6.8: Proteins
- Page ID
- 206964
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
- To recognize amino acids and classify them based on the characteristics of their side chains.
The proteins in all living species, from bacteria to humans, are constructed from the same set of 20 amino acids, so called because each contains an amino group attached to a carboxylic acid. The amino acids in proteins are α-amino acids, which means the amino group is attached to the α-carbon of the carboxylic acid. Humans can synthesize only about half of the needed amino acids; the remainder must be obtained from the diet and are known as essential amino acids. However, two additional amino acids have been found in limited quantities in proteins: Selenocysteine was discovered in 1986, while pyrrolysine was discovered in 2002.
The amino acids are colorless, nonvolatile, crystalline solids, melting and decomposing at temperatures above 200°C. These melting temperatures are more like those of inorganic salts than those of amines or organic acids and indicate that the structures of the amino acids in the solid state and in neutral solution are best represented as having both a negatively charged group and a positively charged group. Such a species is known as a zwitterion.
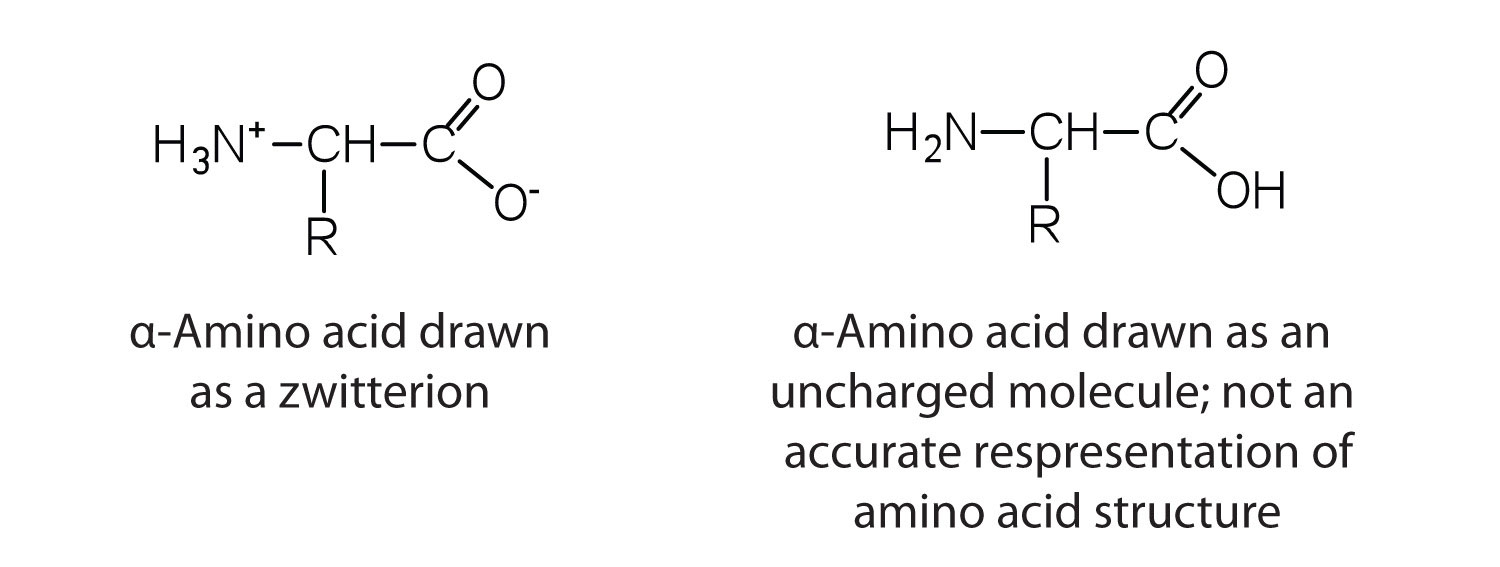
Classification
In addition to the amino and carboxyl groups, amino acids have a side chain or R group attached to the α-carbon. Each amino acid has unique characteristics arising from the size, shape, solubility, and ionization properties of its R group. As a result, the side chains of amino acids exert a profound effect on the structure and biological activity of proteins. Although amino acids can be classified in various ways, one common approach is to classify them according to whether the functional group on the side chain at neutral pH is nonpolar, polar but uncharged, negatively charged, or positively charged. The structures and names of the 20 amino acids, their one- and three-letter abbreviations, and some of their distinctive features are given in Table \(\PageIndex{1}\).
| Common Name | Abbreviation | Structural Formula (at pH 6) | Molar Mass | Distinctive Feature |
|---|---|---|---|---|
| Amino acids with a nonpolar R group | ||||
| glycine | gly (G) | 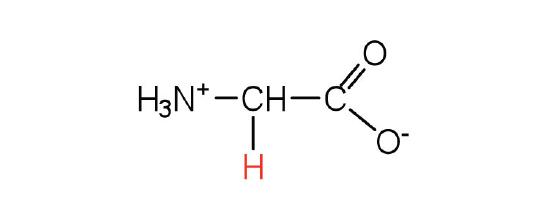 |
75 | the only amino acid lacking a chiral carbon |
| alanine | ala (A) | 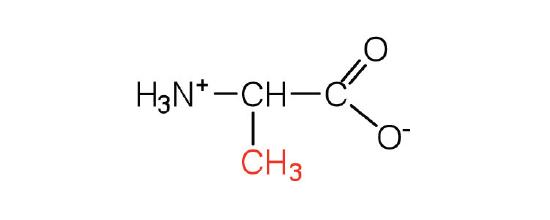 |
89 | — |
| valine | val (V) | 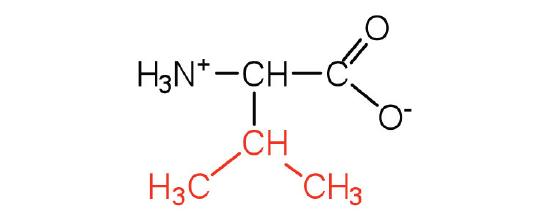 |
117 | a branched-chain amino acid |
| leucine | leu (L) | 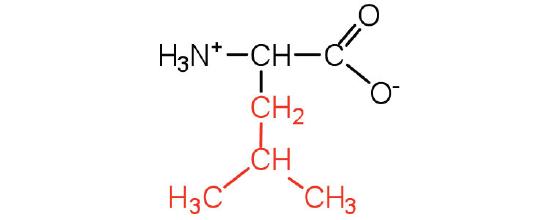 |
131 | a branched-chain amino acid |
| isoleucine | ile (I) | 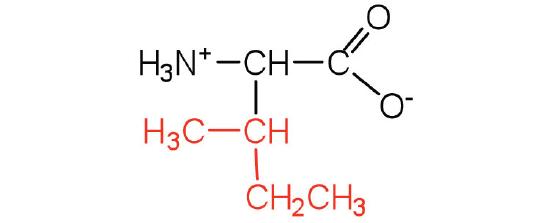 |
131 | an essential amino acid because most animals cannot synthesize branched-chain amino acids |
| phenylalanine | phe (F) | 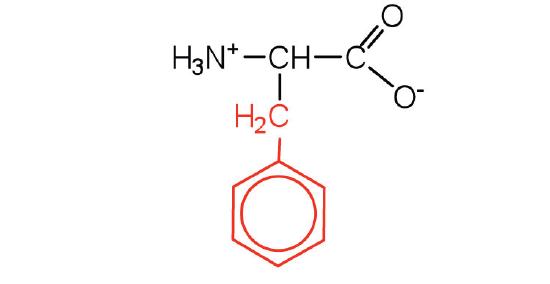 |
165 | also classified as an aromatic amino acid |
| tryptophan | trp (W) | 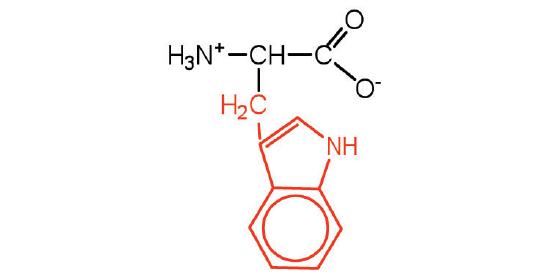 |
204 | also classified as an aromatic amino acid |
| methionine | met (M) | 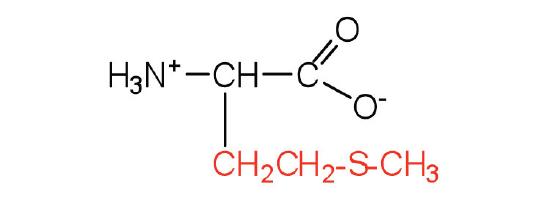 |
149 | side chain functions as a methyl group donor |
| proline | pro (P) | 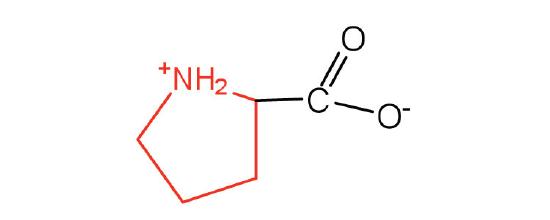 |
115 | contains a secondary amine group; referred to as an α-imino acid |
| Amino acids with a polar but neutral R group | ||||
| serine | ser (S) | 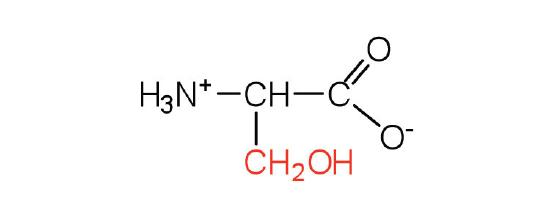 |
105 | found at the active site of many enzymes |
| threonine | thr (T) | 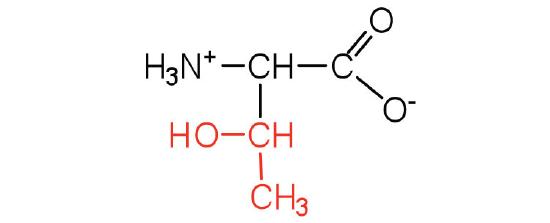 |
119 | named for its similarity to the sugar threose |
| cysteine | cys (C) | 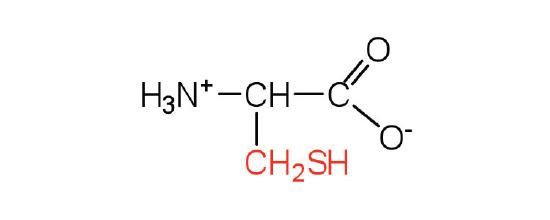 |
121 | oxidation of two cysteine molecules yields cystine |
| tyrosine | tyr (Y) | 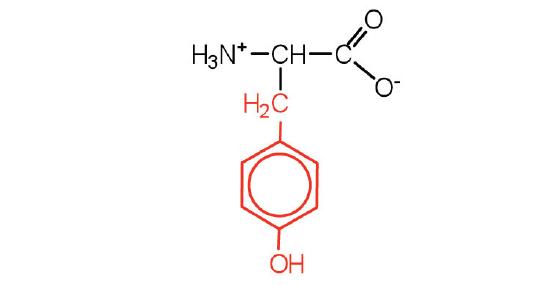 |
181 | also classified as an aromatic amino acid |
| asparagine | asn (N) | 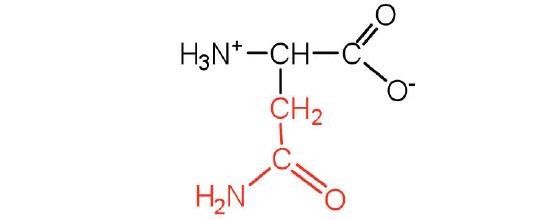 |
132 | the amide of aspartic acid |
| glutamine | gln (Q) | 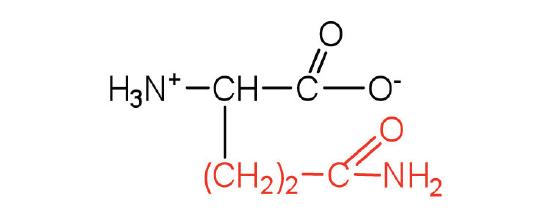 |
146 | the amide of glutamic acid |
| Amino acids with a negatively charged R group | ||||
| aspartic acid | asp (D) | 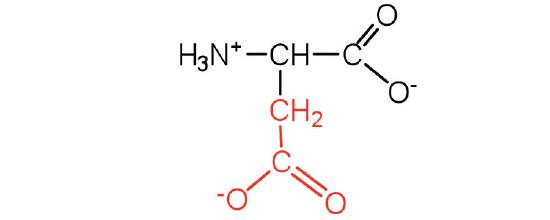 |
132 | carboxyl groups are ionized at physiological pH; also known as aspartate |
| glutamic acid | glu (E) | 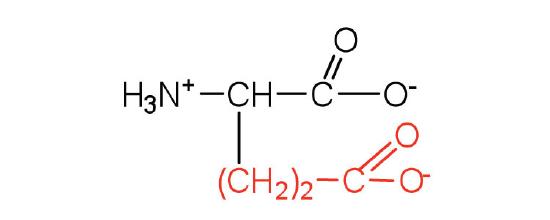 |
146 | carboxyl groups are ionized at physiological pH; also known as glutamate |
| Amino acids with a positively charged R group | ||||
| histidine | his (H) | 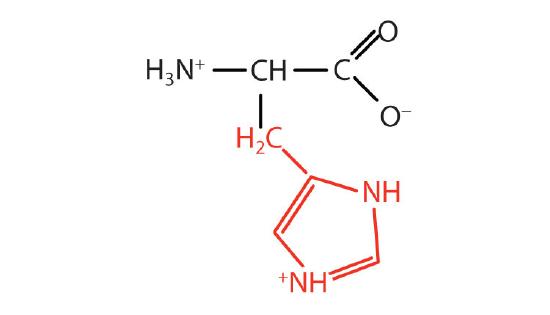 |
155 | the only amino acid whose R group has a pKa (6.0) near physiological pH |
| lysine | lys (K) | 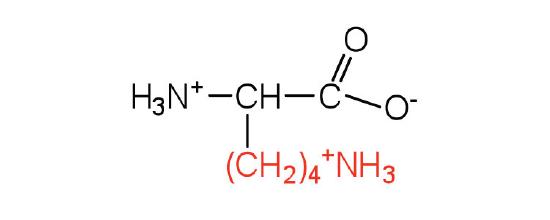 |
147 | — |
| arginine | arg (R) | 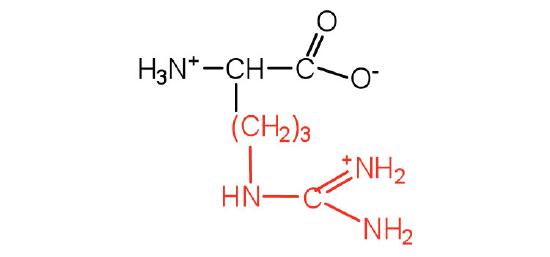 |
175 | almost as strong a base as sodium hydroxide |
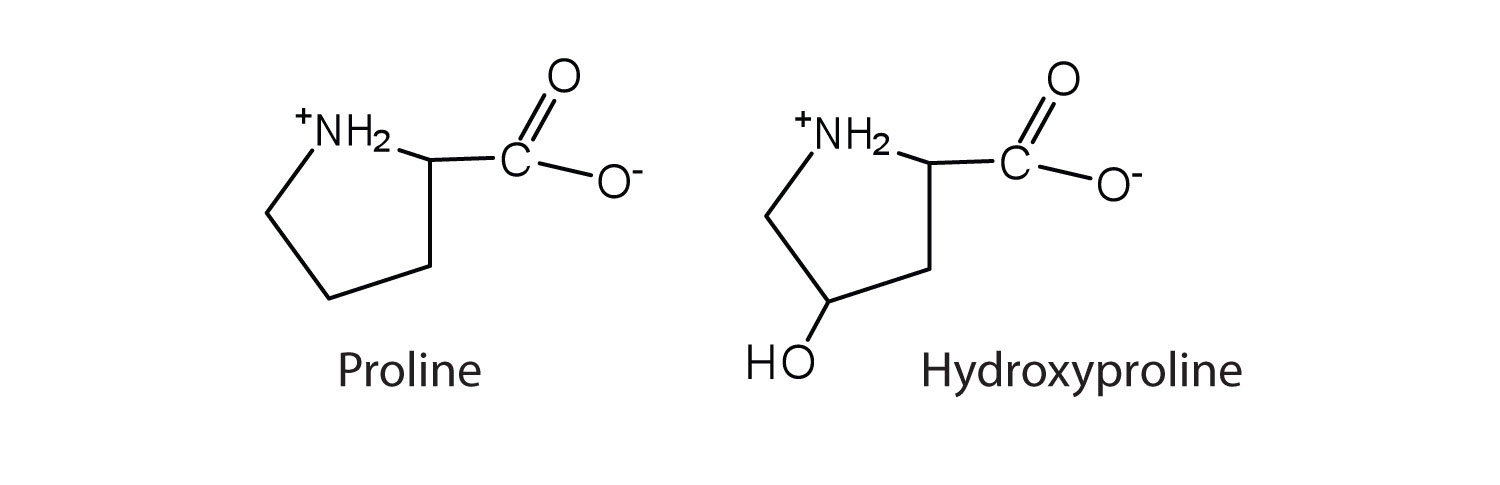
The first amino acid to be isolated was asparagine in 1806. It was obtained from protein found in asparagus juice (hence the name). Glycine, the major amino acid found in gelatin, was named for its sweet taste (Greek glykys, meaning “sweet”). In some cases an amino acid found in a protein is actually a derivative of one of the common 20 amino acids (one such derivative is hydroxyproline). The modification occurs after the amino acid has been assembled into a protein.
Configuration
Notice in Table \(\PageIndex{1}\) that glycine is the only amino acid whose α-carbon is not chiral. Therefore, with the exception of glycine, the amino acids could theoretically exist in either the D- or the L-enantiomeric form and rotate plane-polarized light. As with sugars, chemists used L-glyceraldehyde as the reference compound for the assignment of absolute configuration to amino acids. Its structure closely resembles an amino acid structure except that in the latter, an amino group takes the place of the OH group on the chiral carbon of the L-glyceraldehyde and a carboxylic acid replaces the aldehyde. Modern stereochemistry assignments using the Cahn-Ingold-Prelog priority rules used ubiquitously in chemistry show that all of the naturally occurring chiral amino acids are S except Cys which is R.
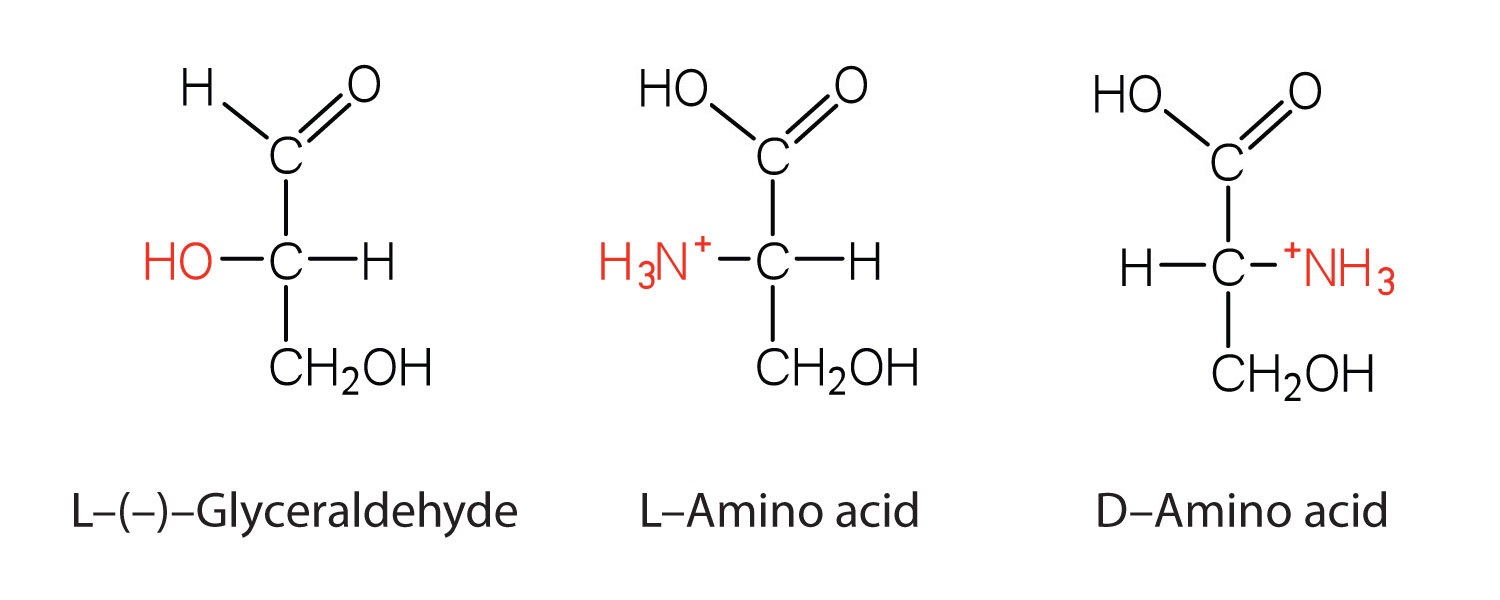
We learned that all naturally occurring sugars belong to the D series. It is interesting, therefore, that nearly all known plant and animal proteins are composed entirely of L-amino acids. However, certain bacteria contain D-amino acids in their cell walls, and several antibiotics (e.g., actinomycin D and the gramicidins) contain varying amounts of D-leucine, D-phenylalanine, and D-valine.
Summary
Amino acids can be classified based on the characteristics of their distinctive side chains as nonpolar, polar but uncharged, negatively charged, or positively charged. The amino acids found in proteins are L-amino acids.
- Describe the four levels of protein structure.
- Identify the types of attractive interactions that hold proteins in their most stable three-dimensional structure.
- Explain what happens when proteins are denatured.
- Identify how a protein can be denatured.
Each of the thousands of naturally occurring proteins has its own characteristic amino acid composition and sequence that result in a unique three-dimensional shape. Since the 1950s, scientists have determined the amino acid sequences and three-dimensional conformation of numerous proteins and thus obtained important clues on how each protein performs its specific function in the body.
Proteins are compounds of high molar mass consisting largely or entirely of chains of amino acids. Because of their great complexity, protein molecules cannot be classified on the basis of specific structural similarities, as carbohydrates and lipids are categorized. The two major structural classifications of proteins are based on far more general qualities: whether the protein is (1) fiberlike and insoluble or (2) globular and soluble. Some proteins, such as those that compose hair, skin, muscles, and connective tissue, are fiberlike. These fibrous proteins are insoluble in water and usually serve structural, connective, and protective functions. Examples of fibrous proteins are keratins, collagens, myosins, and elastins. Hair and the outer layer of skin are composed of keratin. Connective tissues contain collagen. Myosins are muscle proteins and are capable of contraction and extension. Elastins are found in ligaments and the elastic tissue of artery walls.
Globular proteins, the other major class, are soluble in aqueous media. In these proteins, the chains are folded so that the molecule as a whole is roughly spherical. Familiar examples include egg albumin from egg whites and serum albumin in blood. Serum albumin plays a major role in transporting fatty acids and maintaining a proper balance of osmotic pressures in the body. Hemoglobin and myoglobin, which are important for binding oxygen, are also globular proteins.
Levels of Protein Structure
The structure of proteins is generally described as having four organizational levels. The first of these is the primary structure, which is the number and sequence of amino acids in a protein’s polypeptide chain or chains, beginning with the free amino group and maintained by the peptide bonds connecting each amino acid to the next. The primary structure of insulin, composed of 51 amino acids, is shown in Figure \(\PageIndex{1}\).
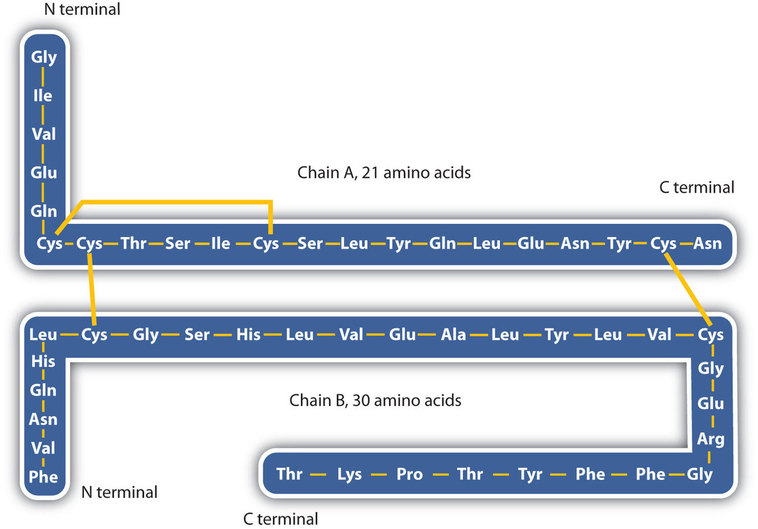
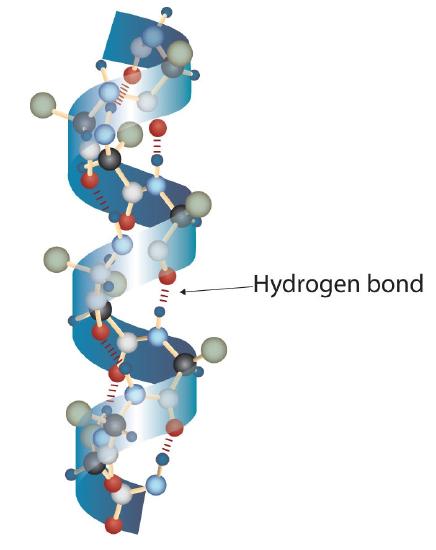
A protein molecule is not a random tangle of polypeptide chains. Instead, the chains are arranged in unique but specific conformations. The term secondary structure refers to the fixed arrangement of the polypeptide backbone. On the basis of X ray studies, Linus Pauling and Robert Corey postulated that certain proteins or portions of proteins twist into a spiral or a helix. This helix is stabilized by intrachain hydrogen bonding between the carbonyl oxygen atom of one amino acid and the amide hydrogen atom four amino acids up the chain (located on the next turn of the helix) and is known as a right-handed α-helix. X ray data indicate that this helix makes one turn for every 3.6 amino acids, and the side chains of these amino acids project outward from the coiled backbone (Figure \(\PageIndex{2}\)). The α-keratins, found in hair and wool, are exclusively α-helical in conformation. Some proteins, such as gamma globulin, chymotrypsin, and cytochrome c, have little or no helical structure. Others, such as hemoglobin and myoglobin, are helical in certain regions but not in others.
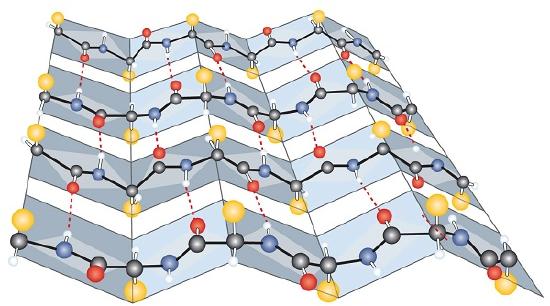
Another common type of secondary structure, called the β-pleated sheet conformation, is a sheetlike arrangement in which two or more extended polypeptide chains (or separate regions on the same chain) are aligned side by side. The aligned segments can run either parallel or antiparallel—that is, the N-terminals can face in the same direction on adjacent chains or in different directions—and are connected by interchain hydrogen bonding (Figure \(\PageIndex{3}\)). The β-pleated sheet is particularly important in structural proteins, such as silk fibroin. It is also seen in portions of many enzymes, such as carboxypeptidase A and lysozyme.
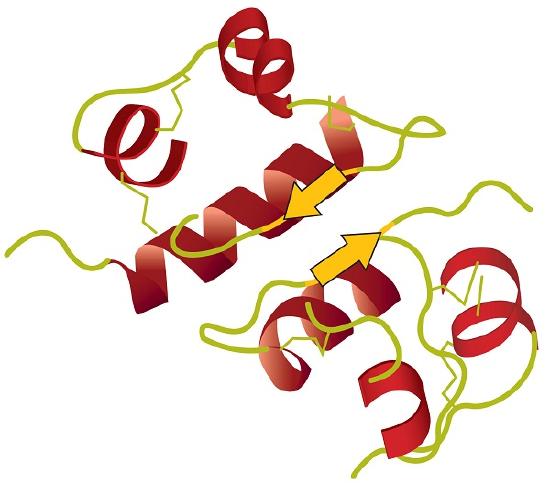
Tertiary structure refers to the unique three-dimensional shape of the protein as a whole, which results from the folding and bending of the protein backbone. The tertiary structure is intimately tied to the proper biochemical functioning of the protein. Figure \(\PageIndex{4}\) shows a depiction of the three-dimensional structure of insulin.
Four major types of attractive interactions determine the shape and stability of the tertiary structure of proteins. You studied several of them previously.
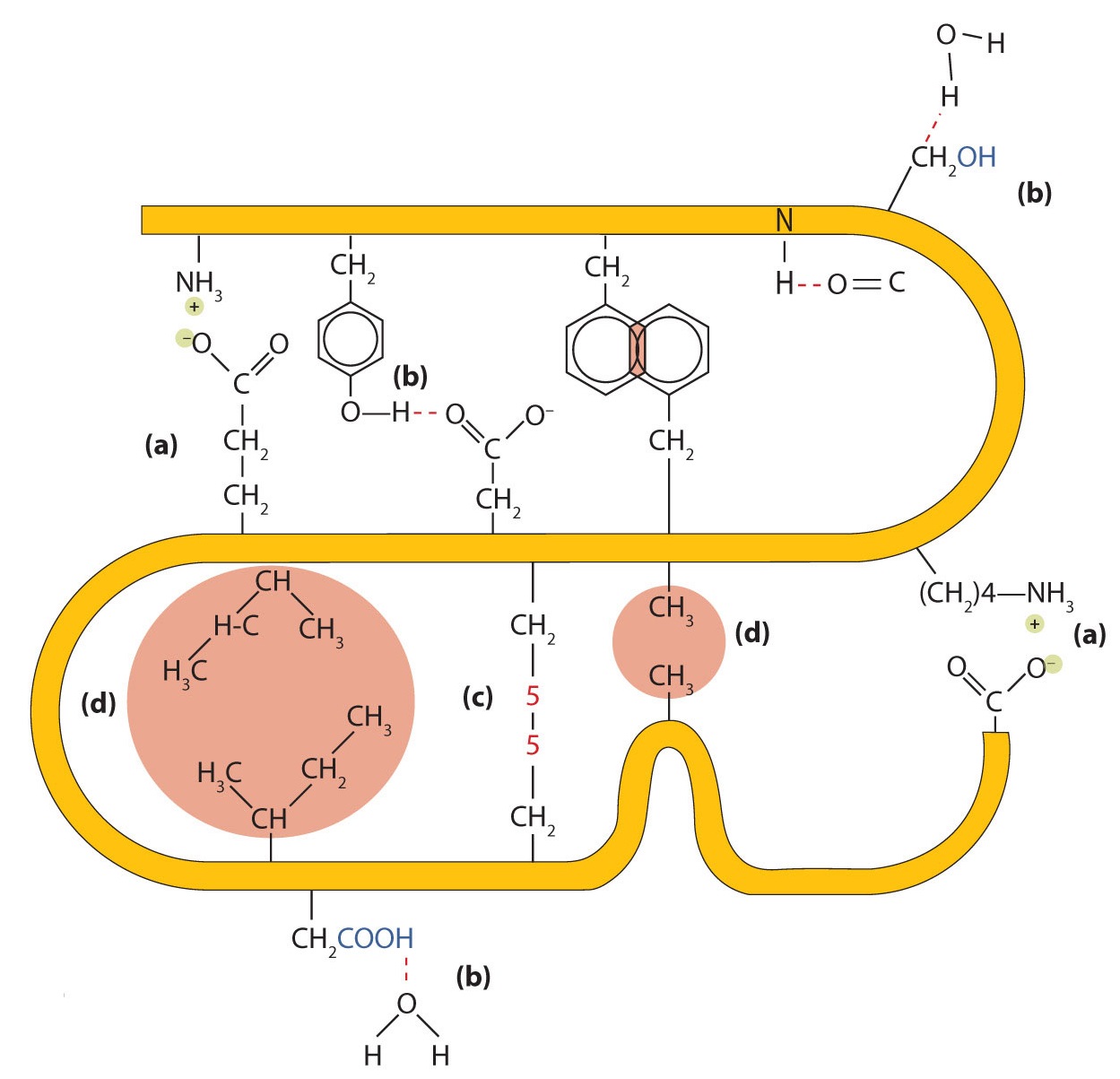
- Ionic bonding. Ionic bonds result from electrostatic attractions between positively and negatively charged side chains of amino acids. For example, the mutual attraction between an aspartic acid carboxylate ion and a lysine ammonium ion helps to maintain a particular folded area of a protein (part (a) of Figure \(\PageIndex{5}\)).
- Hydrogen bonding. Hydrogen bonding forms between a highly electronegative oxygen atom or a nitrogen atom and a hydrogen atom attached to another oxygen atom or a nitrogen atom, such as those found in polar amino acid side chains. Hydrogen bonding (as well as ionic attractions) is extremely important in both the intra- and intermolecular interactions of proteins (part (b) of Figure \(\PageIndex{5}\)).
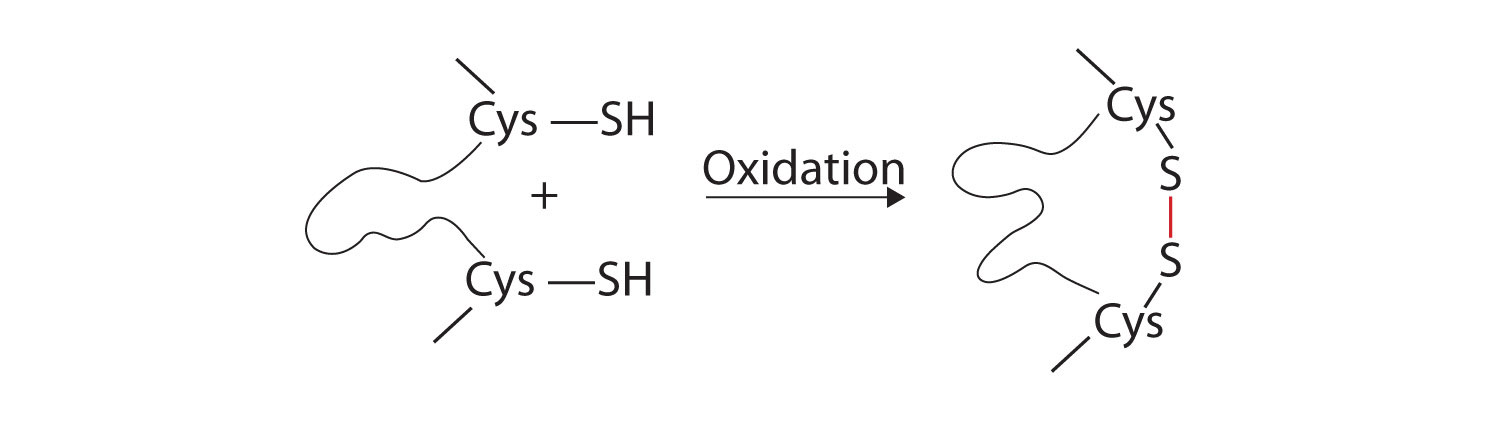
- Disulfide linkages. Two cysteine amino acid units may be brought close together as the protein molecule folds. Subsequent oxidation and linkage of the sulfur atoms in the highly reactive sulfhydryl (SH) groups leads to the formation of cystine (part (c) of Figure \(\PageIndex{5}\)). Intrachain disulfide linkages are found in many proteins, including insulin (yellow bars in Figure \(\PageIndex{1}\)) and have a strong stabilizing effect on the tertiary structure.
- Dispersion forces. Dispersion forces arise when a normally nonpolar atom becomes momentarily polar due to an uneven distribution of electrons, leading to an instantaneous dipole that induces a shift of electrons in a neighboring nonpolar atom. Dispersion forces are weak but can be important when other types of interactions are either missing or minimal (part (d) of Figure \(\PageIndex{5}\)). This is the case with fibroin, the major protein in silk, in which a high proportion of amino acids in the protein have nonpolar side chains. The term hydrophobic interaction is often misused as a synonym for dispersion forces. Hydrophobic interactions arise because water molecules engage in hydrogen bonding with other water molecules (or groups in proteins capable of hydrogen bonding). Because nonpolar groups cannot engage in hydrogen bonding, the protein folds in such a way that these groups are buried in the interior part of the protein structure, minimizing their contact with water.
When a protein contains more than one polypeptide chain, each chain is called a subunit. The arrangement of multiple subunits represents a fourth level of structure, the quaternary structure of a protein. Hemoglobin, with four polypeptide chains or subunits, is the most frequently cited example of a protein having quaternary structure (Figure \(\PageIndex{6}\)). The quaternary structure of a protein is produced and stabilized by the same kinds of interactions that produce and maintain the tertiary structure. A schematic representation of the four levels of protein structure is in Figure \(\PageIndex{7}\).
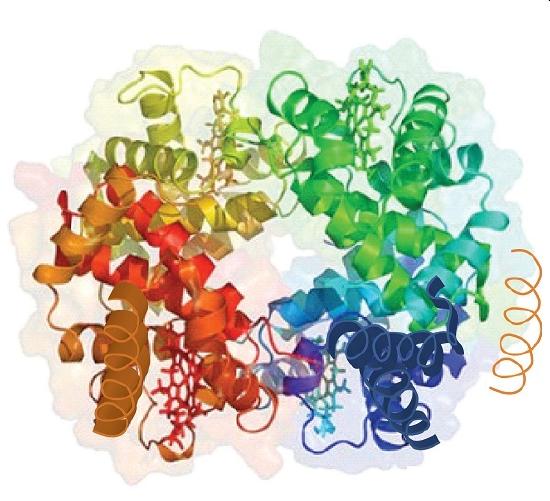
Source: Image from the RCSB PDB (www.pdb.org(opens in new window)) of PDB ID 1I3D (R.D. Kidd, H.M. Baker, A.J. Mathews, T. Brittain, E.N. Baker (2001) Oligomerization and ligand binding in a homotetrameric hemoglobin: two high-resolution crystal structures of hemoglobin Bart's (gamma(4)), a marker for alpha-thalassemia. Protein Sci. 1739–1749).
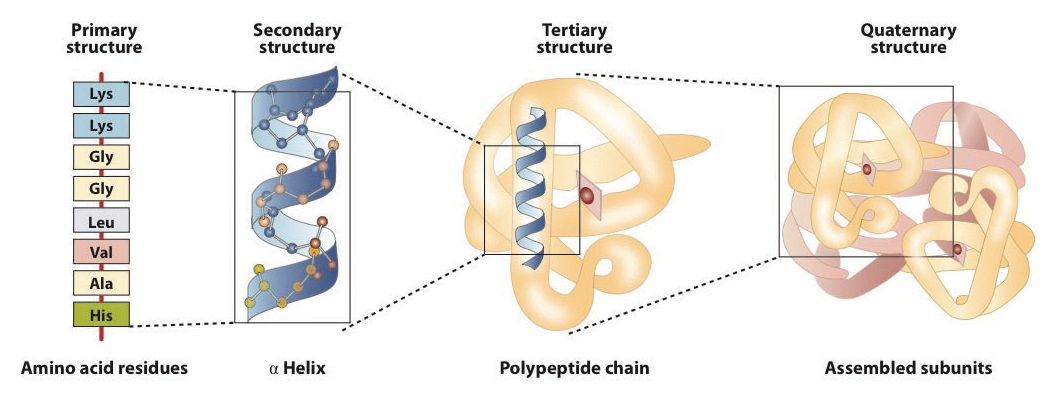
The primary structure consists of the specific amino acid sequence. The resulting peptide chain can twist into an α-helix, which is one type of secondary structure. This helical segment is incorporated into the tertiary structure of the folded polypeptide chain. The single polypeptide chain is a subunit that constitutes the quaternary structure of a protein, such as hemoglobin that has four polypeptide chains.
Denaturation of Proteins
The highly organized structures of proteins are truly masterworks of chemical architecture. But highly organized structures tend to have a certain delicacy, and this is true of proteins. Denaturation is the term used for any change in the three-dimensional structure of a protein that renders it incapable of performing its assigned function. A denatured protein cannot do its job. (Sometimes denaturation is equated with the precipitation or coagulation of a protein; our definition is a bit broader.) A wide variety of reagents and conditions, such as heat, organic compounds, pH changes, and heavy metal ions can cause protein denaturation (Figure \(\PageIndex{1}\)).
| Method | Effect on Protein Structure |
|---|---|
| Heat above 50°C or ultraviolet (UV) radiation | Heat or UV radiation supplies kinetic energy to protein molecules, causing their atoms to vibrate more rapidly and disrupting relatively weak hydrogen bonding and dispersion forces. |
| Use of organic compounds, such as ethyl alcohol | These compounds are capable of engaging in intermolecular hydrogen bonding with protein molecules, disrupting intramolecular hydrogen bonding within the protein. |
| Salts of heavy metal ions, such as mercury, silver, and lead | These ions form strong bonds with the carboxylate anions of the acidic amino acids or SH groups of cysteine, disrupting ionic bonds and disulfide linkages. |
| Alkaloid reagents, such as tannic acid (used in tanning leather) | These reagents combine with positively charged amino groups in proteins to disrupt ionic bonds. |
Anyone who has fried an egg has observed denaturation. The clear egg white turns opaque as the albumin denatures and coagulates. No one has yet reversed that process. However, given the proper circumstances and enough time, a protein that has unfolded under sufficiently gentle conditions can refold and may again exhibit biological activity (Figure \(\PageIndex{8}\)). Such evidence suggests that, at least for these proteins, the primary structure determines the secondary and tertiary structure. A given sequence of amino acids seems to adopt its particular three-dimensional arrangement naturally if conditions are right.

The primary structures of proteins are quite sturdy. In general, fairly vigorous conditions are needed to hydrolyze peptide bonds. At the secondary through quaternary levels, however, proteins are quite vulnerable to attack, though they vary in their vulnerability to denaturation. The delicately folded globular proteins are much easier to denature than are the tough, fibrous proteins of hair and skin.
Summary
Proteins can be divided into two categories: fibrous, which tend to be insoluble in water, and globular, which are more soluble in water. A protein may have up to four levels of structure. The primary structure consists of the specific amino acid sequence. The resulting peptide chain can form an α-helix or β-pleated sheet (or local structures not as easily categorized), which is known as secondary structure. These segments of secondary structure are incorporated into the tertiary structure of the folded polypeptide chain. The quaternary structure describes the arrangements of subunits in a protein that contains more than one subunit. Four major types of attractive interactions determine the shape and stability of the folded protein: ionic bonding, hydrogen bonding, disulfide linkages, and dispersion forces. A wide variety of reagents and conditions can cause a protein to unfold or denature.


