Cytochrome Oxidase
- Page ID
- 97882
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)The electron transport system (ETS) is an energy production system for aerobic organisms found in the mitochondria of cells. It is the third and final step in cellular respiration, which is the process by which the cell produces energy. When an organism takes in carbohydrate nutrients, they are converted to glucose which can be further broken down through a set of chemical reactions to pyruvate through glycolysis. After glycolysis, this pyruvate molecule is oxidized in the Kreb’s cycle which in turn produces ATP and high energy molecules, NADH, and FADH2. Only a few ATP are produced directly from the Kreb’s cycle; however the NADH and FADH2 are transported to the ETS where they can be oxidized to produce more ATP (Figure 1).1
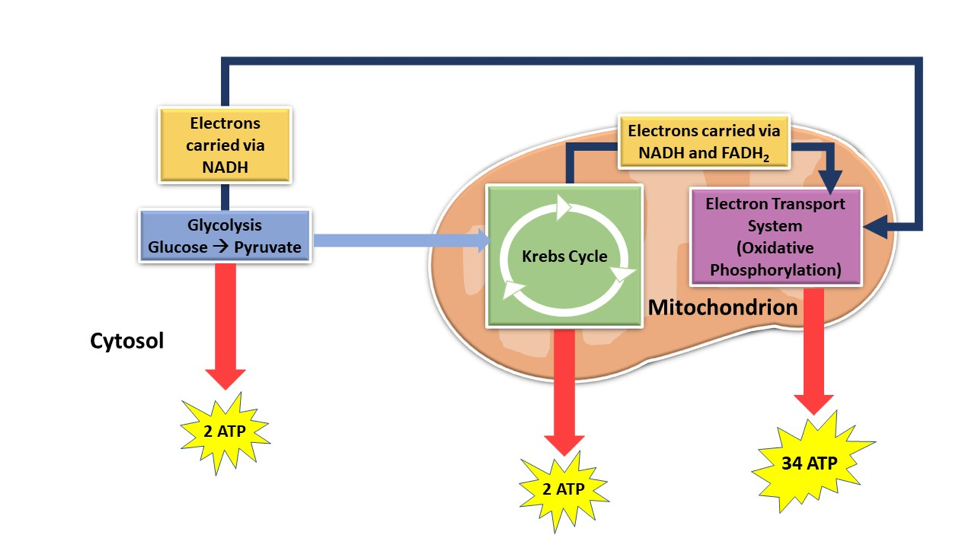
After the NADH and FADH2 are oxidized to NAD+ and FAD, the electrons that are removed from NADH and FADH2 and are transported through the four electron transport complexes of the ETS. The movement of electrons provides energy to pump hydrogen ions (H+) across the inner mitochondrial membrane into the intermembrane space through four complexes.1 The four complexes are NADH-CoQ (ubiquinone reductase), succinate dehydrogenase, cytochrome bc1, and cytochrome oxidase (Figure 2).2

The pumping of hydrogen ions through the four complexes stores potential energy by creating a concentration gradient, with higher hydrogen ion concentration in the intermembrane space. This energy is captured as hydrogen ions cross back over to the matrix of the mitochondria, via the ATP synthase, producing energy for the cell in the form of ATP.1 The hydrogen ion concentration will then increase in the mitochondrial matrix and travel through the three complexes again when another NADH and FADH2 are introduced to the ETS. This process will continue to cycle and produce high numbers of ATP for the cell to use as energy (Figure 2).1
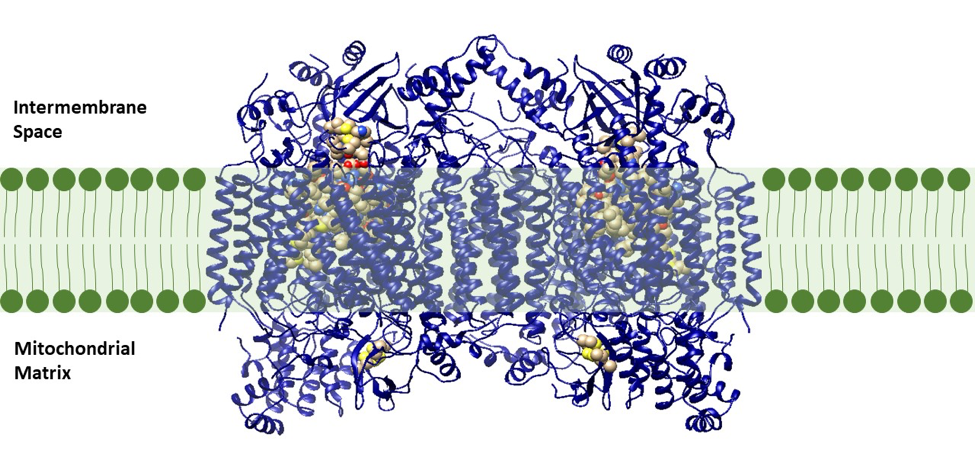
Cytochrome oxidase, also known as complex IV, is the terminal, or final, enzyme of the electron transport system (this does not include ATP synthase). Cytochrome oxidase is a transmembrane molecule found in the mitochondria of eukaryotes and in the cellular space of aerobic prokaryotes. This molecule is a proton pump that plays a vital role in producing energy, in the form of ATP, via the ETS (Figure 3). In the last steps of the energy production process, cytochrome oxidase oxidizes the waste products from the end of the energy making process, converting reactive species, H+ and dioxygen (O2), to a more stable molecule, water (H2O). In the absence of cytochrome oxidase, this energetically favorable reaction could be explosive and no energy production would occur.2 Cytochrome oxidase allows this reaction to occur safely while harnessing energy to form the H+ ion gradient. This energy produced from the reaction is used to pump H+ ions from the matrix to the intermembrane space of the mitochondria (Figure 2).1
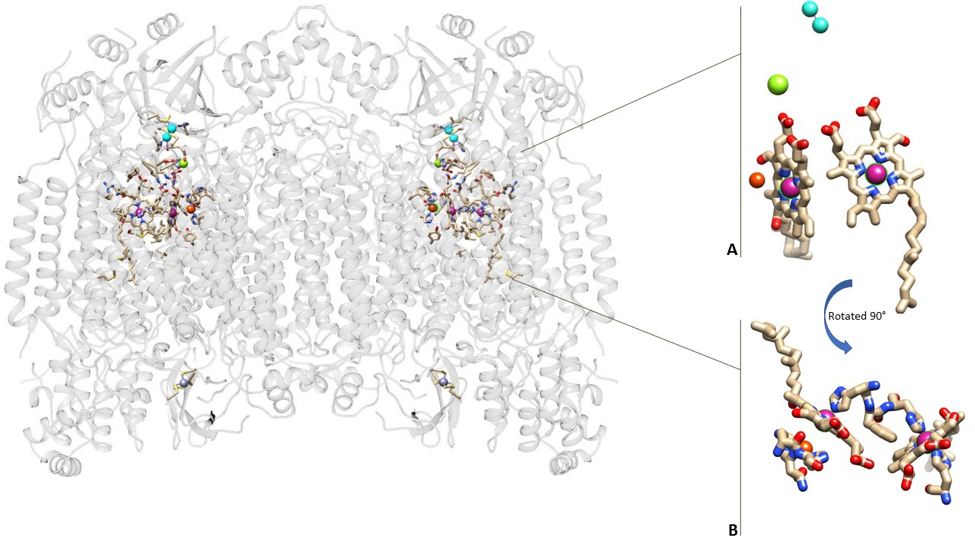
Cytochrome oxidase is a dimer meaning it is made up of two identical proteins. The two proteins in cytochrome oxidase mirror one another. The system uses multiple metals to complete its function including the two irons in the two hemes, three coppers, one magnesium, and one zinc (Figure 4). The irons in the two hemes and the three copper molecules are vital to the success of the enzyme in aerobic respiration.1

2 CuA molecules function in allowing the electrons to enter the enzyme from cytochrome c.2,3 The electrons travel from the CuA molecules to the first heme, also known as hemea. hemea functions in allowing the electrons to be transferred from CuA to the second heme, hemea3. The iron in hemea3 functions in binding oxygen, with the help of CuB. The energy production system selects a heme iron group to bind the oxygen along with the other copper atom and is the location of oxygen reduction (Figure 5).3 The function of magnesium and zinc remains unknown.4
4e- + 8H+ + O2 → 4e- + 4H+ + 2H2O.9 (1)
The overall reduction that occurs in cytochrome oxidase is shown above (Reaction 1). The electron transfers from the electron donor, cytochrome c, to CuA to hemea to hemea3 and finally to the electron acceptor, oxygen, are carried out mostly by redox reactions (Figure 5).3
The electrons travel from cytochrome c to the two CuA molecules in the intermembrane space at a fast rate of 70,000 s-1 and with an Eo’ value of 0.27 V.3,5,6 The Eo’, or standard reduction potential, is a value indicating the likelihood a chemical species will be reduced, with a more positive value meaning it is more likely to be reduced. Literature has shown that CuA has a mixed oxidation state of I and II (or 1.5 each). Because of this mixed oxidation state when the electrons enter the cytochrome oxidase the charge is shared between the two CuA. Only a single electron is transferred from the cytochrome c to the CuA.6 In the cell the environment is different than laboratory conditions, therefore the Eo’ value would be about +0.05 V.3
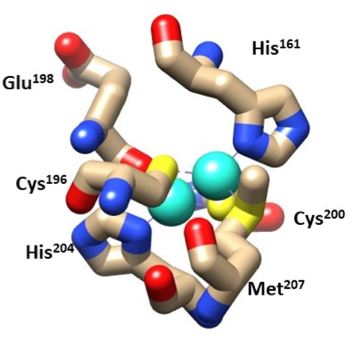
The two CuA molecules are what bring the electrons from the cytochrome c to hemea. They both have tetradentate ligands with a tetrahedron shape.7 CuA1 is bound by His204, Cys200, Cys196, and Glutamic acid198. CuA2 is bound by His161, Cys200, Cys196, and Met207. Both Coppers are bound by Cys196 and Cys200, this chelating effect helps stabilize the molecule (Figure 6). The Hard Soft Acid Base (HSAB) theory helps explain the stability of the hemes. HSAB theory characterizes metals as hard, borderline, or soft acids and bases based upon their charge density. HSAB shows that chemical species like to bind with similar species, for example hard acids like to pair with hard bases and borderline acids like to pair with borderline bases. CuA applies to the HSAB theory as the copper binds to the imidazole residue on histidine which is borderline for HSAB. The two cysteine and glutamic acid residues that bind with Cu (I) or (II) are soft and hard bases, respectively, for HSAB and may still bind as they are next to borderline. Therefore HSAB may still be a selective method for the Cys and Glu ligands.3
As Cu (II) is a d10 metal its d orbital is full of electrons. As the d orbital is full the LFSE (ligand field stabilization energy) would be zero. The LFSE is a number that defines the energy of the electrons compared to the average energy of the orbitals. A more negative LFSE indicates lower energy and a more stable complex. As Cu(II) has an LFSE of zero this indicates that the geometry of the metal and its ligands it likely dependent upon sterics. Therefore the tetrahedral geometries of the CuA1 and CuA2 is a result of reducing steric hindrance between the ligands. CuA1 and CuA2 would have an electron count of 17-18. Both coppers each have four ligands with no charge thus having 8 ligand electrons. Copper can have an oxidation state of (I or II), and therefore will have 9-10 d electrons bringing the electron count to 17-18.3
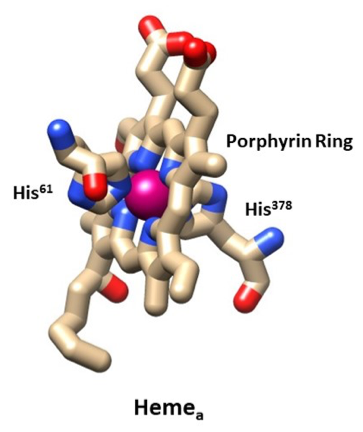
After the electrons have entered the cytochrome oxidase from the cytochrome c to the CuA molecules they will then transfer to hemea (Figure 7). This transfer is made possible by 14 covalent bonds and 2 hydrogen bonds spanning 19.5 Å and taking about 20,000 s-1.3 The iron transfers the electrons one at a time and changes from Fe (III) to Fe (II). However, literature shows that the iron only changes its charge about 10-15% as the change in charge is spread over the porphyrin ring decreasing the amount of energy needed.6
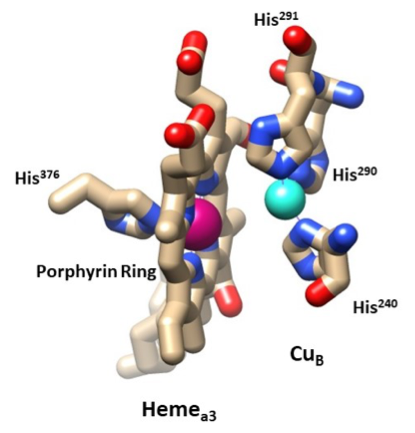
Next, the electrons transfer from hemea to hemea3 (Figure 8). The hemes are perpendicular to one another and the edges are located 7 Å apart.8 How the transfer of electrons from hemea to hemea3 occurs is still not fully understood. Literature shows that electron tunneling stops at hemea, and from hemea to hemea3 the electrons are transferred from the edges of the heme and propelled by protons pumping to the intermembrane space.6
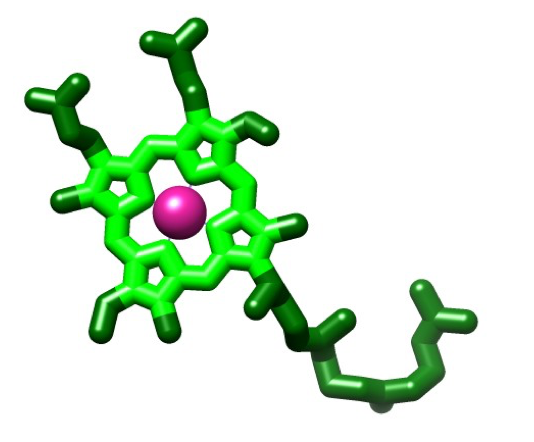
The structure of the heme molecule is a porphyrin ring that is tetradentate chelating of an iron molecule with four nitrogens. Iron can form up to six bonds due to steric constraints that cost the molecule energy by not having enough space for the electrons which repel one another. A porphyrin ring is the building block for heme and is a planar structure composed of four pyrrole rings (Figure 9). Four of the bonds are with the nitrogens of the porphyrin ring, as previously stated, while the fifth coordination site is occupied by an imidazole in histidine, and the sixth coordination site is occupied by another imidazole for hemea, or O2 for hemea3 (Figure 10). Both irons have a planar shape with the heme molecules, but with the addition of their ligands (two imidazole residues for hemea or an imidazole and an oxygen for hemea3) they becomes an octahedral shape.3 The geometry of the hemes can be attributed to LFSE as the the LFSE equals -0.4 for Fe(II) and -2.0 for Fe(III). This indicates that the geometry of the molecule may be a result of the high stability due to the lower energy of the electrons and their orbitals.8
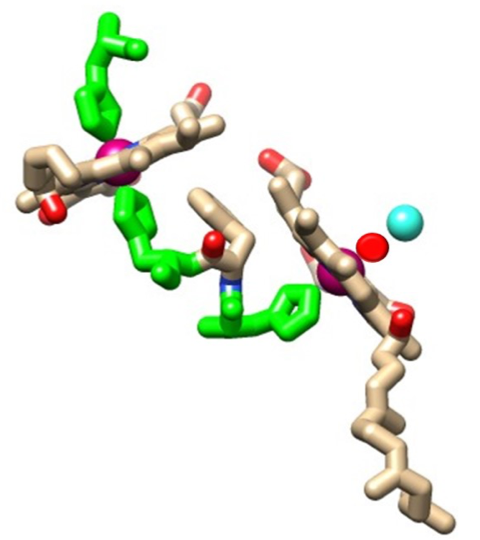
Thermodynamic theories are important for ML binding as they provide ideas about the energy needed for breaking bonds and the stability of a molecule. The two hemes in cytochrome oxidase both maintain their stability largely due to their macrocyclic porphyrin ring. This porphyrin ring is made up of 4 pyrroles that bind their nitrogens to the iron. In cytochrome oxidase the iron never needs to leave the hemes and the stability of the porphyrin ring may help the iron stay in its place to properly transfer electrons to oxygen (Figure 10).9
The iron does not leave the porphyrin ring for either hemes and the ligands of hemea remain bound as well. The heme molecules are very stable due to the chelating effect of the porphyrin ring and is relatively inert meaning it holds its metal ion and maintains its planar geometry. The iron in hemea3 has an imidazole ligand that remains bound to the iron, but also has a position for oxygen. The oxygen iron bond is broken when electrons are transferred to the oxygen and bound to two H+ ions.3 The high lability, or ability of the metal-ligand bond (Fe (II) and O2) to be broken, is necessary for the water to be released from hemea3 and for the cytochrome oxidase to reset for the next set of electrons. The high lability of the Fe (II) is likely a result of the low LFSE value of -0.4. The rate constant is unknown for the iron bound to oxygen. The HSAB theory applies to the iron and the porphyrin ring as Fe (II) and N (in the pyrrole ring) are both classified as having borderline acid character. Like prefers like, indicating that Fe (II) may be using this to select the right ligand to help it bind oxygen, which is hard base for the HSAB theory. The nitrogen in the pyrrole ring is likely a borderline base as it is in a ring structure like pyrimidine and imidazole which are borderline nitrogen containing ring structures as well.8
The iron in hemea transitions between two oxidation states, Fe (III) and Fe (II). The iron hemea3 transitions from Fe (II, III, and IV).10 The two irons in the heme a and a3 both have an electron count of 16-18, as the oxidation state changes. Both irons are bound by four nitrogens from a porphyrin ring, an imidazole nitrogen from His and an O2 molecule. All of the nitrogen donor atoms are neutral. With six ligands bound to the Fe there would be a total of 12 ligand electrons (6e- x 2=12e-). Knowing that the iron may be missing 2-4 electrons, based on its oxidation state, there remains 4-6 d-electrons giving a total electron count of 16-18 (12 ligand e- + 4-6 d e- = 16-18 e-) (Figure 11). Literature has shown that the iron in hemea is low spin as it is a 3d metal with a 3+ charge and because it has imidazole ligands that are pi acceptors which are strong field. While the other iron metal center in hemea3 is high spin as it is a 3d metal with a 2+ charge.10,11
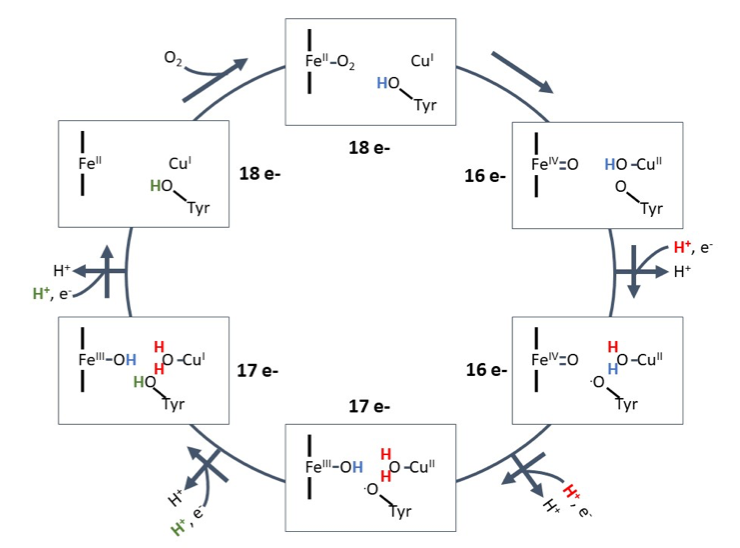
Lastly, the electrons transfer from hemea3 to O2. This transfer is made possible by the iron on the hemea3, CuB, and a tyrosine. These set of molecules are also known as the bionuclear center (BNC).3 Once the electron reaches the hemea3, Fe(III) becomes Fe (II) and the oxygen binds to the Fe (II). The H+ ion from the tyrosine ligand of CuB transfers to the oxygen bound to CuB (I) changing it to CuB (II). During this state the Fe (II) double bonds to the oxygen and becomes Fe (IV). The first H+ and e- enter the BNC and the H+ binds to the oxygen on the CuB (II). During this step, while the electron places on the tyrosine oxygen the tyrosine oxygen moves to the CuB(II) oxygen. Next, the second H+ and e- set enter with the new H+ replacing an H+ on the CuB(II) site and the e- placing on the Fe (IV) changing it to Fe (III). The replaced H+ moves to the Fe (IV). The third set of H+ and e- move in with the third H+ going on the tyrosine oxygen and the electron going on the CuB (II) causing it to become CuB (I). The final H+ and e- enter the BNC and place on the Fe (III) causing it to become Fe(II). This stage is also where the two water molecules bound to Fe (III) and CuB(I) fall off. The cycle then restarts when oxygen binds to the Fe (II) (Figure 11).12
CuB is vital to the reaction of cytochrome oxidase and has tridentate ligands. All three ligands are imidazole side chains (His240, His290, and His291). With the three imidazole ligands, CuB forms a trigonal planar shape and is stabilized by the chelating His290 and His291 ligands (Figure 8).3 Similar to the CuA ions, CuB has an LFSE of 0 and has a trigonal planar geometry due to steric hindrance of the three His ligands. Copper can have an oxidation state of (I) or (II).10 CuB would have an electron count of 14-16. This copper has three imidazole ligands with no charge therefore it has 6 ligand electrons. The HSAB theory also applies to the Cu (II) (CuB) center and the 3 imidazole ligands as both are considered borderline acids and bases, respectively, for HSAB and like attracts like, indicating that Cu (II) may be using this to select the right ligands. Cu (I) also occurs and may still be using HSAB to select the imidazole groups as it is close to borderline. Copper can have an oxidation state of +I, or +II, and therefore will have 9-10 d electrons bringing the electron count to 15-16. CuB takes on an oxygen and releases it after two H+ ions are bound to it producing one molecule of water (Figure 11). The rate constant of the O-CuB bond is 3.5 x 108 M-1s-1 showing that the breaking of the bond is quick as and that it is a weak bond. As previously mentioned, the CuB has an LFSE of 0 indicating it is highly labile which correlates with the CuB’s ability to react quickly with the oxygen.3
The reduction potential of the cytochrome oxidase reaction (Reaction 1) is +0.82 V. Beginning with the initial electron donor, cytochrome c, and ending with the final electron acceptor, O2, we see that the reduction potential went from +0.27 V to +0.82 V. This increase in reduction potential from the electron donor to the electron acceptor makes the overall reactions of cytochrome oxidase spontaneous.6
Cytochrome oxidase is a colored protein. CuA has absorption bands at about 480 nm, 530 nm, and even one in the infrared spectrum at about 830 nm.12 Literature has shown that although these absorbances reflect a blue/purple color in the lab, in an intact cytochrome oxidase the hemes hide the CuA absorbances with their own absorbances. CuA absorptions are charge transfer absorptions as ε>1000 cm-1 (ε=2000 cm-1).12
Hemea has been found to have absorbances at 446 nm and 605 nm. hemea3 has absorbances at 441 nm, 611 nm, and 656 nm.12 The absorbances for both hemes mean the reflected light gives the hemes a reddish/purple color. The absorbances and colors of both hemes have been to found to be likely a result of charge transfers; however no ε value was found in the literature. CuB is not colored as it does not have any absorbances in the visual spectrum.12
Cytochrome oxidase is a key molecule in aerobic respiration and without a properly functioning enzyme can lead to cell death. The primary disorder associated with cytochrome oxidase is cytochrome oxidase deficiency which is a relatively rare, autosomal recessive disorder.13 Leigh’s Syndrome is a human disease caused by cytochrome oxidase deficiency and its symptoms include poor muscle tone and muscle movement, mental impairment, and lack of growth. These symptoms begin to manifest in infancy and the disorder usually results in death within two to three years of life. Although there is no cure for Leigh’s Syndrome, treatment with Vitamin B1 has been shown to help prolong the patient’s life. Therefore, cytochrome oxidase is very important to humans and all other aerobic organisms as it is the primary reason we breathe oxygen and is what helps our bodies produce energy to keep us alive.14
References
- Alberts B, Johnson A, Lewis J, et al. 2002. Molecular Biology of the Cell., 4th edition. New York: Garland Science.
- Goodsell, D. 2000. Cytochrome c Oxidase. Protein Database (PDB). (accessed Jan. 23, 2018).
- Michel, H., Behr, J., Harrenga, A. Kannt, A. 1998. Cytochrome c oxidase: structure and spectroscopy. Annu. Rev. Biophys. Biomol. Struct., 27.
- Florens L. et al. 1999. The Role of Magnesium and Its Associated Water Channel in Activity and Regulation of Cytochrome c Oxidase. The Phototrophic Prokaryotes. Springer, Boston, MA.
- Ludwig, B., Bender, E., Arnold,S., Hüttemann, M., Lee, I., and Kadenbach, B. 1999. Cytochrome c Oxidase and the Regulation of Oxidative Phosphorylation. Biochemistry, 38 (46): 15129–15140.
- Kaila, V. R. I., Verkhovsky, M. I., and Wikstrom, M. 2010. Proton-Coupled Electron Transfer in Cytochrome Oxidase. Chem. Rev., 110: 7062–7081.
- Youfen Li, Jeong-Soon Park, Jian-Hong Deng, and Yidong Bai. (2006). Cytochrome c oxidase subunit IV is essential for assembly and respiratory function of the enzyme complex. J Bioenerg Biomembr, 38 (5-6): 283–291.
- Carter, K. Palmer, G. 1982. Models of the Two Heme Centers in Cytochrome Oxidase. The Journal of Biological Chemistry, 257(22): 13507-13514.
- Crofts, A. 1994. Cytochrome oxidase: Complex IV. University of Illinois at Urbana-Champaign. Accessed 4 April 2018.
- Tsukihara, Tomitake, et al. 1996. Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 angstroms. Science, 269: 1136-1144.
- Kepp, K. 2017. Heme: From quantum spin crossover to oxygen manager of life. Coordination Chemistry Reviews, 344: 363-374.
- Hellwig, P., Soulimane, T., Buse, G., and Mantele, W. 1999. Electrochemical, FTIR, and UV/VIS Spectroscopic Properties of the ba3 Oxidase from Thermus thermophilus. Biochemistry, 38: 9648-9658.
- Shoubridge, E. A. 2001. Cytochrome c oxidase deficiency. American Journal of Medical Genetics, 106 (1): 46-52.
- U.S. National Library of Medicine: Genetics Home Reference. (2018, March 6). Leigh Syndrome. Retrieved from https://ghr.nlm.nih.gov/condition/leigh-syndrome.
Contributed by
This work was originally written by Allyson Slusser, Spring 2018: Allyson is currently (as of 2018) a senior biology major at Saint Mary's College in Notre Dame, IN.
This work was originally edited by Dr. Kathryn Haas (Assistant Professor) and Madison Sendzik (Teaching and Research Assistant) at Saint Mary's College.


