Potassium Channels
- Page ID
- 98123
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)K+ Channels
Ion channels are large membrane-spanning proteins that allow selective transport of an ion. This can occur for ions like potassium, calcium, and sodium.1 These channels are responsible for the electrical conduction in nervous systems to allow nerve and muscle cells to function properly. Neuron plasma membranes contain ion channels that have the ability to open and close based on these electric signals received from neuron activity. Electrical activity of neurons that send signals to the plasma membrane include conduction, transmission, and receiving of electric signals.2 When there is an unequal distribution of charge across a membrane, ions will spontaneously diffuse across the membrane to equalize the charge (Figure 1).1 Transport of ions across a membrane can be either active or passive depending on if energy is required. Ion channels are passive, or otherwise known as spontaneous, which allows the flow of ions to be transported across the a membrane based on when the signals are received.1-2 The flow of the ions goes from the cytoplasm to the extracellular solution, and thus depolarizing the cell. Potassium channels are one of many different types of ion channels. Main cellular functions include cell volume regulation and electrical impulse formation.3
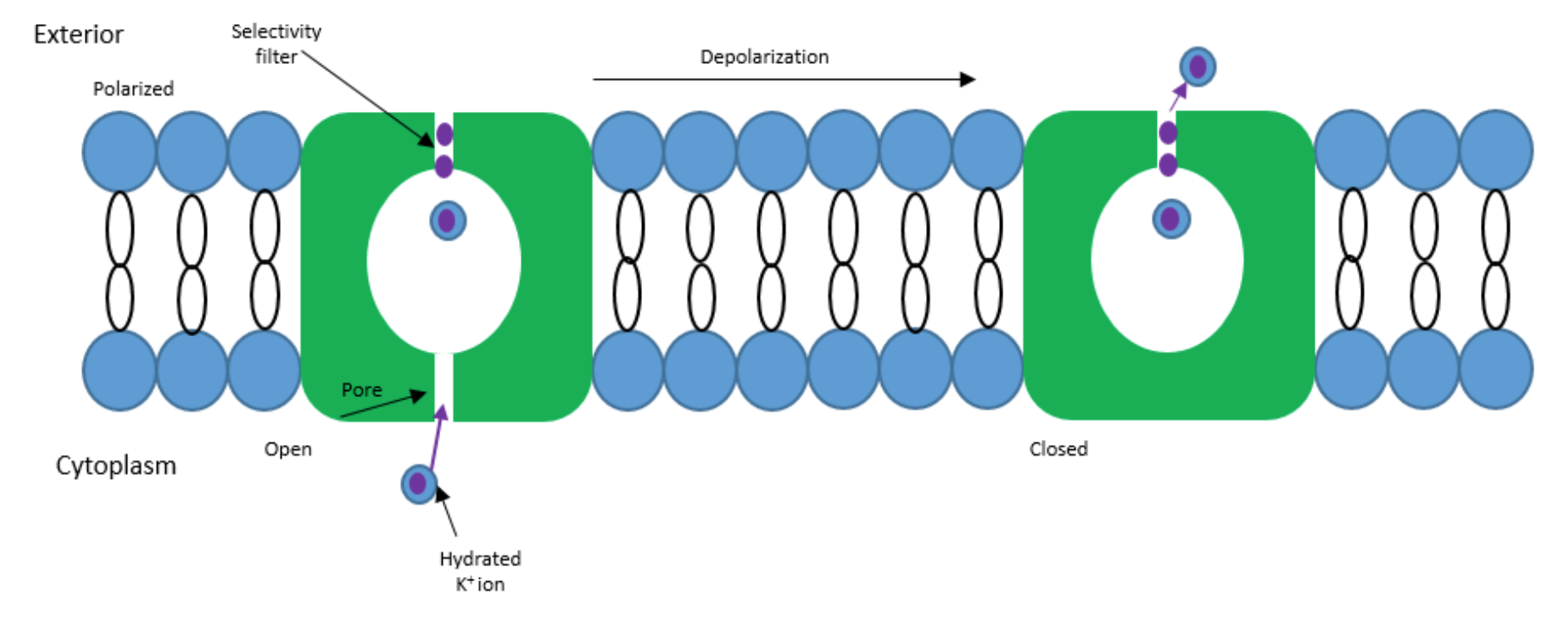
There are many types of ion channels, but this paper will be focusing on potassium channels.The human genome contains over 90 genes that encode for potassium channels.3 Dysfunction of potassium channels have been shown to link an individual to diseases such as the hyper-and hypokalemic periodic paralysis, myotonia, and epilepsy.4 Potassium channels specifically allow potassium ions to penetrate the membrane when there is a response to a change in the membrane potential. This “specificity” means that no other ions can pass through this channel including sodium, which is very similar to potassium in regards to charge and size.3 This selection is completed at rates that approach the diffusion limit. There are four main types of potassium channels which are as followed: calcium activated, inwardly rectifying, tandem pore domain, and voltage-gated. The differences between these types are mainly with how the gate receives its signal, whereas the structure of these channels is similar. A general structure for potassium channels can be seen below in Figure 2.3
Potassium
Potassium is an alkali metal with a valence electron configuration of ns1 and contributes only one element to the molecular orbital band. There are some common trends that can be found from the periodic table, and there are specific ones that are for alkali metals. Alkali metals contain low first ionization energies and will want to form M+ metals increasingly down the group. Potassium’s thermodynamic tendency to become K+ is due to the large and negative standard potentials that occur. Oxidation becomes more favorable going down the group because the enthalpies of sublimation and ionization decrease. Group 1 element ions are hard Lewis acids and mainly form complexes with hard donors like O and N.1
KcsA Channel: Structure
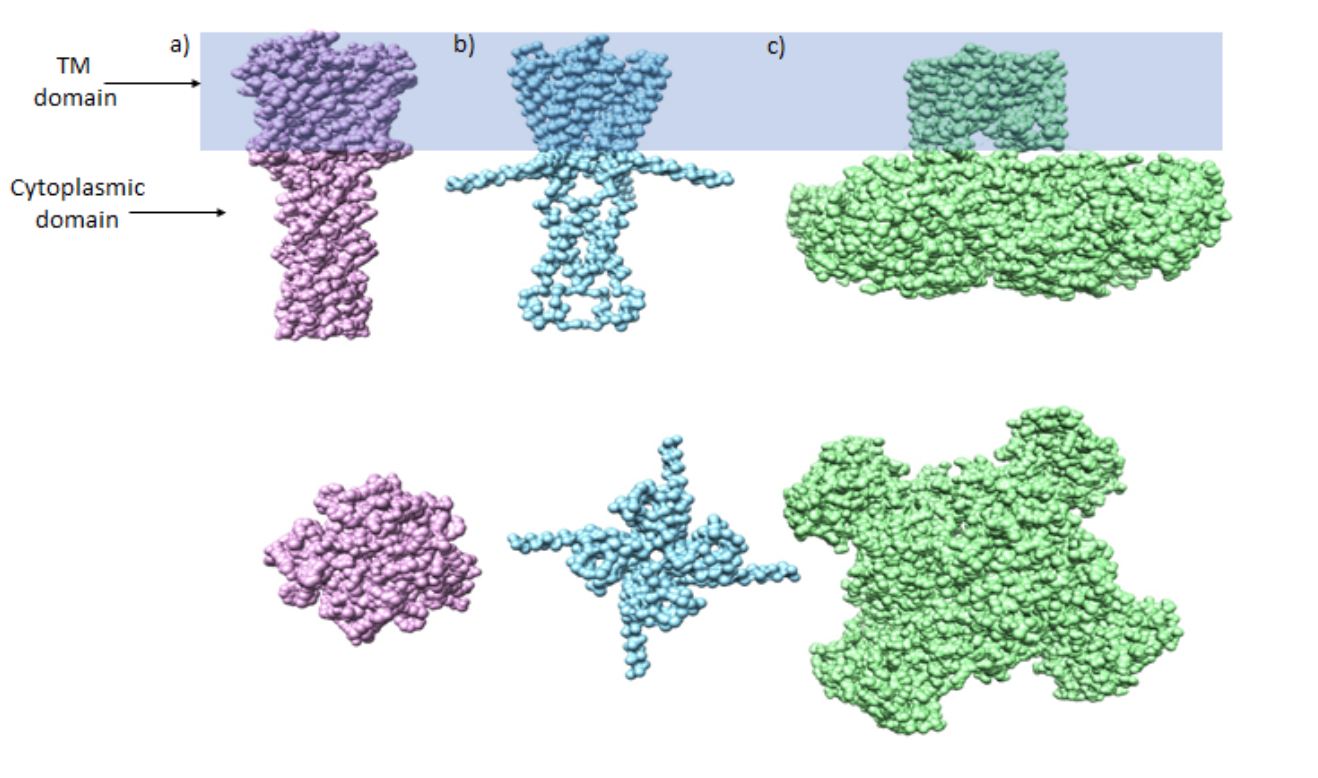
This paper will be focusing on KcsA, a potassium channel from Streptomyces lividans, which contains two domains. The first being the transmembrane portion, which contains 120 residues. This will be discussed in detail below. The second domain is a cytoplasmic domain that contains about 40 residues. The cytoplasmic domain contributes to the stability of the closed state. This is concluded based on data that shows a decrease in efficiency of channel folding and assembly and thermal stability being affect when the cytoplasmic domain is removed from the crystal structure.5 As seen in Figure 2, the water filled C-terminal cytoplasmic domain is 4-helix bundle structure that extends linearly towards the cytoplasm.5-6 It has been shown by electron paramagnetic resonance, EPR, data produced by Uysal, that the C-terminal has a stretch that bulges out where the C-terminal domain meets the transmembrane domain.5 This stretch of helix that connects the two has a higher degree of flexibility than the other parts that potentially plays a role during the opening of the channel.5 There is still much research that needs to be conducted to fully understand the functionality of the C-terminal domain. A suggested function of the domain is that it may serve as a receptor for an unidentified plasmic activator protein or ligand, but others argue that there is no significant effect on the permeation path for the ion. The structure does not yet contain orientation of any side chains, but the arrangement of the helices suggest electrostatic interactions in the subunit.6 Figure 3, demonstrates how the cytoplasmic domain compares between an open channel vs. a closed channel with two bacteria channels.7 The rest of the paper will be focusing on the transmembrane domain that selects for potassium.
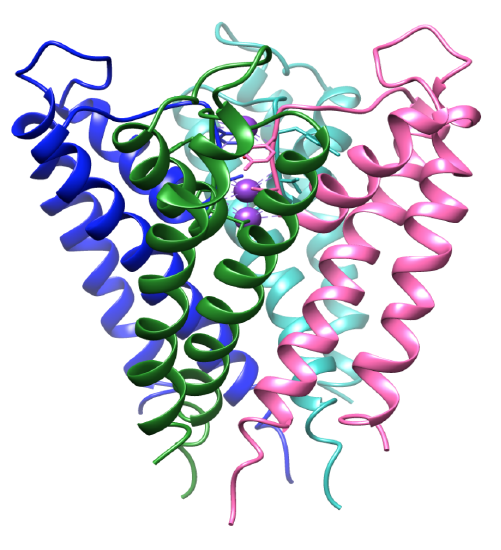
The selectivity for potassium ions relies heavily on the structure of the potassium channel selectivity filter. The coordination environment of the channel has selective ligands that bind specifically to potassium ions.8 The potassium channel protein is a tetramer, that contains four identical subunits that form a central pore.8-9 The ions flow into a pore that opens and closes, directing the ions to the central cavity. The subunits contain two alpha helices where one faces the central pore, inner helix, and one faces the lipid membrane, otherwise known as the outer helix.3 Along with the inner and outer helices, each subunit includes a pore helix.3,8 The subunits are tilted, as seen in Figure 3, where the selectivity filter is near the extracellular surface of the membrane. The four inner helices are structured in a way where they come together near the intracellular surface. Both the intra- and extracellular entryways are negatively charged by acidic amino acids. This allows for a higher concentration of cations near the membrane and less anions because of the opposing charges.8

The pore region was first identified with pore-blocking scorpion toxins. They interact with amino acids at the entrance of the pore causing blocking and dysfunction. The pore is where the potassium ion enters, where then it is transferred to the central cavity, as seen in Figure 4. During this time the potassium ion stays hydrated.8 The intracellular pore is the feature of the channel that allows potassium ions to enter the cell when it is open, or be blocked from entering when closed. The channels opens when there is a change in membrane potential, allowing a influx of potassium ions into the cavity filter. Once selective binding occurs in the selectivity filter, the potential drops across the membrane, thus closing the pore.1 The pore and the cavity are hydrophobically lined, allowing the ability to lower electrostatic barrier. There needs to be a lower electrostatic barrier so the energetic cost of dehydration is compensated due to the large energy barrier for ion permeation (dielectric barrier). Thus, the water filled cavity allows for an energetically favourable environment.3,8,10,11
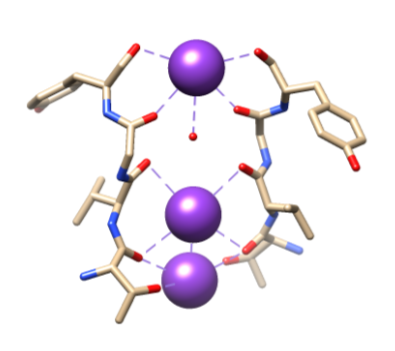
Next, the ions need to be selected into the selectivity filter (SF), which is lined with polar main chain atoms that belong to amino acids. The SF is so narrow that the ions must first be dehydrated before entering it. There are two essential features in the SF. First, the main chain atoms create suitable dimensions for coordinating a dehydrated potassium ion by stacking, thus making sequential oxygen rings. This creates a very small distance that the potassium ion must travel to reach each site while traveling through the filter. This forms a sequence of four cubic eightfold coordination sites, as seen in Figure 6.3,8 At any time, these sites are occupied by two K+ ions and two water molecules in an alternating sequence The second feature is the protein packing around the SF, which follows a sequence of T-V-G-Y-G, shown in Figure 5.9 Conduction can then occur when two potassium ions enter the SF, causing a balance between attractive force that occurs from the SF to the ion with the repulsive force that occurs between the two potassium ions.8 This repulsion helps potassium ions overcome its intrinsic affinity for binding tightly to the binding sites.3 Further discussion of the selection mechanism will be discussed below.
K+ Selectivity
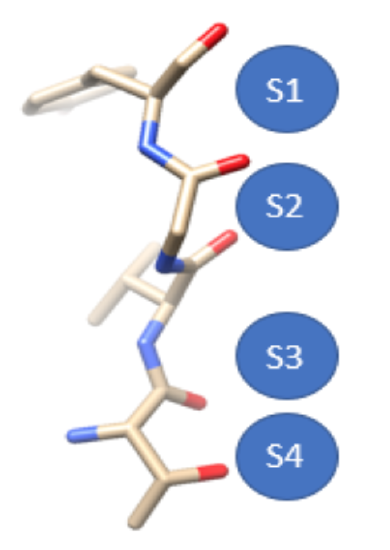
The SF consists of four sites (S1-S4) that are formed by oxygen molecules, as seen in Figure 7.11 The selectivity of potassium channels is of functional importance due to the fact that if sodium ions were to be selected, membrane repolarization would not occur. Even though sodium ions are smaller than potassium, ~.95 Å and ~1.33 Å respectively, they are not able to penetrate the membrane.12-13 This selection of potassium ions occurs by a factor of more than 1000 compared to sodium ions. Much research has been conducted to elucidate the mechanistic nature of how exactly potassium channels select for potassium ions, in which conclusions can vary by the type of potassium channel. This description will be based on KcsA..9, 12, 13
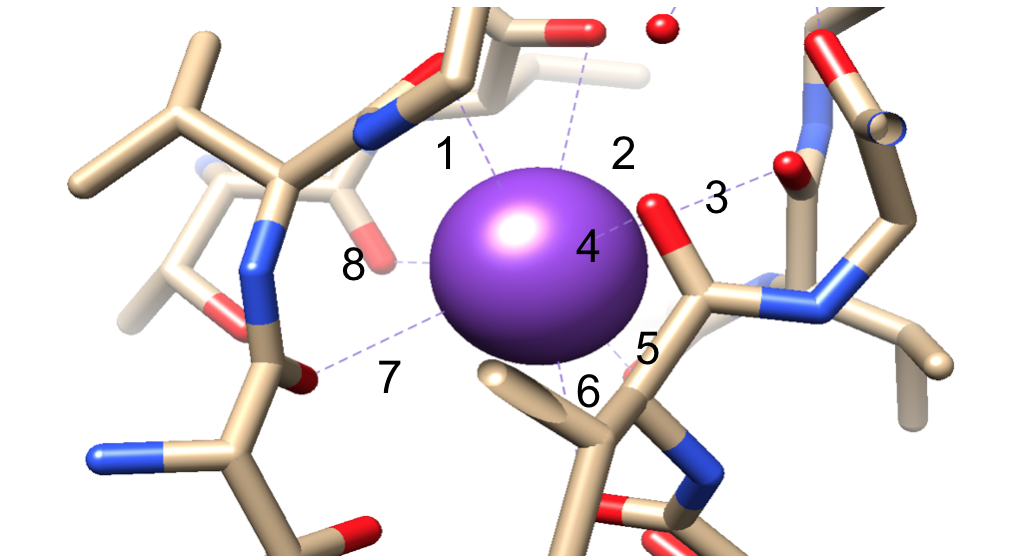
Selectivity of potassium ions has been attributed to the sequence motif TVGYG in positions 75-78 of each monomer. As stated above, potassium channels bind to ligands through an eight coordinate, octahedral coordination environment.11 These are optimally placed to allow for the interaction with potassium ions, making the transfer of energy from the central cavity to the SF extremely low because the environment is not drastically changing.8,11 When a potassium ion binds to the SF, eight water molecules are released from a hydrated potassium ion. The ion is then bound to eight carbonyl oxygens at each of the four sites in the SF. At the last site, the ion binds to four of the carbonyl oxygens as well as four water molecules, keeping the eightfold coordination number. Once the ion has completely diffused through the channel, the ion becomes rehydrated once again in the extracellular matrix.3,8 This mechanism can be viewed in Figure 8.

Figure 8. Mechanism of the diffusion of potassium ions through the channel, emphasizing when the potassium channel is hydrated. The ion first goes through the pore where it resides in the central cavity. At this point the ion is still hydrated. When the ion is selected, it then moves to through the selectivity filter, where it becomes dehydrated. Following, the ion enters the extracellular matrix hydrated where it is hydrated once more.
This simple reaction is favorable because of low energy cost to lose the water molecules and bind the to the selectivity filter. Potassium ions are normally surrounded by eight water molecules, which are stripped before entering the filter. The filter is designed to have similar dimensions that mimic the shell of water that encloses potassium by lining the pore with eight oxygen atoms, as stated above. These are optimally placed to allow for the interaction with potassium ions. This makes the transfer of energy from the central cavity to the selectivity filter extremely low because the environment is not drastically changing.This static environment completely compensates for the energetic cost of K+ ion dehydration when it enters the cavity.9 The selection of the right metal allows for the ability for potassium channels to function correctly.
Atomic Radii: Na+ vs. K+
Sodium is a metal that is very similar in charge and size, as stated above, but sodium ions are not selected. This is due to the size of the ions.3,8,11 Sodium ions are smaller in ionic radius. As seen in Figure 9, sodium ions have a smaller ion radius resulting in the inability to coordinate with the all eight oxygens at the first site. This demonstrates why sodium ions are not selected to go through the channel, even though the ions are smaller than potassium ions. This results in sodium ions favoring the shell of water.9,12,13
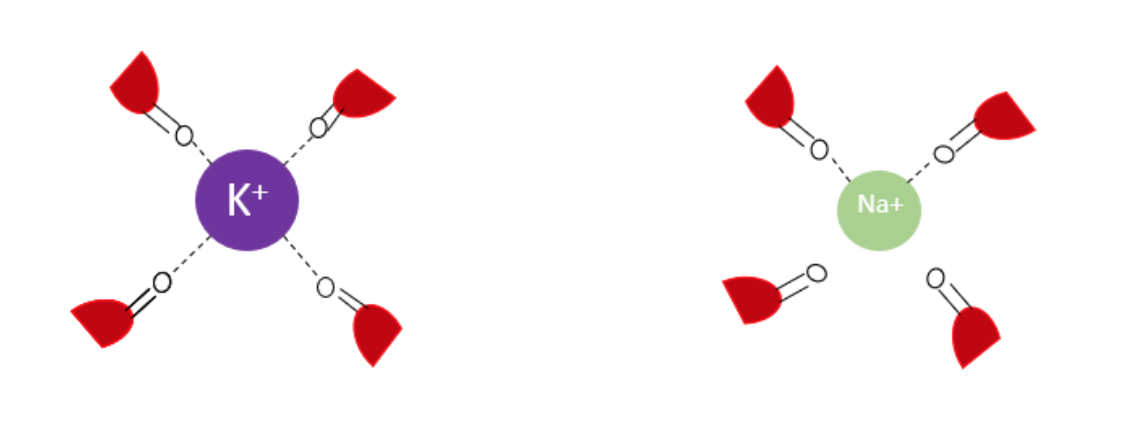
Figure 9. Comparison of potassium and sodium ion binding to a site in the selectivity filter. The sodium ion has too small of an ionic radius to be fully coordinated to all the carboxylic groups in the filter. It should be noted that the figure for easier visualization only shows only four of the eight potential carbonyl oxygens available at each site. The red represents the amino acid.
Additional Bio-inorganic Properties to Explain K+ Binding/Release
HSAB Theory
The potassium ion binds to carbonyl oxygens which are hard bases.14 Hard Soft Acid Base theory characterizes metals as either “hard” or “soft”. This is dependent on the charge density, polarizability, and the type of bonding interactions. For example, hard metals tend to have higher charge density and/or form more electrostatic interactions, while soft metals tend to have lower charge density and/or more covalent bonding. Since potassium is characterized as a hard metal due to its electrostatic interactions, it would most likely bind to ligands that are hard. This theory helps explain why potassium binds to Thr 75, Val 76, Gly 77, and Tyr 78, which are mainly hard ligands.11,14
Lability
Lability refers to the ability for a metal-ligand bond to break. When a compound has bonds that are easily broken between the metal-ligand, then that metal is referred to as labile. A common trend that occurs is that alkali metals tend to be more labile compared to other metals that have higher charges. When metals have higher charges, they tend to be stronger Lewis acids. This would allow the metal to bind the ligands more tightly, resulting in harder bonds to break. Another trend that has been observed is that metals that have no d electrons tend to be more labile because of there are no electrons in the way for a new ligand to donate its electrons. Potassium channels are extremely labile because it has a +1 charge, as well as a 0 ligand field stabilization energy (contains zero d electrons). Potassium being labile is extremely important since its diffusion rate is so high that the ion needs to be able to go from one site to the next. If the metal was not labile, the channel would not be able to flow potassium ions as quickly as it can because the metal-ligand complexes would not be as easily breakable.
Conclusions
Potassium channels are necessary for cellular function, especially assisting in neuronal signaling. There are many features that make up the channel, but the key feature is the selectivity filter. The SF is what keeps the channel functioning by specifically selecting potassium ions. Overall, this selectivity for potassium over other ions can be attributed to the size of the ion and HSAB theory. The SF is the the correct fit to allow an eightfold coordination environment with potassium ions. The specific ligands assist with the binding of the potassium channels because the HSAB theory suggest that like binds to like and both the ligands and metals are characterized as hard. To conclude, there are many features of potassium channels, and this paper only began to touch on the functionality of the channels.
Sources
- Shriver, D F, P W. Atkins, and Cooper H. Langford. Inorganic Chemistry. Oxford: Oxford University Press, 1994. Print.
- Lodish, Harvey, et al. The Action Potential and Conduction of Electric Impulses. 2000. www.ncbi.nlm.nih.gov, https://www.ncbi.nlm.nih.gov/books/NBK21668/.
- MacKinnon, R. Potassium Channels. FEBS Letters 2003, 555 (1), 62–65.
- Dworakowska, B.; Dołowy, K. Ion Channels-Related Diseases. Acta Biochim. Pol. 2000, 47 (3), 685–703.
- Uysal, S.; Vásquez, V.; Tereshko, V.; Esaki, K.; Fellouse, F. A.; Sidhu, S. S.; Koide, S.; Perozo, E.; Kossiakoff, A. Crystal Structure of Full-Length KcsA in Its Closed Conformation. Proc Natl Acad Sci U S A 2009, 106 (16), 6644–6649.
- Cortes, D. M.; Cuello, L. G.; Perozo, E. Molecular Architecture of Full-Length KcsA. J Gen Physiol 2001, 117 (2), 165–180.
- Goodsell, D. S. Potassium Channels. RCSB Protein Data Bank 2003.
- Doyle, D. A.; Morais Cabral, J.; Pfuetzner, R. A.; Kuo, A.; Gulbis, J. M.; Cohen, S. L.; Chait, B. T.; MacKinnon, R. The Structure of the Potassium Channel: Molecular Basis of K+ Conduction and Selectivity. Science 1998, 280 (5360), 69–77.
- Kuang, Q.; Purhonen, P.; Hebert, H. Structure of Potassium Channels. Cell Mol Life Sci 2015, 72, 3677–3693.
- Roux, B. The Cavity and Pore Helices in the KcsA K+ Channel: Electrostatic Stabilization of Monovalent Cations. Science 1999, 285 (5424), 100–102.
- Sansom, M. S. P.; Shrivastava, I. H.; Bright, J. N.; Tate, J.; Capener, C. E.; Biggin, P. C. Potassium Channels: Structures, Models, Simulations. Biochimica et Biophysica Acta (BBA) - Biomembranes 2002, 1565 (2), 294–307.
- Nimigean, C. M.; Allen, T. W. Origins of Ion Selectivity in Potassium Channels from the Perspective of Channel Block. The Journal of General Physiology 2011, 137 (5), 405–413.
- Thompson, A. N.; Kim, I.; Panosian, T. D.; Iverson, T. M.; Allen, T. W.; Nimigean, C. M. Mechanism of Potassium Channel Selectivity Revealed by Na+ and Li+ Binding Sites within the KcsA Pore. Nat Struct Mol Biol 2009, 16 (12), 1317–1324.
- Noskov, S. Y.; Berneche, S.; Roux, B. Control of Ion Selectivity in Potassium Channels by Electrostatic and Dynamic Properties of Carbonyl Ligands. Nature 2004, 431 (7010), 830.
Contributed By:
This work was originally written by Courtney Weston, Spring 2018; Courtney recently (as of May 2018) received her B.S. in Chemistry from Saint Mary's College (Notre Dame, IN) and will finish her second degree in Environmental Engineering next Spring.
This work was originally edited by Dr. Kathryn Haas (Assistant Professor), Madison Sendzik (Teaching and Research Assistant).


