virulence and for cells to suppress pathogens, intracellular zinc is depleted and exported out of the cell to make it less available for pathogens to scavenge.1, 8-9 To do this, the host cell uses specific zinc transporter proteins such as ZIPs whose function is to either import or export zinc. Similarly, pathogens also have zinc transporter proteins, such as ZntB that are used to scavenge the zinc from the host.
ZntB is a zinc transporter protein found in most prokaryotic organisms that has been observed to import and export Zn2+.1,2 ZntB is a transmembrane protein and there is debate as to whether ZntB operates as an efflux or influx zinc transporter.2 Regardless of whether ZntB is an imports or exports zinc, the protein plays a crucial role in the homeostasis of intracellular zinc concentrations in bacteria. ZntB is of particular interest because it has been isolated from several pathogenic bacteria, including Escherichia coli (E. Coli), Cupriavidus metallidurans, Salmonella typhimurium and Enterobacteriaceae, and it may be a critical virulence factor in these pathogens.2 As ZntB helps to regulate bacterial zinc concentrations, it could be used as a target for new antimicrobial drugs. Since zinc is critical to both human and pathogenic bacterial viability, inhibiting zinc uptake could be used as a method to combat illness.The exact mechanism as to how would require further research but one could speculate that inhibition and/or substitution could be two of the possible approaches. However, although ZntB could be a potential target in antimicrobial drug development, it is a poorly understood protein. Until recently, ZntB was misunderstood to function like other members of the CorA protein family.
Most of the understanding of ZntB is based upon the understanding of the prokaryotic CorA family of Mg2+ transporter proteins.1,4 ZntB is classified as a member of the CorA protein family based on sequence homology. The CorA proteins contain multi-metal binding sites, located both in the pore of the protein and in the extracellular/intracellular domains, and are not specific for Mg2+: they have been observed to transport several other divalent metals such as Ca2+, Mn2+, Co2+ and Zn2+.4 CorA proteins form funnel-shaped pentameric pores in the plasma membrane that transport metals as either their fully-hydrated or partially-hydrated ions.1,2,4 In other words, the metal ions are bound to water molecules as they pass through the protein pore.
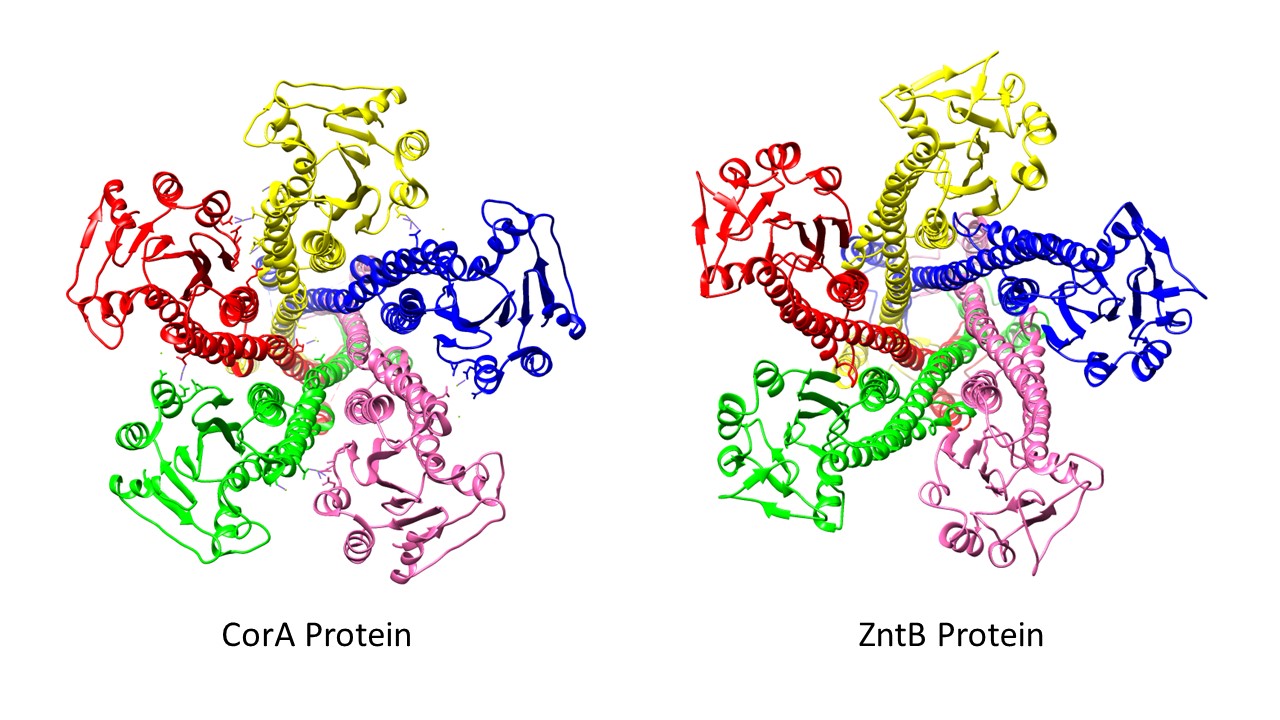
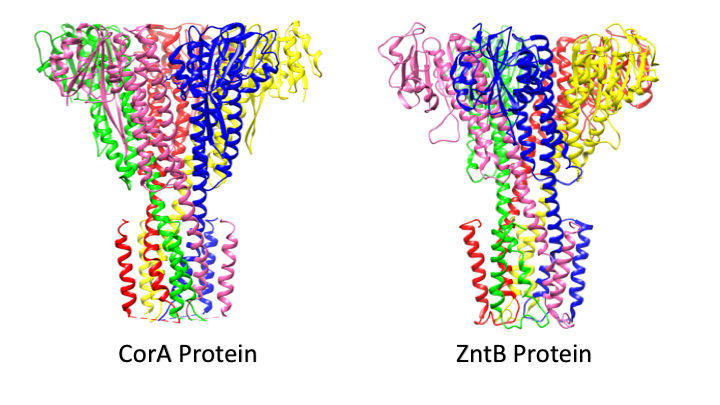
Despite ZntB being a member of the CorA protein family, ZntB shows striking differences to the CorA proteins. The major difference between ZntB and CorA is that CorA proteins can transport several divalent metal ions whereas the ZntB protein is specific for Zn2+.2 It is proposed that the selectivity for the metals is primarily based on to the size and shape of the Zn2+ solvation shells and its unique rate of water exchange.2 Additionally, the pore sizes have been found to be different, as ZntB tends to have a larger pore size than CorA.1,2 In CorA proteins that are selective for specific divalent metals, amino acid interactions with partially hydrated metal ion molecules within the protein pore is found to be critical in cation selectivity.3 However, unlike CorA, there have been no metal binding sites identified for ZntB. As a result, the transportation of Zn2+ is believed to act more as an ion transport channel, with with transport through the pore being relatively simple.1,5 Further, evidence indicates Zn2+ transport by ZntB is coupled with the transfer of an H+ ion, which is also unique to ZntB. Additionally, the ZntB pore in which Zn2+ is transported is embedded with 25 Cl- ions, which is believed to contribute to the electrostatics of the metal ion transport.1 Additional studies are necessary to delineate the exact mechanism of transport with any level of confidence.
Transportation of Zn(II) by ZntB
Based on the current understanding the ZntB system, there is no evidence to suggest that Zn(II) actually binds to the ZntB protein for transport. To date, there have been no identified binding residues. Rather, Zn(II) is thought of as being transported hydrated and acting more as an ion channel.2 In order to transport hydrated Zn(II) through its pore, ZntB have been observed to undergo a conformational change of the pore.This is thought to be caused by the electrostatics of the protein and used to help narrow the selectivity for Zn(II) over other transition metals.
Description of ZntB protein
ZntB is a funnel-shaped pentamer, much like the shape of the CorA protein shown in Figure 1, that spans the plasma membrane.1,2 The CorA and ZntB structures are very similar, with the exception of the ZntB pore being slightly wider than CorA (see figure below). ZntB has a very large N-terminal cytoplasmic domain composed of an αβα motif that then feeds into the α-helices that make up the transmembrane region.2 ZntB has been isolated in a variety of bacterial strains. The resolved cytoplasmic domain structures between the Vibrio parahaemolyticus (VP-ZntB) and Salmonella typhimurium (ST-ZntB) differ significantly with a difference of 9.5 A°.2 This is indicative of two different states of the transport cycle as the sequencing of the proteins are nearly identical.2 Having two identified transport states suggests that the ZntB protein undergoes a conformational change. In addition to different crystalline structures, a change in electrostatics has been to seen to play a role in the conformation and selectivity of ZntB.
ZntB has been crystallized in its full length form as well as just the cytoplasmic domain. The two ribbon structures are shown below. Researchers noted two primary differences between the structures. Despite both being absent of Zn(II), the cytoplasmic domain was crystallized with 25 Cl- atoms. The Cl- atoms were embedded into the pockets of the β-sheets and ⍺-helices and observed to be bound to specific amino acid residues.1 The binding of the Cl- atoms is thought to contribute to the widening of the pore by adjusting the hydrogen bonding and salt bridges between protein components, therefore affecting the overall protein pentamer pore structure. Static field potentials of both structures have been calculated and the results indicate that the chlorine atoms contribute significantly to a more negative potential across the protein pore.1 Electrostatic free energy of transferring a cation from bulk into the pore (ΔΔGint) have also been calculated and revealed that the affinity of a positive divalent cation to the protein pore relies heavily on the presence of the chlorine atoms.1 Therefore, the binding of the Cl- atoms to specific amino acids in the pore to cause a slight widening of the protein pore and the change in potential is expected to be essential in Zn(II) selectivity and transport. Shown below is a comparison of the full length ZntB structure with the cytoplasmic ZntB structure that has the embedded chloride ions.
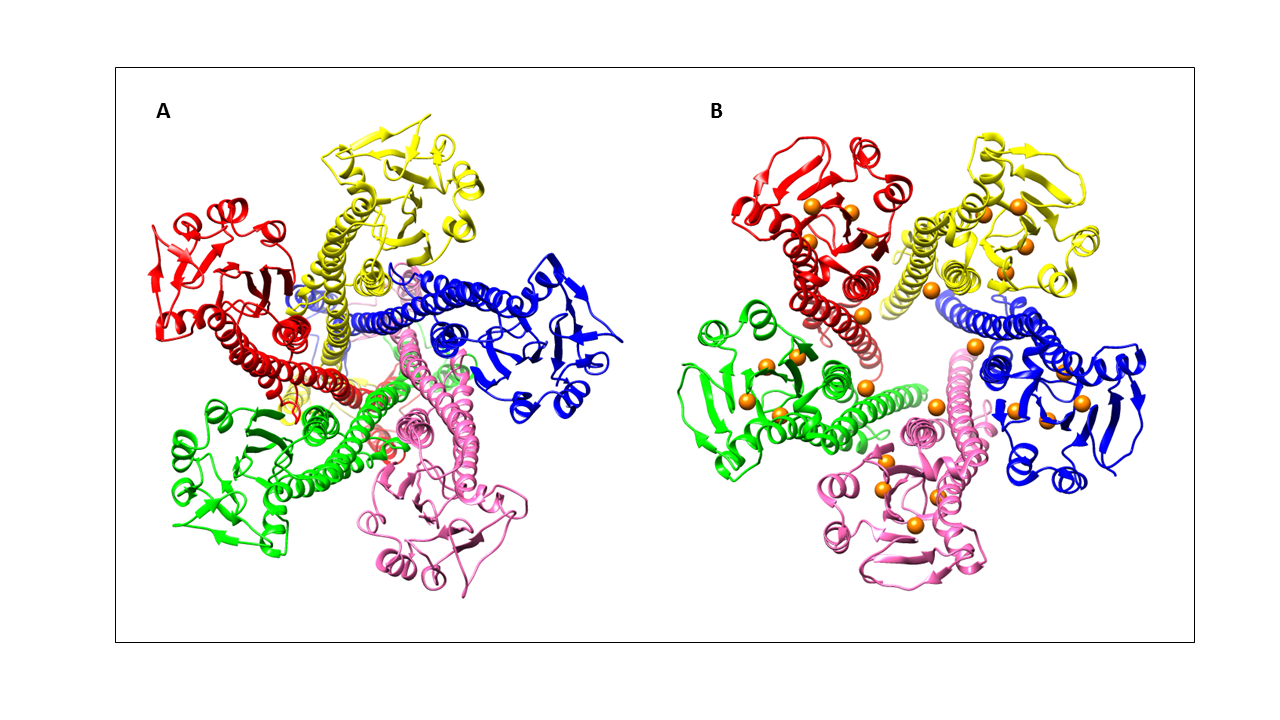
Figure 2: A: Full length ZntB structure (PDB: 5N9Y). Structure was resolved with no Zn(II) and no Cl- atoms. B: Cytoplasmic domain of ZntB structure (PDB: 3CK6). Structure was resolved without Zn(II) bound but 25 Cl- atoms (shown in orange) are bound to specific amino acid residues.
In addition to the transport of Zn(II), a H+ transfer has also been observed and is proposed to be key factor for Zn(II) uptake by ZntB. The transport of Zn(II) by ZntB has been observed to be coupled with a proton transfer through experiments that monitor Zn transport by ZntB under different pH gradients. These experiments have shown that the uptake of Zn(II) by ZntB is stimulated by a decrease in the pH gradient, suggesting an increase in the presence of protons.2 The evidence demonstrates that when H+ ions are transported from the extracellular region to the intracellular region, this also stimulates Zn(II) transport by ZntB, indicating that the proton and zinc ion are transported together and in the same direction. To validate these results, further experiments were run in more basic conditions. At the point where Zn(II) uptake had reached a plateau, a proton ionophore FCCP was introduced extracellularly to the system which consequently stimulated additional uptake of Zn(II) by ZntB.2 Further, when the pH was higher intracellularly than it was extracellularly and the proton ionophore was introduced, the efflux of Zn(II) was stimulated.2 Compoundingly, this suggests that successful transport of Zn(II) by ZntB is coupled with a proton transfer. However, the exact mechanism of transport of zinc by ZntB is still elusive. In addition to the transport of Zn(II) being coupled with a proton transfer and the lack of Zn(II) binding to ZntB, Zn(II) is suggested to be transported fully hydrated. The hydration can be either full or partial, with fully hydrated being thought as the more probable condition at this time.
Aquated Zinc
Due to the zinc ion being transported either fully or partially hydrated, it can be described as an aquated zinc ion. Zn2+ is a borderline metal (acid), and according to Hard Soft Acid Base Theory, it should prefer to bind to borderline ligands (bases). However, the oxygen atom of water is a hard base. Due to Zn2+ being a borderline metal, it is reasonable that it could interact with hard ligands. In aqueous solutions, Zn(II) tends to behave more like a hard acid.6 In other words, in the strong presence of water and no readily available soft ligands, zinc is likely to bind to the hard oxygen in water. This is not as much due to preference by the zinc ion, rather the abundance of water acting as the driving force according to Le Chatelier's principle. The reaction of zinc with water is illustrated below. Additionally, water is a neutral ligand. In an ideal situation, Zn(II) would prefer to bind to a ligand with a negative charge because it itself has a positive charge. Therefore, when water is abundant, it will drive the hydration process, which then allows Zn2+ to become hydrated in physiological conditions.6
Zn2+ + nH2O → Zn(H2O)n
The reaction shown above is in fact favorable, as evidenced by a negative enthalpy of hydration. Zn(II) has a relatively high Zeff compared to other 3d transition metals, and would form the most stable complexes with water among other 3d transition metals according to the Irving-Williams series.
Based on Taube’s rules we can expect Zn(II) to be relatively labile. Zn(II) has a full d-orbital with an ligand field stabilization energy (LFSE) of zero. With a full d-orbital, there are electrons in the antibonding orbitals which promotes lability and the lack of LFSE energy does not make the zinc molecule more stable when bound to ligands. Water is a borderline pi donor/acceptor ligand. Pi donors tend to increase lability because they decrease Δ, which decreases the LFSE values. However, since Zn(II) already has a full d-orbital, the water molecules can’t donate to the antibonding d-orbitals and therefore, the ligand effects on lability and overall stability are minimal in this case. The splitting diagram for Zn(II) is shown below.
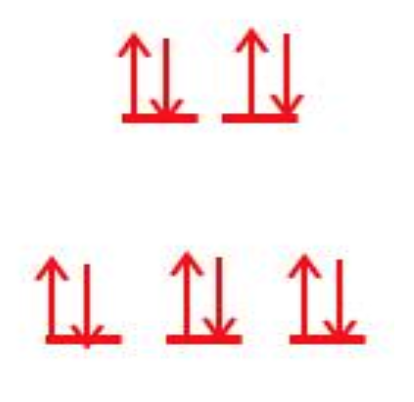
Figure 3: The d-orbital splitting diagram of Zn(II). Zinc(II) has 10 valence electrons, which fills all 5 of the d-orbitals.
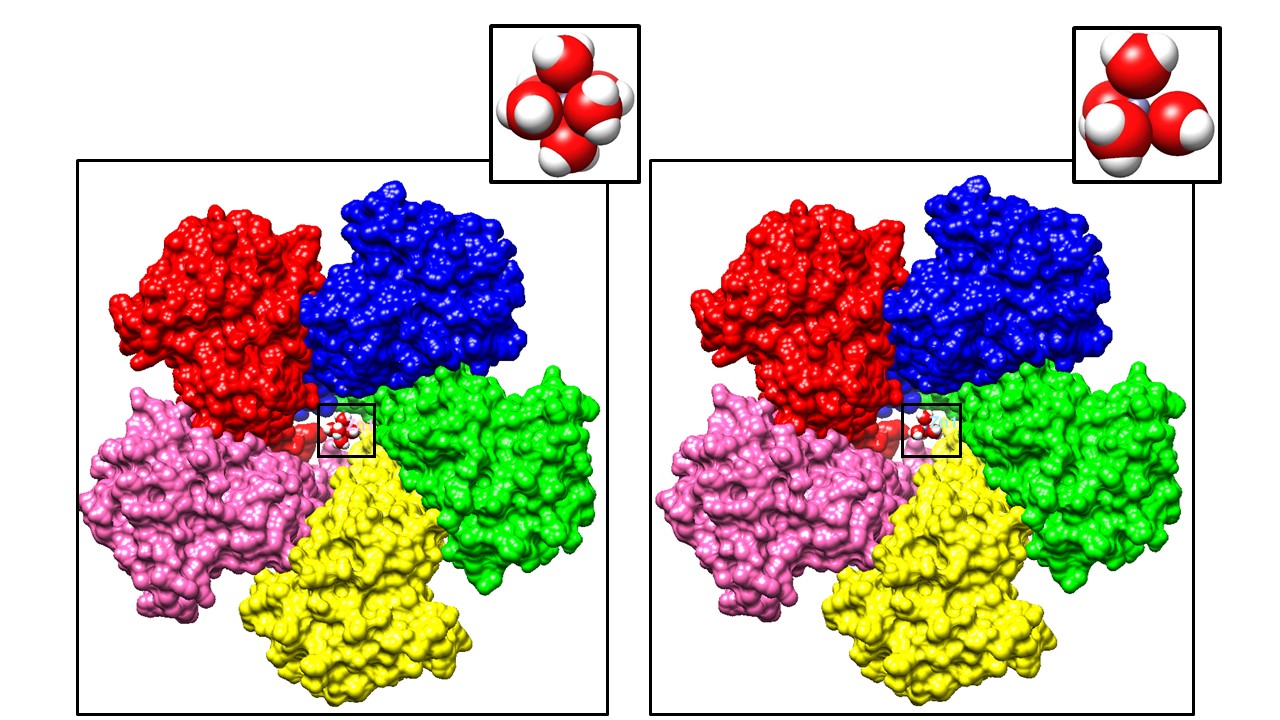
As mentioned previously, Zn(II) has a full d-shell and as a result, is not subjected to the consequences of LFSE.6 Due to lack of LFSE, Zn(II) has no preference for any specific geometry. In fact, Zn(II) is primarily found as a mixture of tetrahedral, octahedral, and/or trigonal bipyramidal geometries.7 Approximately 95% of hydrated zinc ions are found in either the tetrahedral, octahedral or trigonal bipyramidal geometries.7 However, of the three geometries, tetrahedral is the most common, followed by octahedral, which is then followed by trigonal bipyramidal. Based on this observed trend of the three geometries, one can conclude that the tetrahedral geometry is the most stable, octahedral is the second most stable, and trigonal pyramidal is the least stable. Interestingly, all the geometries are similar in energy to one another according to experimental evidence, which allows the hydrated ion to easily interconvert between these geometries.7 Despite the similarity in energy, the more stable tetrahedral geometry makes sense based on steric factors. In the tetrahedral coordination environment, the water molecules are less compacted, decreasing the amount of repulsion between them than in both the octahedral and trigonal pyramidal geometry. Decreasing the repulsion between water molecules makes the overall molecule more stable. Due to there not being a major difference between the sterics of octahedral and trigonal pyramidal geometries, one can speculate that it is possible that a driving force in Zn(II) coordination preference is electron density. In the octahedral geometry, there would be more electron density from more ligands interacting with the 2+ charge of the zinc ion than in the trigonal pyramidal geometry. Since tetrahedral is the more commonly observed geometry, sterics seem to play the primary role in geometry preference which is followed by electron density saturation. Since ZntB is thought to transport zinc as a fully hydrated ion with no indication of protein coordination, it is feasible that the zinc ion could be in either of the three commonly observed geometries. Shown below are the two most stable geometries, as calculated using density functional theory.7

These two coordination geometries for aquated Zn have different radii where Zn(H2O)6 has a radius of 2.10 A° and Zn(H2O)4 has a radius of 1.98 A°.7 Although there are slight differences in their radii, the difference is small, and it might be expected that either one of these hydration spheres could fit through the ZntB pore. Considering the flexibility of the Zn coordination environment due to zero LFSE and its lability, it is probable that the coordination geometry can adapt its geometry however might be necessary to reach the ideal specifications to travel through the pore. On the other hand, other metal ions that are less geometrically flexible (like Co2+) and/or have significantly different radii (like Ca2+) may be prevented from traveling through the pore. Proposed earlier was that the rate of water exchange during transport was a selectivity filter for Zn(II) in the ZntB system. Metal ions such as Mn and Mg, which are similar in ionic size to Zn(II) do not have the lability of Zn(II) due to their d-orbital filling and the stronger ligand effects of water. Their LFSE values are also negative and promote a more stable and inert hydrated molecule. Further, based on Mn and Mg LFSE values, there would be a preferred octahedral binding geometry.11 For the ZntB system, it seems imperative to have an ion that is stable but also labile so once transported in or out the cell, the metal ion can perform any number of functions.
Little is still known about the ZntB system. More information is needed to better understand the selectivity of Zn(II) and the mechanism of transport. However, it is known that ZntB is a vital system to sustain cell integrity in prokaryotic systems. Although much is still to be learned about the system, ZntB shows promise for potential bacterial antibiotic targets and other disease treatment due to its role in maintaining intracellular zinc concentrations. As harmful bacterial pathogens rely on intracellular zinc for survival, using ZntB as a target could help decrease their viability within hosts and maintain host cell health. However, better understanding of the zinc transport by ZntB is necessary in order to pursue further applications.
Sources
(1) Tan, K.; Sather, A.; Robertson, J. L.; Moy, S.; Roux, B.; Joachimiak, A. Structure and Electrostatic Property of Cytoplasmic Domain of ZntB Transporter., Structure and Electrostatic Property of Cytoplasmic Domain of ZntB Transporter. Protein Sci 2009, 18, 18 (10, 10), 2043, 2043–2052.
(2) Gati, C.; Stetsenko, A.; Slotboom, D.; Scheres, S.; Guskov, A. The structural basis of proton driven zinc transport by ZntB | Nature Communications, 8, 1313. DOI: 10.1038/s41467-017-01483-7
https://www.nature.com/articles/s41467-017-01483-7 (accessed Jan 22, 2018).
(3) Dudev, T.; Lim, C. Importance of Metal Hydration on the Selectivity of Mg2+ versus Ca2+ in Magnesium Ion Channels. Journal of the American Chemical Society, 2013, 135, 17200-17208.
(4) Worlock, A. J.; Smith, R. L. ZntB Is a Novel Zn2+ Transporter in Salmonella Enterica Serovar Typhimurium., ZntB Is a Novel Zn2+ Transporter in Salmonella Enterica Serovar Typhimurium. J Bacteriol 2002, 184, 184 (16, 16), 4369, 4369–4373.
(5) Dalmas, O.; Sandter, W.; Medovoy, D.; Frezza, L.; Bezanilla, F.; Perozo, E. A repulsion mechanism explains magnesium permeation and selectivity in CorA. PNAS, 111, 8, 3002-3007.
(6) Moore, J. W. Heavy Metals in Natural Waters: Applied Monitoring and Impact Assessment; Springer Series on Environmental Management; Springer-Verlag New York, 1984.
(7) Bock, C.; Katz Kaufman, A.; Gluskar, J. Hydration of Zinc Ions: A Comparison with Magnesium and Beryllium Ions. Journal of the American Chemical Society, 1995, 117, 3754-3765.
(8) Antimicrobial resistance http://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance(accessed Apr 30, 2018).
(9) Prentice, A. M.; Ghattas, H.; Cox, S. E. Host-Pathogen Interactions: Can Micronutrients Tip the Balance? J Nutr 2007, 137(5), 1334–1337.
(10) An Element of Life: Competition for Zinc in Host-Pathogen Interaction. Immunity 2013, 39(4), 623–624.
(11) Senguen, F. T.; Grabarek, Z. X-Ray Structures of Magnesium and Manganese Complexes with the N-Terminal Domain of Calmodulin: Insights into the Mechanism and Specificity of Metal Ion Binding to an EF-Hand. Biochemistry 2012, 51(31), 6182–6194.
Contributed By:
Kate McMahon, a junior chemistry major at Saint Mary's College.


