18.3: Amino Acids
- Page ID
- 86302
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- To recognize amino acids and classify them based on the characteristics of their side chains.
- Identify which amino acids are chiral.
The proteins in all living species, from bacteria to humans, are polymers constructed from the same set of 20 amino acids. Humans can synthesize only about half of the needed amino acids; the remainder must be obtained from the diet and are known as essential amino acids. However, two additional amino acids have been found in limited quantities in proteins: Selenocysteine was discovered in 1986, while pyrrolysine was discovered in 2002.
Amino Acid Structure
Every amino acid contains an amino group, (–NH2), a carboxyl group, (–COOH), and a side chain or R group, which are all attached to the alpha (\(\alpha\)-) carbon (the one directly bonded to the carboxyl functional group). Therefore, amino acids are commonly called alpha-amino (\(\alpha\)-amino) acids. Figure \(\PageIndex{1}\) below shows the structure of a generic \(\alpha\)-amino acid.
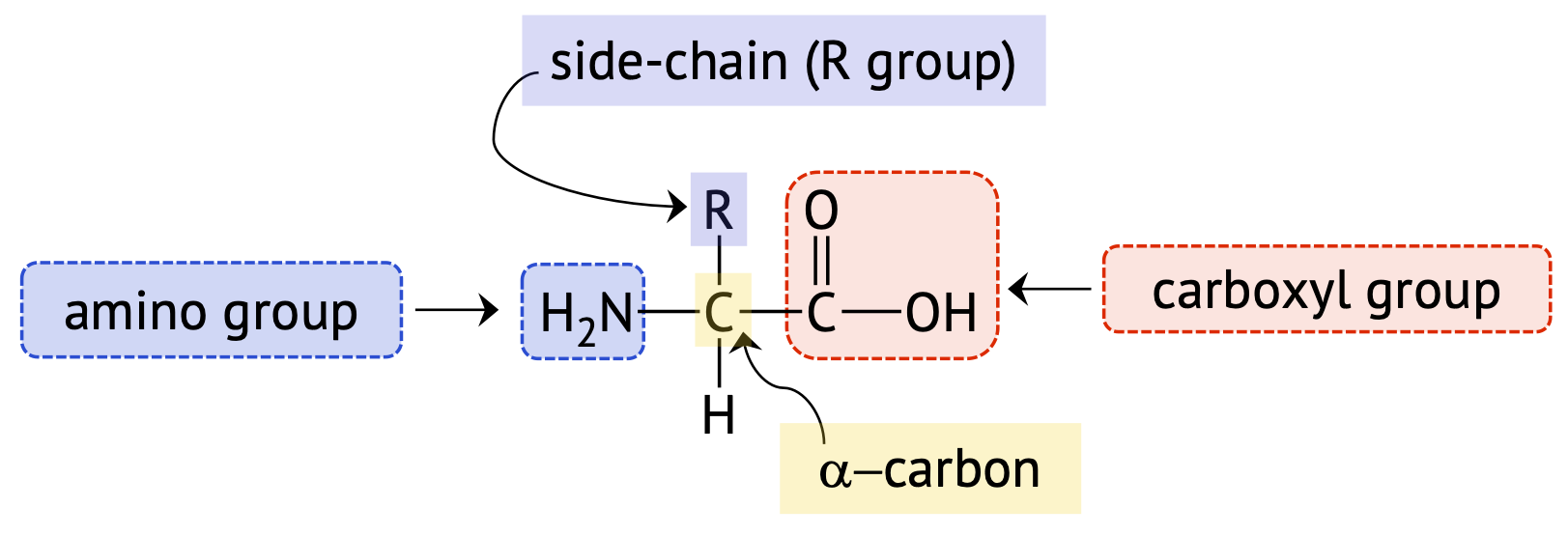
Amino Acid Side Chains
Amino acid side chains or R groups can range from a single hydrogen atom (as in glycine), to a simple hydrocarbon chain, to a hydrocarbon containing a functional group. Each R group has differences in size, shape, solubility, and ionization properties, which contributes to the unique properties of an individual amino acid, and can have an effect on the overall structure and function of a protein.
Table \(\PageIndex{1}\) below lists the 20 common amino acids along with their names, their three- and one-letter codes, structures, and distinctive features. The three-letter codes are generally the first three letters of the amino acid name except in a few cases, such as isoleucine (Ile) and tryptophan (Trp). Similarly, the one-letter code is usually the first letter in the amino acid name, but where the letter is not unique, a letter that is phonetically similar the amino acid name is used: F for Fenylalanine, R for aRginine, and W for tWiptophan. This table also groups the amino acids according to whether the side chain at neutral pH is nonpolar, polar uncharged, positively charged, or negatively charged.
| Common Name | Three-letter (one-letter) Code | Systematic (IUPAC) Name | Structural Formula (at pH 6) | Isoelectric Point (pI) | Distinctive Feature |
|---|---|---|---|---|---|
| Amino acids with a nonpolar R group | |||||
| glycine | Gly (G) | aminoethanoic acid | 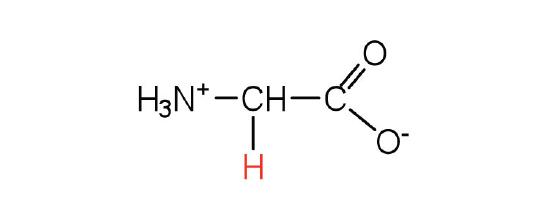 |
6.0 | the only amino acid lacking a chiral carbon |
| alanine | Ala (A) | 2-aminopropanoic acid | 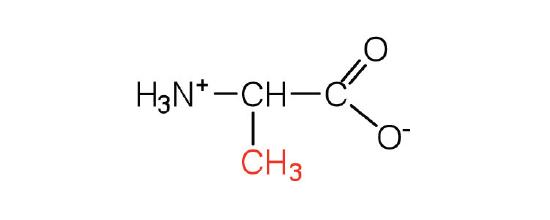 |
6.0 | a methyl group, it is the second smallest side chain |
| valine | Val (V) | 2-amino-3-methylbutanoic acid | 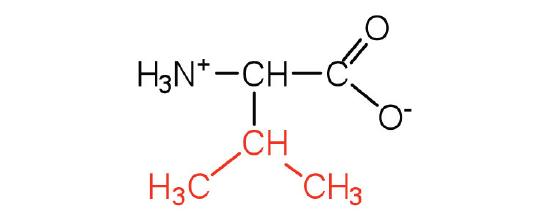 |
6.0 | a branched-chain amino acid |
| leucine | Leu (L) | 2-amino-4-methylpentanoic acid | 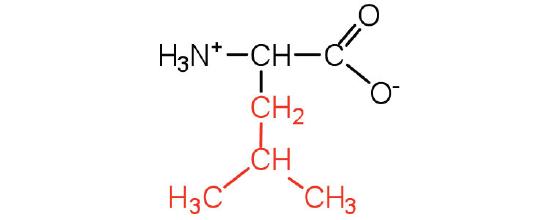 |
6.0 | a branched-chain amino acid |
| isoleucine | Ile (I) | 2-amino-3-methylpentanoic acid | 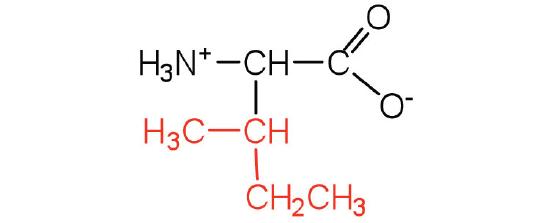 |
6.0 | an essential amino acid because most animals cannot synthesize branched-chain amino acids |
| phenylalanine | Phe (F) | 2-amino-3-phenylpropanoic acid | 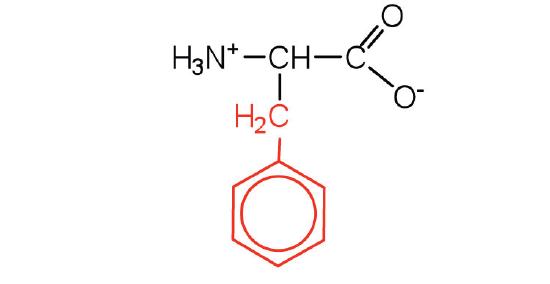 |
5.5 | also classified as an aromatic amino acid |
| tryptophan | Trp (W) | 2-Amino-3-(1H-indol-3-yl)-propanoic acid | 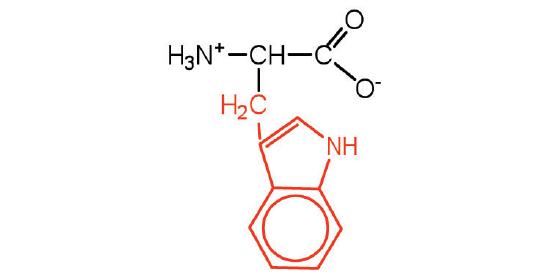 |
5.9 | also classified as an aromatic amino acid |
| methionine | Met (M) | 2-amino-4-(methylthio)butanoic acid | 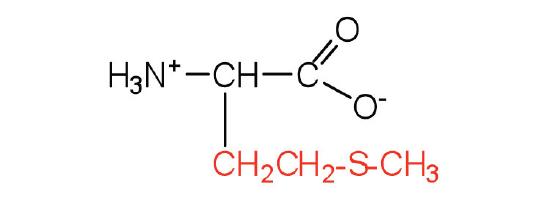 |
5.7 | side chain functions as a methyl group donor |
| proline | Pro (P) | pyrrolidine-2-carboxylic acid | 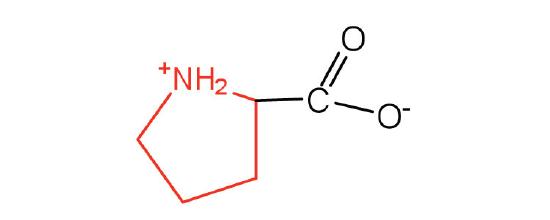 |
6.3 | contains a secondary amine group; referred to as an α-imino acid |
| Amino acids with a polar but neutral R group | |||||
| serine | Ser (S) | 2-amino-3-hydroxypropanoic acid | 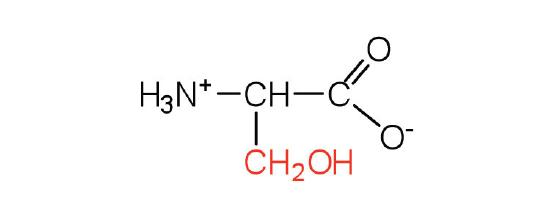 |
5.7 | found at the active site of many enzymes |
| threonine | Thr (T) | 2-amino-3-hydroxybutanoic acid | 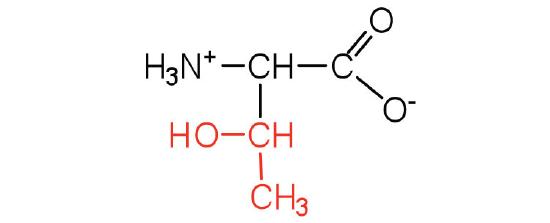 |
5.6 | named for its similarity to the sugar threose |
| cysteine | Cys (C) | 2-amino-3-mercaptopropanoic acid | 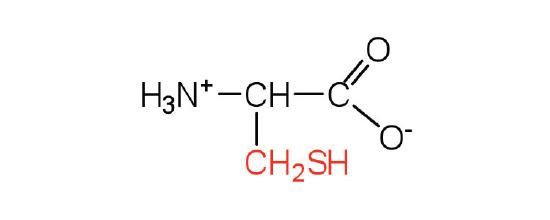 |
5.0 | oxidation of two cysteine molecules yields cystine |
| tyrosine | Tyr (Y) | 2-amino-3-(4-hydroxyphenyl)-propanoic acid | 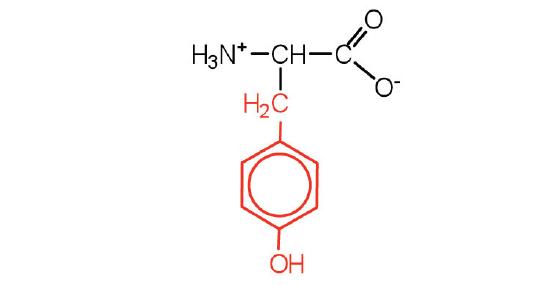 |
5.7 | also classified as an aromatic amino acid |
| asparagine | Asn (N) | 2-amino-3-carbamoylpropanoic acid | 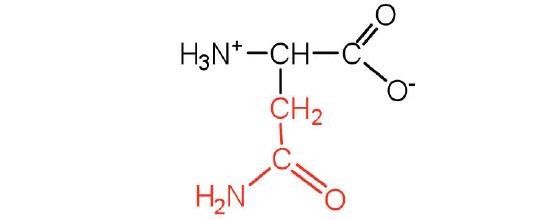 |
5.4 | the amide of aspartic acid |
| glutamine | Gln (Q) | 2-amino-4-carbamoylbutanoic acid | 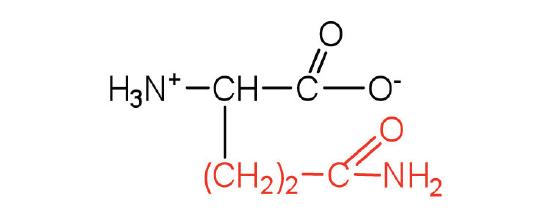 |
5.7 | the amide of glutamic acid |
| Amino acids with a negatively charged R group | |||||
| aspartic acid | Asp (D) | 2-aminobutanedioic acid | 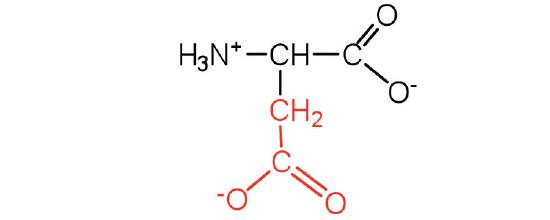 |
3.0 | carboxyl groups are ionized at physiological pH; also known as aspartate |
| glutamic acid | Glu (E) | 2-aminopentanedioic acid | 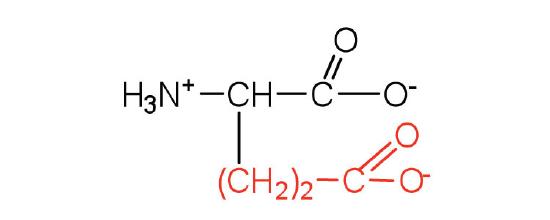 |
3.2 | carboxyl groups are ionized at physiological pH; also known as glutamate |
| Amino acids with a positively charged R group | |||||
| histidine | His (H) | 2-Amino-3-(1H-imidazol-4-yl)-propanoic acid | 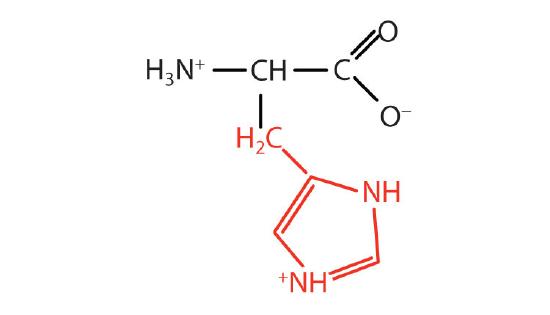 |
7.6 | the only amino acid whose R group has a pKa (6.0) near physiological pH |
| lysine | Lys (K) | 2,6-diaminohexanoic acid | 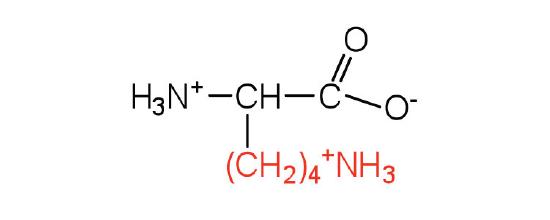 |
9.7 | is somewhat amphipathic due to the long hydrocarbon tail and positively charged amino group on the \(\epsilon\) carbon |
| arginine | Arg (R) | 2-amino-5-guanidinopentanoic acid | 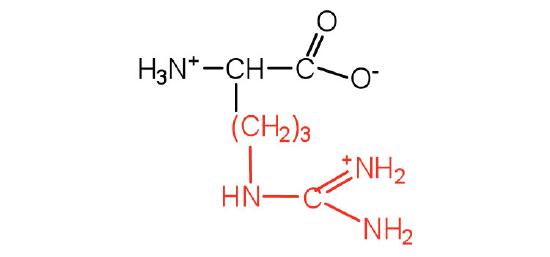 |
10.8 | almost as strong a base as sodium hydroxide |
The first amino acid to be isolated was asparagine in 1806. It was obtained from protein found in asparagus juice (hence the name). Glycine, the major amino acid found in gelatin, was named for its sweet taste (Greek glykys, meaning “sweet”). Glutamic acid is named as such because it was first isolated from gluten. The crystalline salt of glutamic acid is called monosodium glutamate (MSG), which is naturally occuring in some foods but is also added to as a savory or "umami" flavor enhancer.
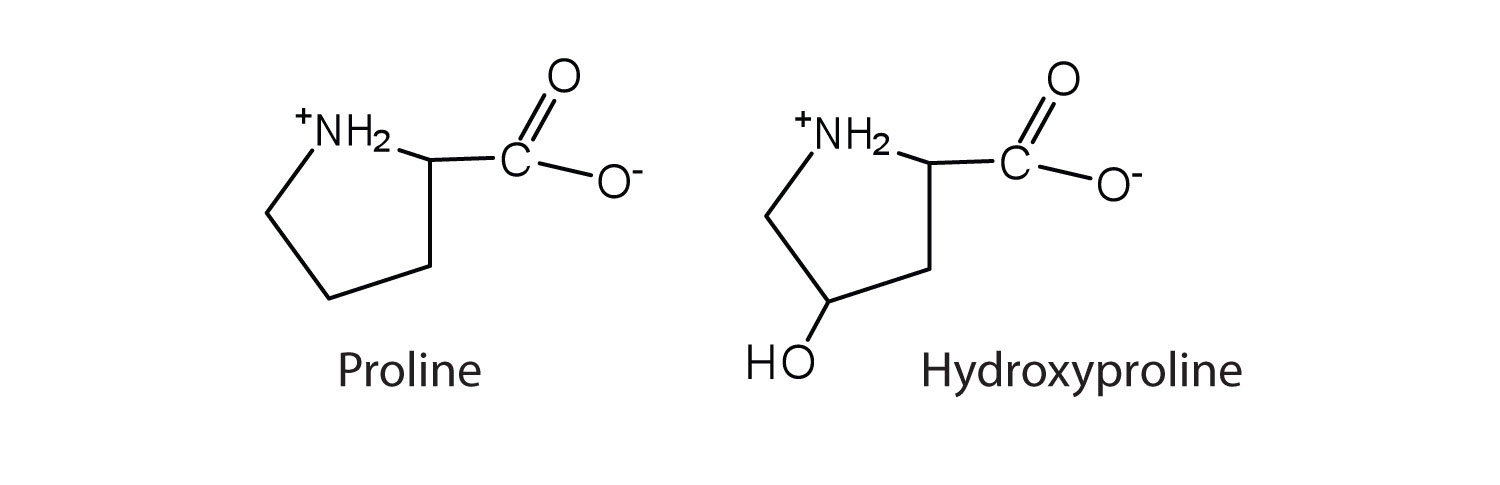
In some cases an amino acid found in a protein is actually a derivative of one of the common 20 amino acids (one such derivative is hydroxyproline). The modification of proline occurs after the amino acid has been assembled into a protein.
Chirality of Amino Acids
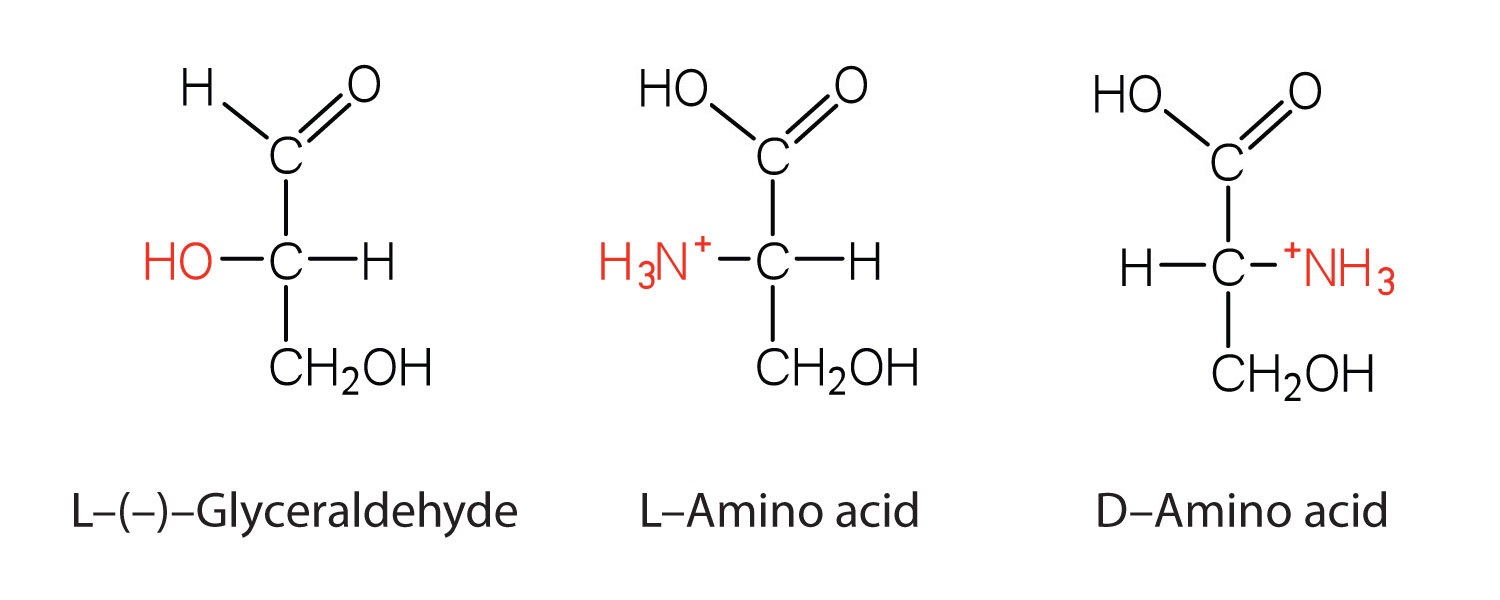
Notice in Table \(\PageIndex{1}\) that glycine is the only amino acid whose (\alpha\)-carbon is not chiral, in other words the molecule and the mirror-image of glycine are identical. All other amino acids have two forms that are mirror images of each other, they are enantiomers. As you can see in the figure below, the "left-handed" form of the molecule is known as the L-amino acid and the "right-handed" form is the D-amino acid.
Summary
Amino acids can be classified based on the characteristics of their distinctive side chains as nonpolar, polar but uncharged, negatively charged, or positively charged. The amino acids found in proteins are L-amino acids.


