13.E: Reactions at the α-Carbon, Part II (Exercises)
- Page ID
- 106631
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)P13.1: Tetrahydrolipastatin, a potent inhibitor of lipase enzymes (see section 11.6) is being tested as a possible anti-obesity drug. Lipastatin, a close derivative, is synthesized by the bacterium Streptomyces toxytricini. The biosynthetic pathway involves the following step shown below - draw a likely mechanism. (J. Biol. Chem. 1997, 272, 867).
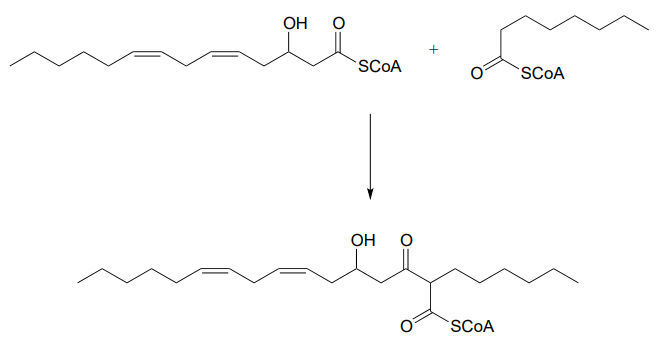
P13.2: The metabolism of camphor by some bacteria involves the step below. Draw a likely mechanism. (J. Biol. Chem 2004, 279, 31312)

P13.3: The glucogeogenesis pathway, by which glucose is synthesized from pyruvate, begins with a reaction catalyzed by pyruvate carboxylase. The enzyme requires the \(CO_2\)-carrying biotin to function, but the final step is thought to be the simple carboxylation of pyruvate by free carbon dioxide (Biochem. J. 2008, 413, 369). Draw a mechanism for this step.
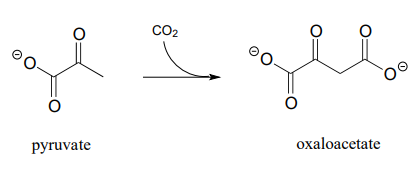
P13.4: Draw a reasonable mechanism for this decarboxylation step in tryptophan biosynthesis (EC 4.1.1.45). Hint: a tautomerization step precedes the decarboxylation.
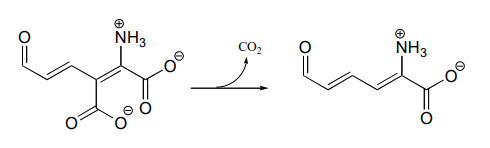
P13.5: The biosynthetic pathway for the antibiotic compound rabelomycin begins with the condensation of malonyl \(CoA\) and acetyl \(CoA\). Predict the product of this reaction, and propose a likely mechanism. (Org. Lett. 2010 12, 2814.)
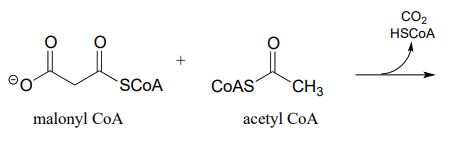
P13.6: Predict the product of this decarboxylation step in the biosynthesis of the amino acid tyrosine. Hint: think about comparative stability when you are considering where protonation will occur!
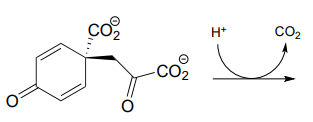
P13.7: Show a likely mechanism for this reaction from lysine biosynthesis:
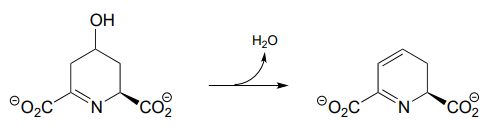
P13.8: Compound A undergoes hydrolytic cleavage in some fungi to form the products shown. Predict the structure of A. (J. Biol. Chem. 2007, 282, 9581)
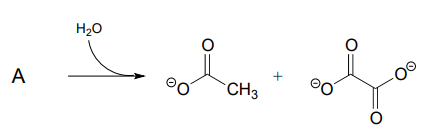
P13.9: Propose a mechanism for the following reaction from the gluconeogenesis pathway (EC 4.1.1.32):
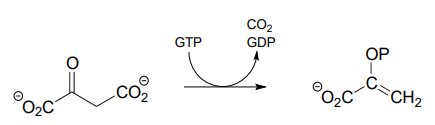
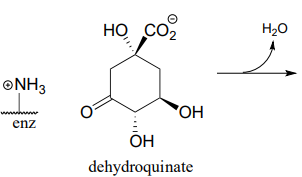
P13.10: Dehydroquinate undergoes dehydration (EC 4.2.1.10) in aromatic amino acid biosynthesis. Experimental and genomic evidence points to a lysine-linked iminium intermediate. More than one dehydration product is possible for dehydroquinate, but in this case the most stable product is the one that forms. Predict the structure of the product, explain why it is the more stable of the possible dehydration products, and draw a mechanism for its formation.
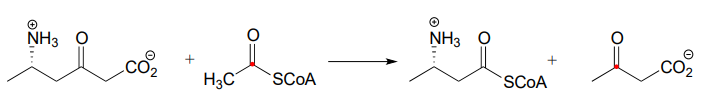
P13.11: The enzyme catalyzing the reaction below, thought to participate in the fermentation of lysine in bacteria, was recently identified and characterized (J. Biol. Chem. 2011, 286, 27399). Propose a likely mechanism. Hint: the mechanism involves two separate carbon-carbon bond forming and bond breaking steps. \(C_1\) of acetyl \(CoA\)is identified with a red dot to help you trace it through to the product.
P13.12: Menaquinone (Vitamin K) biosynthesis in bacteria includes the following step:
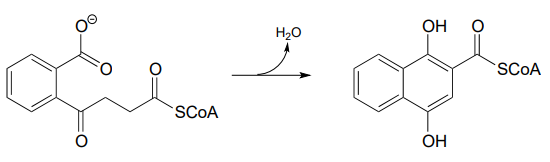
Propose a likely mechanism. Hint: the mechanism involves a Claisen condensation step which is unusual in that the electrophile is a carboxylic acid group rather than a thioester. What is the driving force that allows this unusual step to occur? (J. Biol. Chem. 2010, 285, 30159)
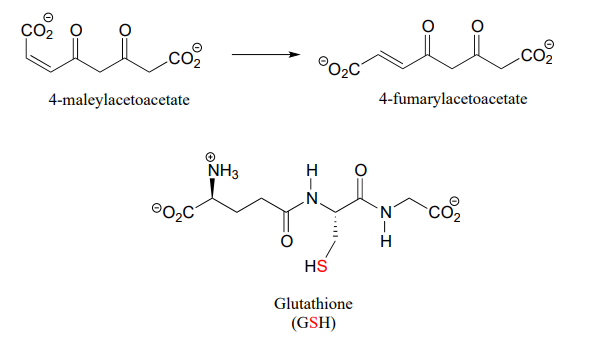
P13.13: 4-maleylacetoacetate isomerase (EC 5.2.1.2) catalyzes the following cis to trans alkene isomerization as part of the degradation of the aromatic amino acids phenylalanine and tyrosine.
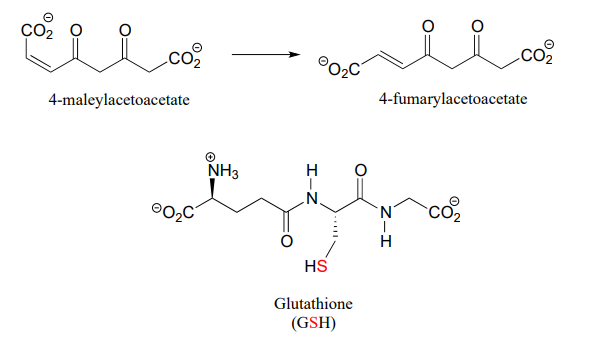
The enzyme uses the thiol-containing coenzyme glutathione, which is also involved in the formation of disulfide bonds in proteins, but in this case glutathione serves as a 'thiol group for hire'. The mechanism for the reaction is essentially a reversible conjugate addition of glutathione. Draw out the steps for this mechanism, showing how the cis-trans isomerization could be accomplished. Also, explain why the equilibrium for this reaction favors 4-fumarylacetoacetate. The structure of glutathione is shown, but you can use the abbreviation GSH in your mechanism.
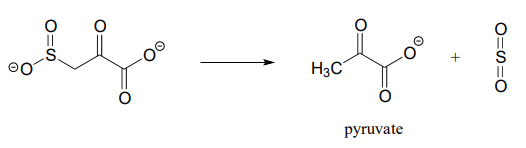
P13.14: Based on the mechanistic patterns you have studied in this chapter, propose a likely mechanism for this final reaction in the degradation of the amino acid cysteine in mammals.
P13.15: Propose a mechanism for the following carboxylation reaction (EC 6.4.1.4) in the leucine degradation pathway. The complete reaction is dependent on the \(CO_2\)-carrying coenzyme biotin as well as ATP, but assume in your mechanism that the actual carboxylation step occurs with free \(CO_2\) (you don’t need to account for the role played by biotin or ATP).

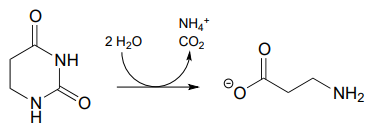
P13.16: Propose a mechanism for the following reaction, which is part of the degradation pathway for the nucleotide uridine.
(
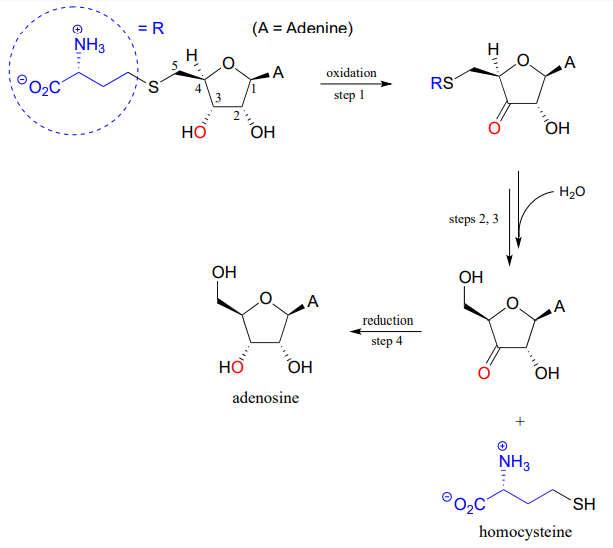
P13.17: Illustrated below is a series of reactions in the degradation pathway for the amino acid methionine. In step 1, an alcohol group on \(C_3\) is oxidized to a ketone, and in step 4 the ketone is reduced back to an alcohol - we will study these reactions in chapter X. In steps 2 and 3, the thiol (homocysteine) is replaced by water - but this does NOT involve a nucleophilic substitution process.
- Draw a likely mechanism for step 2
- Draw a likely mechanism for step 3
- How does the involvement of the redox steps (steps 1 and 4) provide evidence that overall substitution of water for homocysteine is not a nucleophilic substitution?
P13.18: (Challenging!) A recently discovered reaction in the biosynthesis of rhizoxin, a potent virulence factor in the rice-seedling blight fungus Rhizopus microsporus, is illustrated below (Angewandte Chemie 2009, 48, 5001). The reaction takes place at the intersection of two 'modules' of a multi-enzyme complex, and provides an example of a biochemical conjugate addition step that results in the formation of a new carbon-carbon bond (conjugate addition of a carbon nucleophile is referred to as a Michael addition). Draw a likely mechanism.
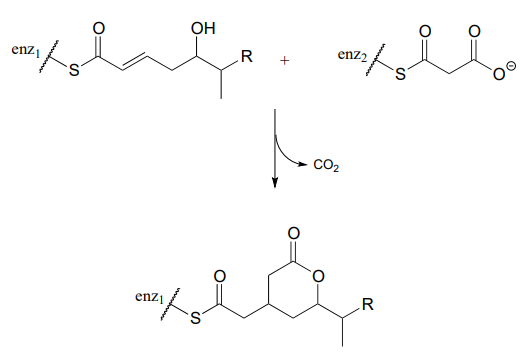
P13.19:
- The 'acetoacetic ester synthesis' is a useful carbon-carbon bond-forming reaction in the laboratory. The reaction mechanism is described as a-carbon deprotonation to form an enolate, followed by SN2 alkylation, ester hydrolysis, and decarboxylation. Below is an example:
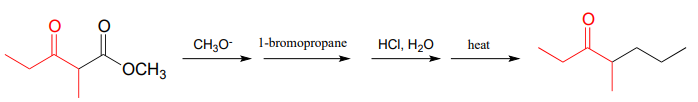
Draw out a reasonable mechanism, taking care to propose reactive intermediates that are appropriate given the basic or acidic conditions present (note that the reaction starts under basic conditions, then is later acidified).
- Suggest starting compounds for the synthesis of 4-phenyl-2-butanone by the acetoacetic ester reaction.
- A very useful ring-forming reaction in laboratory synthesis is called 'Robinson annulation' (Sir Robert Robinson was an English chemist who won the 1947 Nobel Prize in Chemistry, and the term ‘annulation’ comes from the Latin annulus, meaning ‘ring’.) The reaction, which takes place in basic conditions, consists of a conjugate (Michael) addition step, followed by aldol addition and finally dehydration (\(\beta\)-elimination of water). A typical example is shown below, with carbons numbered to help you to follow the course of the reaction.
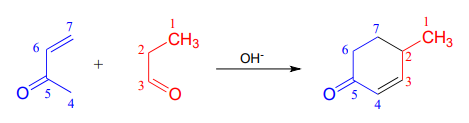
Draw a mechanism for this reaction (when proposing intermediate species, keep in mind that the reaction is occurring in a basic environment, and choose protonation states accordingly).
- Propose starting compounds for the Robinson annulation synthesis of the following product:
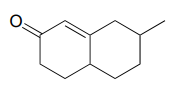
P13.20: The reaction shown below, catalyzed by orotidine monophosphate decarboxylase (EC 4.1.1.23), is one of the most extensively studied enzymatic transformations. It is known to occur without the participation of any coenzymes.
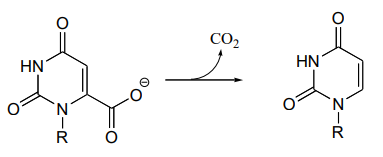
- Look at the reaction closely: what is unique about it?
- In 1997, a paper was published in which the authors predicted, based on theoretical calculations, that this reaction proceeded through a carbene intermediate (carbenes are not covered in this text – you may need to look them up). This was prior to the publication of an x-ray crystal structure. What kind of active site environment does this imply?
- When the crystal structure was published a few years later, we learned that an aspartate residue (predicted to be negatively charged) is positioned very near the substrate carboxylate group, and a lysine residue (predicted to be positively charged) is positioned nearby on the opposite side (see figure below). What roles do you think were predicted for these two active site residues?
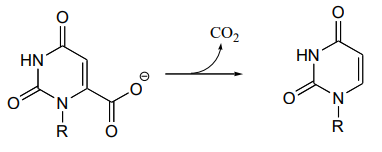
P13.21: In the histidine degradation pathway, histidine undergoes elimination of ammonia to form trans-urocanate. The enzyme catalyzing this reaction (E.C. 4.3.1.3) has been shown to use an unusual 'coenzyme', 4-methylideneimidazole-5-one (MIO), which is formed from the cyclization of an alanine-serine-glycine stretch of the enzyme itself.
A mechanism has been proposed in which the MIO coenzyme plays the role of electron sink, and the intermediate shown below forms.
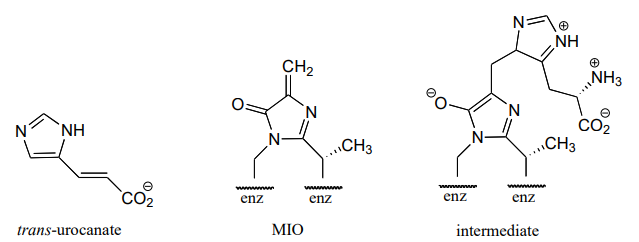
Propose a full mechanism for this reaction according to this information.
P13.22: The product that forms in the reaction between benzaldehyde and acetophenone (along with a catalytic amount of sodium hydroxide) has a 1H-NMR spectrum in which all of the signals are between 7-8 ppm. Give the structure of product.


