Antioxidant: Cu/Zn Superoxide dismutase (SOD1)
- Page ID
- 67336
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)This is a video introduction to the Cu,Zn superoxide dismutase (SOD1), which acts as an antioxidant in the cytoplasm of mammalian cells. (PDB 1sxz)
The role of the copper-zinc superoxide dismutase system (SOD1) is to catalyze the conversion of the potentially toxic superoxide anion, O2·-, to the less toxic substance hydrogen peroxide, H2O2. Dioxygen, O2, is also a product of this reaction.1 Superoxide is produced in aerobes as a product of metabolism. The superoxide anion can be an intermediate or a byproduct of metabolic processes such as respiration or photosynthesis.2 Superoxide, O2·-, is toxic when in the protonated form, HO2·, and can cause damage to the cells.1 Superoxide is involved in the oxidative damage phenomena also known as oxidative stress.2 Oxidative stress is harmful and it is caused by radicals, such as superoxide, that can attack lipids, proteins, and DNA within the body. Oxidative stress is linked to many human diseases including hypertension, diabetes mellitus, ischemic diseases, and malignancies.3 Cells use superoxide dismutase in order to prevent oxidative damage by removing superoxide free radicals.2
Superoxide dismutase participates in a dismutation reaction in order to convert superoxide into the less toxic substances of hydrogen peroxide and dioxygen. A dismutation reaction is a reaction in which one substance is oxidized and the other is reduced simultaneously.5 There are two redox reactions that occur in the copper zinc superoxide dismutase system, and copper catalyzes these reactions. In one reaction Cu2+ is reduced to Cu1+, while the superoxide molecule, O2-, is oxidized to O2. In the second step Cu1+ is oxidized to Cu2+, while a second superoxide molecule is reduced to hydrogen peroxide, H2O2.7 A simplified catalytic cycle can be seen in the Figure below.
The overall reduction potential of the overall reaction is dependent on the half reactions which are:
\[O_2 → e^- + O_2^-\]
with E = -0.16 V
\[O_2 + e^- → 2H^+ + H_2O_2\]
with E = +0.89 V
Therefore, the overall reduction potential of this reaction must be within -0.16V and 0.89V.6
The overall reaction catalyzed by superoxide dismutase enzymes is shown below. When catalyzed by Cu,Zn Superoxide Dismuase (SOD1), the reaction happens in two steps: In one step, Cu+2 at the enzyme active site is reduced to Cu+1 as superoxide is oxidized to dioxygen. (see right of simplified reaction cycle shown below) In another step, Cu+1 is oxidized to Cu+2 as superoxide is reduced to hydrogen peroxide. This step also requires two hydrogen ions (see left of simplified reaction cycle shown below).
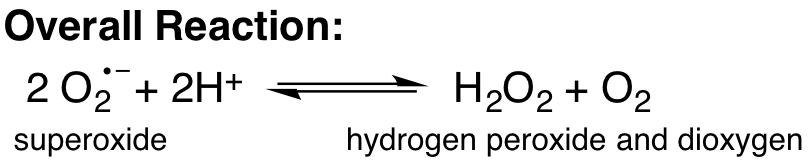
Simplified Catalytic Cycle:

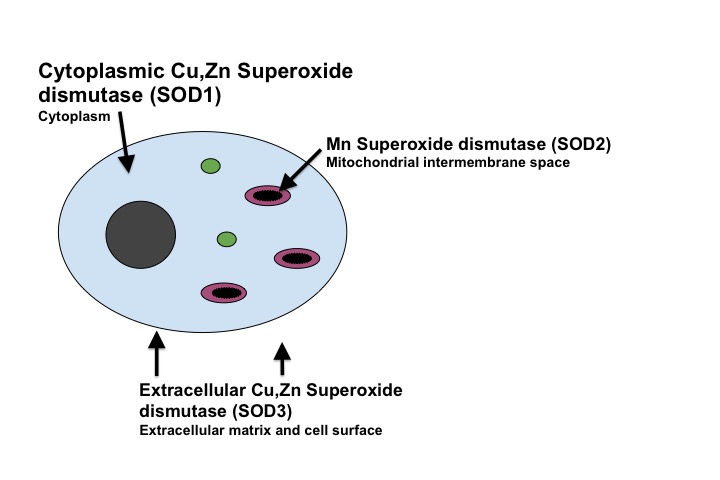
There are three different types of superoxide dismutase that are found in mammals. The primary form of superoxide dismutase is Cu,Zn superoxide dismutase, which is referred to as SOD1. SOD1 is primarily found in the cytoplasm of cells but a small portion is found in the intermembrane space of the mitochondria. A key feature of SOD1 is that is possesses a copper cofactor, which is used to catalyze the superoxide dismutation reaction. A second type of superoxide dismutase is manganese superoxide dismutase, which is referred to as SOD2. Manganese superoxide dismutase is found in the mitochondrial matrix of the cell. SOD2 is produced in the cytoplasm and then transported to the mitochondrial matrix. Manganese (Mn) is the metal cofactor used to catalyze the conversion of superoxide in SOD2. A third type of mammalian superoxide dismutase is the extracellular Cu,Zn superoxide dismutase, referred to as SOD3. SOD3 is primarily found in tissues on the extracellular matrix and on cell surfaces. There is a small amount of SOD3 that is found in plasma and extracellular fluids. SOD3 is similar to SOD1 in that it poseses copper at its active site.4 This page focuses specifically on SOD1.
This figure illustrates the three types of SOD found in mammals, and their primary localization in reference to the cell.
Amyotrophic lateral sclerosis (ALS), the disease commonly referred to as Lou Gehrig’s disease, can be caused by a genetic mutation in SOD1. About one tenth of all cases of ALS are hereditary, and in these hereditary cases of ALS there are mutations in the SOD1 gene. Much research has been aimed at understanding how this mutation in SOD1 causes ALS, with hopes of developing better treatments or a potential cure for this disease.5
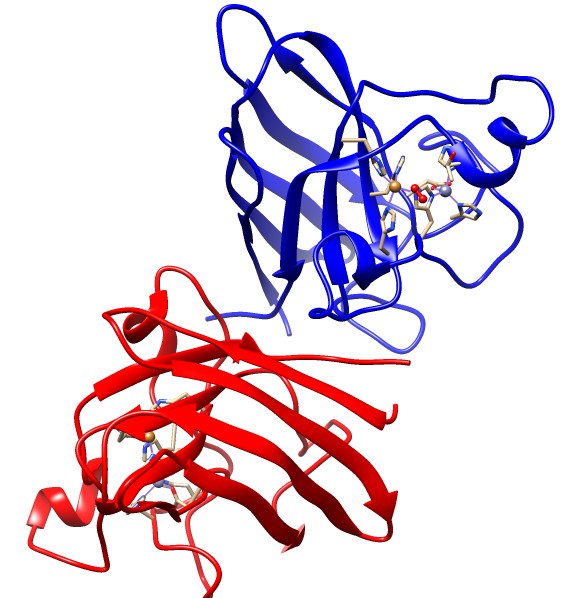
The SOD1 protein is a dimer composed of two identical subunits (homodimer). The two subunits are differentiated in red and blue in the Figure below. The two subunits are oriented so the two active sites are facing away from each other. Each subunit is composed of 151 amino acid residues. These amino acids are arranged in eight β-sheets and three exterior loops. Each subunit contains a copper ion and a zinc ion at the active site. The copper ion acts as an electron transfer center in this catalyst, while the zinc ion serves as a structural scaffold and a contributes to the positive charge of the active site, which attracts the negatively charged superoxide anion.
Ribbon diagram of the reduced form of Cu,Zn SOD with two subunits differentiated in blue and red. Active site side chains and metal cofactors are shown with superoxide bound. (Modified from PDB 1sxz). 6
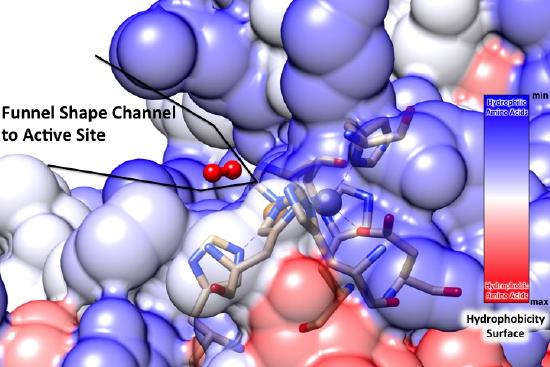
The placement of the active site of this complex as well as the geometry surrounding the copper metal centers are critical for this reaction to occur. The active site of this complex is at the bottom of a channel. The copper ion is exposed at the bottom of this channel to allow binding of superoxide to the copper ion, while the zinc ion is buried within the protein.8 This funnel shape leading to the active site is very important to the superoxide dismutase function.9 Superoxide dismutase has a very small active site compared to the entire size of the enzyme. The small funnel shape is selective towards small molecules such as superoxide, as seen in the Figure below. In the funnel shaped channel are copper and zinc cations as well as two positively charged hydrophilic amino acids, histidine and arginine. The strong positive charge provided by these amino acids and metal ions attracts the negatively charged superoxide to the active site where the superoxide can be converted into less toxic substances.5
At the active site of Cu,Zn superoxide dismutase, copper provides a redox-active binding site for the superoxide anion, while zinc provides structural support and positve charge.1 There are several histidine amino acids that surround the copper and zinc metal centers. The copper ion is bound to either three or four histidine side chains, and may also be bound to either water or superoxide, depending on the copper oxidation state and the step in the catalytic cycle (see Figure below). The zinc coordination is more static, and zinc is consistently bound to three hisidine and one aspartate side chain (see Figure below).
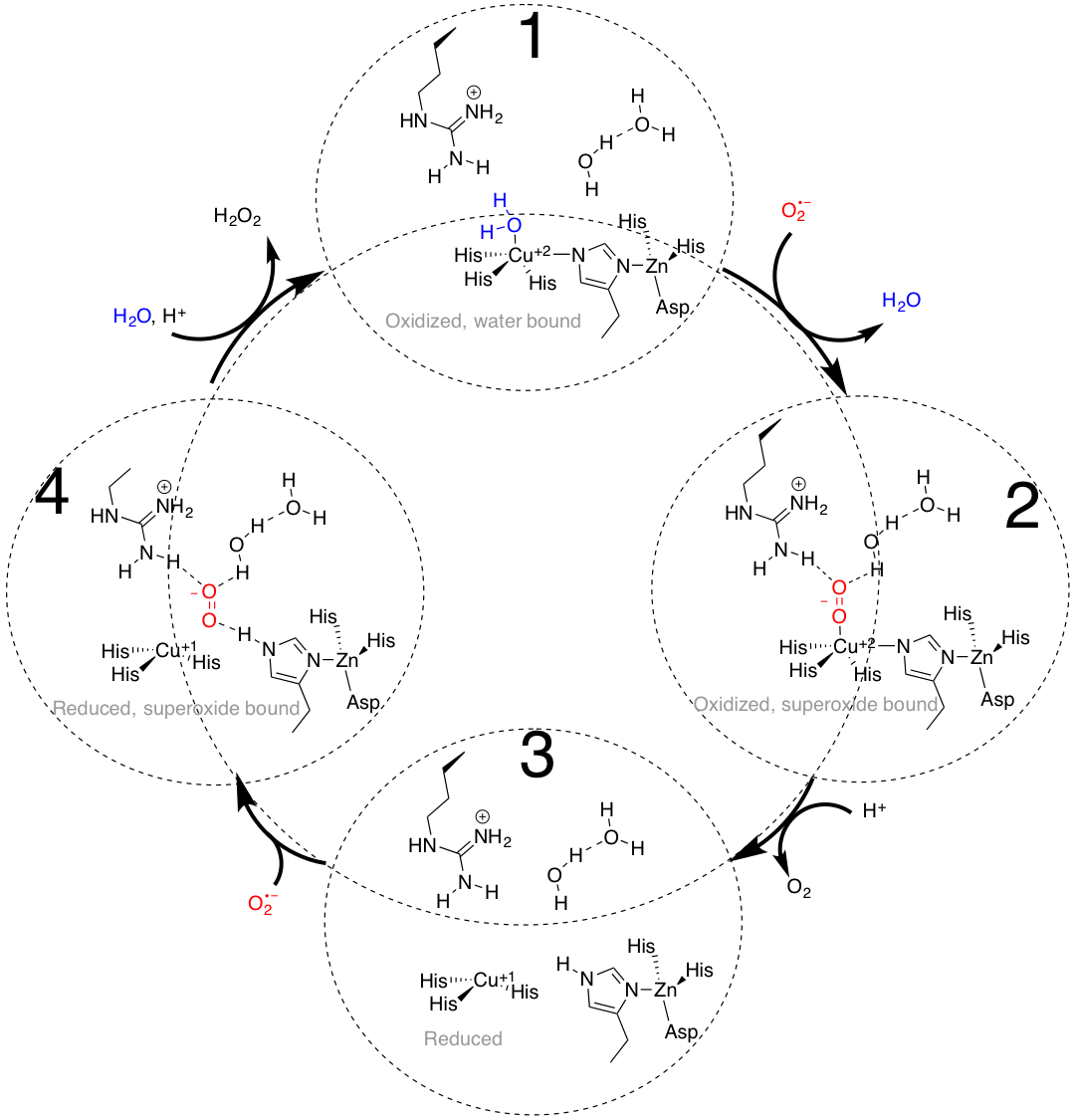
The four steps of the detailed catalysts cycle are shown below. This catalytic cycle alternates between electron counts of 19 and 16. Structure 1 below is of the copper(II) metal center with a four histidine ligands and a water ligand. This five coordinate system yields an electron count of 19. Then the water molecule is displaced as the superoxide molecule binds to copper(II). This is a ligand substitution reaction and once again the structure that is formed, seen as the structure on the right of Figure 13, has 19 electrons. When copper(II) is reduced to copper(I), there are only three histidine ligands that are attached. This copper(I) structure in a trigonal planar geometry is illustrated below as tStructure 3. This copper(I) structure with three ligands attached yields an electron count of 16. Then superoxide binds near copper in the active site, and the copper(I) is oxidized to copper(II), while the superoxide molecule is reduced to hydrogen peroxide. A water binds to the oxidixed Cu+2, to return to structure 1 with an electron count of 19. This continuous cycle converts the superoxide molecule to hydrogen peroxide and dioxygen.9,10
The detailed catalytic cycle at the active site of SOD1. Adapted from reference 9.
Hard-soft acid-base (HSAB) theory predicts that "like" classes of acids and bases prefer to interact (ex: hard acid + hard base is better than hard + soft), thus this binding site of primarily borderline bases is ideal for their borderline metal cations. The aspartate, which is bound to zinc and is a hard donor atom, may provide charge balance.According to Ligand Field Theory (LFT), histidine amino acids are strong σ donors and are π acceptors. Thus, histidine is a relatively strong field ligand ,compared to water and other amino acid side chain ligands, and would encourage a relatively large Δ. The aspartate is both a σ donor and π donor, and is a weak field ligand.
Ligand field theory would predict the geometry surrounding the copper(II) metal center as Jahn-Teller distorted octahedron. This would create either a square pyramid or square plane geometry. The geometry that surrounds the copper(II) metal center, however, is a distorted square pyramid when ether water or superoxide is bound (see structures 1 and 2 in the figure above).1 This distorted from the ideal geometry for a d9 metal center such as copper(II). However, this causes a slight destabilization of the copper(II) metal center, which helps promote copper into its d10, monovalent oxidation state. In other words, the destabilization of Cu(II) allows redox cycling to occur more easily (this is also related to Marcus theory, and this article should be elaborated to describe application of Marcus Theory). If the geometry was a perfect square pyramid, the Cu2+ would be more stable, and it would be more difficult to reduce to Cu+1. In the redox reaction, the copper(II) metal center oxidizes the superoxide molecule to dioxygen, while the copper(II) metal center is reduced to copper(I). This step (from 2 to 3 in the figure above) occurs via an inner sphere electron transfer as the superoxide molecule is bound directly to the copper(II) metal center.8 Upon reduction of copper, the dioxygen product is immediately released (structure 3, figure above). According to hard-soft acid-base theory, hard acids are attracted to hard bases and soft acids are attracted to soft bases. The superoxide molecule is a hard base and copper(II) is a borderline acid. Thus the binding of superoxide directly to the copper(II) metal is not surpising based on their close hard-soft acid-base character.
Ligand field theory cannot predict a preferred geometry surrounding the copper(I) metal center, as Cu+1 is a d10 metal, and there is no ligand-field stabilization energy. The geometry that surrounds the copper(I) metal center in SOD1 is a distorted trigonal plane (structures 3 and 4, figure above).1 In conversion from structure structure 3 to 4 in the figure above, superoxide binds in the active site through H-bonding and electrostatic attraction, however does not directly coordinate to the copper center. In conversion of structure 4 to 1, the copper(I) metal center reduces the superoxide molecule to hydrogen peroxide, while copper(I) is oxidized to copper(II).9 The fact that superoxide binds directly to copper(II) in the first part of the cycle (structure 1 to 2 above), but does not bind directly to copper(I) in the second part of the cycle (structure 3 to 4 above) is not surprising based on hard-soft acid-base principles. According to hard-soft acid-base theory, the soft copper(I) ion is not a preferred acid for the hard superoxide base.
A "snapshot" of the Cu,Zn SOD active site, adapted from PDB structure 1SXZ, is shown below. In the PDB structure, 1SXZ, azide is bound to the reduced copper center. We have converted azide to superoxide.
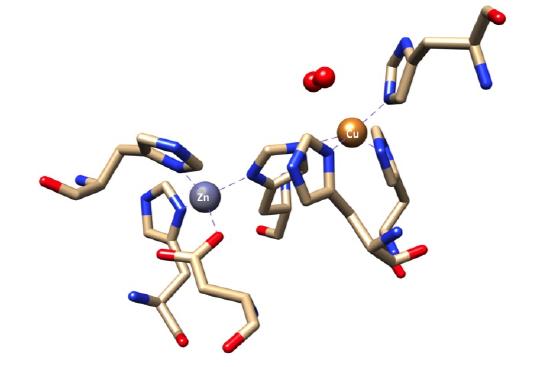
The active site of SOD1 has a zinc metal center that is present for structural support and stabilization. Ligand field theory does not play a role in the determination of the geometry surrounding the zinc metal center since zinc is a d10 metal and does not have ligand field stabilization energy. Instead. the geometry is determined by steric preference of ligands and structural constraints imposed by the protein. For example, the tetrahedral geometry, with bond angels of 109.5° is preferred in terns of sterics compared to a square plane, with smaller bond angles of 90°. Thus, the geometry that surrounds the zinc metal center is distorted tetrahedron which is the favored geometry due to sterics and structural constraints imposed by the polypeptide.1
The crystal field splitting diagrams for the copper metal center is a distorted square pyramid geometry and the zinc metal center in a tetrahedron geometry are shown below.
Cu2+ is a d9 metal; therefore, there is an empty orbital for an electron to move up in energy as seen in Figure 11. Therefore, d-d transitions can occur in this molecule when superoxide dismutase is in its reduced form, Cu2+. However, when superoxide dismutase is in its oxidized form, Cu1+, d-d transitions cannot occur. This is because Cu1+ is a d10 metal. Therefore, all of the orbitals will be filled. Figure 12 is the crystal field splitting diagram of Zn2+ in a tetrahedral geometry. Zn2+ is a d10 metal; therefore, all of the orbitals are filled so d-d transitions cannot occur here. The square pyramidal crystal field splitting diagram indicates that there are electrons in the anti-bonding orbitals, therefore, weakening the bonds. This makes the molecule more labile and more reactive, making the reaction more likely to occur.
The copper-zinc superoxide dismutase system is a necessary molecule in the body that converts the toxic superoxide molecule into the less toxic substances of hydrogen peroxide and dioxygen. The superoxide molecule can cause oxidative damage in the body. There has recently been discovered a connection between a mutation in SOD1 and a hereditary form of Lou Gehrig's disease. This molecule catalyzes the dismutation reaction of the conversion of superoxide. This reaction occurs due to the destabilization of the copper(II) metal center due to the distorted square pyramid geometry and the lability of copper. The zinc metal center is a structural scaffold for this molecule. The copper, zinc superoxide dismutase system is a necessary for the body in order to prevent oxidative damage that can affect lipids, proteins, and DNA.
Sources:
1Miessler, G.L.; Tarr, D.A.; Inorganic Chemistry. 2010. 644-647.
2Pelmenschikov, V.; Siegbahn, P. Copper-zinc superoxide dismutase: theoretical insights into the catalytic mechanism. Inorg. Chem. 2005, 44, 3311-3320.
3Yoshikawa, T.; Naito, Y. What is oxidative stress? JMAJ. 2002, 45, 271-276.
4Fukai, T.; Ushio-Fukai, M. Superoxide dismutases: role in redox signaling, vascular function, and diseases. Antioxid Redox Signal. 2011, 15, 1583-1606.
5Goodsell, D. Superoxide dismutase. Protein Data Bank. 2007.
https://pdb101.rcsb.org/motm/94.
6Ferraroni, M.; Rypniewski, W.; Bruni, B.; Orioli, P.; Mangani, S. Crystallographic determination of reduced bovine superoxide dismutase at pH 5.0 and of anion binding to its active stie. J. Biol. Inorg. Chem. 1998, 3, 411-422.
7Azab, H.A.; Banci, L.; Borsari, M.; Luchinat, C.; Sola, M.; Viezzoli, M.S. Redox chemistry of superoxide dismutase. Cyclic voltammetry of wild-type enzymes and mutants on functionally relevant residues. Inorg. Chem. 1992, 31, 4649-4655.
8Tainer, J.A.; Getzoff, E.D.; Richardson, J.S.; Richardson, D.C. Structure and mechanism of copper, zinc superoxide dismutase. Nature. 1983, 306, 284-287.
9Gray, H.B.; Stiefel, E. I.; Valentine, J.S.; Bertini, I. Biological Inorganic Chemistry: Structure and Reactivity. 2007. 331-337.
10Rakhit, R; Charkrabartty, A. Structure, folding, and misfolding of Cu, Zn superoxide dismutase in amyotrophic lateral sclerosis. Biochimica et Biophysica Acta. 2006. 1762: 1025-1037.
Contributed by:
This work was originally written by Margaret Mraz, Fall 2016: Margaret is currently (as of 2016) a senior chemistry major at Saint Mary's College in Notre Dame, IN.
This work was originally edited by Dr. Kathryn Haas, Assistant Professor at Saint Mary's College.