7.5: Tertiary structure of proteins
- Page ID
- 425777
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
- Define the tertiary structure of proteins and understand the interactions that form and maintain the tertiary structure.
What is the tertiary structure of proteins?
The secondary structure of proteins represents the folding of portions of the polypeptide held primarily by hydrogen bonding between \(\ce{C=O}\) and \(\ce{N-H}\) groups of the polymer backbone. The overall protein chain also folds in a configuration that causes more stabilizing interaction primarily between the groups on the side chains (\(\ce{-R}\)) and the between side chain groups and the groups on the polymer backbone.
The three-dimensional arrangement of all the atoms of a single polypeptide chain in space, held together by stabilizing interactions between the side chain and the backbone groups, is called the tertiary structure of proteins.
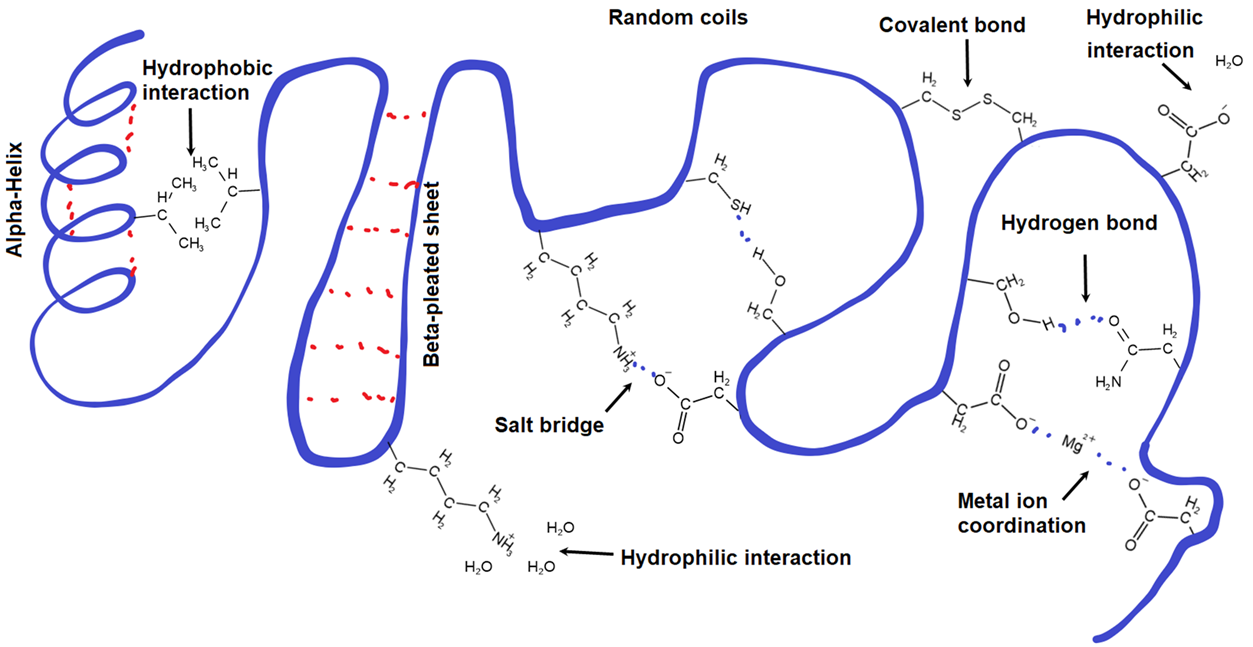
The interactions stabilizing the tertiary structure include disulfide linkage, salt bridge, coordinate bonds with metal ions, hydrogen bonding, and hydrophobic interaction, as shown in Figure \(\PageIndex{1}\). and explained below.
Disulfide linkage
When thiol (\(\ce{-SH}\)) groups of two cysteine residues come close to each other in a folded protein chain, they can make disulfide linkage ((\(\ce{-S-S{-}}\)), i.e., the only covalent bond, besides peptide bond, holding amino acid residues together in proteins.
\(\ce{R-SH + HS-R' ->[Oxidation] R-S-S-R + 2H^{+} + 2e^{-}}\)
Salt bridge
The side chain of acidic amino acids, e.g., Asp, loses protons and becomes an anion, and that of basic amino acids, e.g., Lys, gains a proton and becomes a cation. When the opposite ions come close to each other, they make an ionic bond, like that in salts, called salt-bridge, e.g., \(\ce{...-CH2-COO^{-} ^{+}NH3-(CH2)4-...}\)) can form between Asp and Lys residues. This is an ionic bond holding opposite charges together.
Coordinate bond
Some metals can act as Lewis acids and make a coordinate covalent bond by accepting a lone pair of electrons from Lewis bases. Oxygen and nitrogen atoms in proteins have lone pairs of electrons and can act as Lewis bases. For example, oxygen atoms of two carboxylate \(\ce{-COO^{-}}\) groups can make coordinate covalent bonds with \(\ce{Mg^{2+}}\), resulting in a coordinate covalent linkage (\(\ce{-COO^{-}\!\!\!\!\!->\!\!Mg^{2+}\!\!\!\!\!\!\!\!<-\!\!\!\!\! ^{-}OOC{-}}\)) between the amino acid residues.
Hydrogen bonding
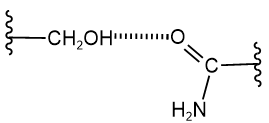
Hydrogen atoms bonded to oxygen or nitrogen atoms, e.g., in \(\ce{-OH}\) and \(\ce{-NH2}\), can make hydrogen bonds with any oxygen or nitrogen atom. These bonds could be between side chains, backbone groups, or side chains and backbone groups. For example, \(\ce{-OH}\) in the side chain of serine can make hydrogen with the carbonyl oxygen of the amide group (\(\ce{-CONH2}\)) of asparagine residue, as shown below.
Hydrophobic interactions
As shown below, amino acids with aliphatic or aromatic hydrophobic side chains, e.g., valine and phenylalanine, establish hydrophobic interaction.
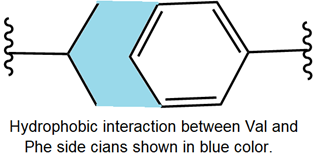
Hydrophobic interaction is London dispersion forces between nonpolar groups. The physiological medium is predominantly water. As a nonpolar liquid like oil, when mixed with a polar liquid like water, the oil drops merge, expelling water. Similarly, hydrophobic groups in proteins come close to each other due to the hydrophobic interaction and tend to stay in the interior of the protein, expelling water away from those regions. Ionic and polar compounds dissolve in water. Similarly, ionic and polar groups on the side chains stay on the proteins' outer surfaces, interacting more with water.
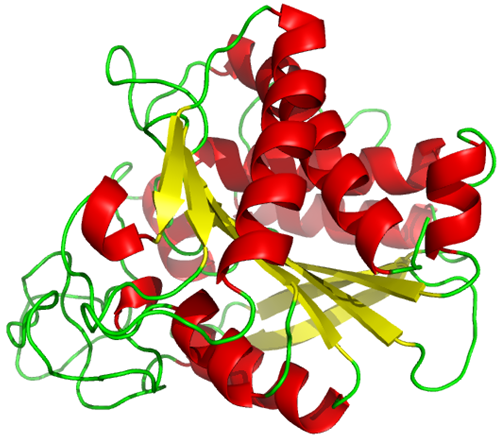
The tertiary structure of proteins may be predominantly \(\alpha\)-helix, \(\beta\)-pleated sheets, or random coil, but often it is a mix of these, for example, Carboxypeptidase A from bovine pancreas, shown in Figure \(\PageIndex{2}\).