7.3: Complex II
- Page ID
- 190047
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Complex II is another group of proteins that serves as a second entry point into the electron transport chain, which is involved in the additional production of ATP to power cellular processes. The electron transport chain releases minute amounts of energy with each electron transfer, and the transport is coupled to the pumping of protons across the mitochondrial membrane. Eventually, the electrons are delivered to molecular oxygen, which is reduced to water, in Complex IV.
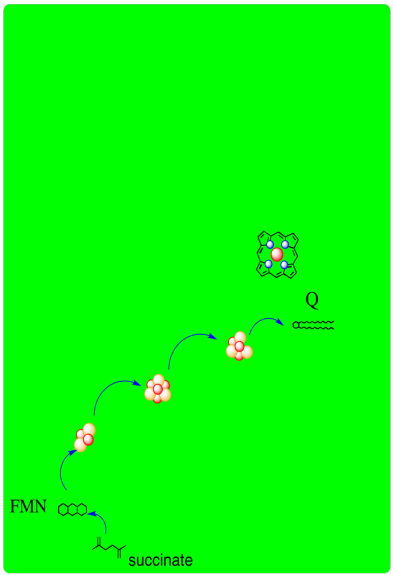
A schematic of Complex II is provided below. It shows electrons provided by succinate traveling first to FAD and then along a series of iron sulfur clusters to a ubiquinone acceptor.
- Complex II, like Complex I, is an entry point into the electron transport chain.
- In Complex II, two electrons are delivered from succinate.
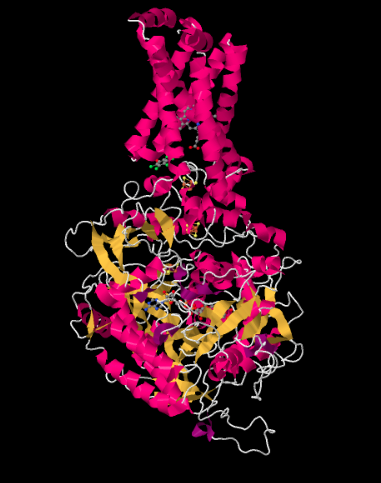
The X-ray crystal structure of Complex II is shown below. It is depicted in cartoon form to highlight the structure, with pink helices, yellow sheets and white loops. Just as in the structure of Complex I, the structure here is shown oriented so that the inner mitochondrial membrane would stretch across the top of the picture; if it were shown, we would see it in cross section, from side to side. Once again, there is a hydrophobic domain in the upper part of the complex that would be embedded in that membrane. Below that lies a much larger hydrophilic domain, which would extend into the mitochondrial matirx.
Complex II actually plays a part in another biochemical process, the citric acid cycle. One of the steps of the citric acid cycle is the oxidation of succinate to fumarate. It is during this oxidation that additional electrons are introduced into the electron transport chain via Complex II. At the same time, two protons are delivered from succinate as well. The loss of two protons and two electrons from succinate corresponds to a formal loss of H2 (formal, meaning a molecule of H2 is not actually formed, but the pieces are equivalent to a full H2 molecule). A double bond forms in the new fumarate molecule, which is said to be "dehydrogenated" with respect to the original succinate. Complex II is the enzyme that carries out this transformation in the citric acid cycle, and so the complex is also called "succinate dehydrogenase".
- Complex II is actually an integral part of the TCA cycle, which takes place in the mitochondrial matrix.
- Conversion of succinate to fumarate is a two-electron, two-proton process.
Shown below, the proteins are stripped out of the picture to reveal some of the working parts inside. Having already seen Complex I, you will probably recognize a few iron sulfur clusters, shown again in red and yellow. At the bottom of the picture, close to the matrix, you can see a flavenoid structure, a little like the FMN you saw in Complex I. At the top, actually buried within the hydrophobic portion (you can just see the edge of it if you look between two of the pink helices in the picture above), is an iron heme complex; it's a little like the active site of hemoglobin. You can see the iron atom, coded in red, and the four nitrogen atoms around it, coloured blue. However, it is not clear what role this heme actually plays in electron transport through Complex II.
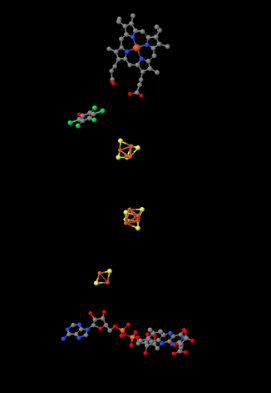
In between the highest iron sulfur cluster and the iron heme is a molecule of pentafluorophenol. That's not supposed to be there in the cell. It's an inhibitor; the researchers added it to hold the place of a ubiquinone while they were growing the crystal that they would analyse to get this structure. There is actually a second inhibitor in the structure, an oxaloacetate, which holds the place of the succinate molecule that would deliver electrons to the complex. However, the oxaloacetate is not clearly visible because it's partly obscured by the flavenoid at the bottom of the picture.
- Like in Complex I, electrons eventually arrive at ubiquinone in Complex II.
- Ubiquinone is a mobile, lipid-soluble, two-electron carrier.
Complex II is unique in a another important way. Unlike Complexes I, III, and IV, it takes part in the electron transport chain but does not actively pump protons across the inner mitochondrial membrane. Instead, the electrons that it harvests contribute to building the proton gradient once they arrive at Complex III (and later at Complex IV).
- Complex II does not pump protons directly.
- Complex II does send two protons on to Complex III in the form of the reduced ubiquinol.
In terms of how those electrons make their arrival, the picture is somewhat similar to Complex I. Succinate is a hydride donor in this reaction, so it is donating two electrons. As in Complex I, a series of FeS clusters will eventually relay the electrons, so we have a matching problem. To step down from a two-electron process to a one-electron process, another flavenoid, FAD, acts as an intermediary. Like the related FMN, FAD can accept either one or two electrons at a time, and its reduced form, FADH2, can donate either one or two electrons at a time.
- Just like in Complex I, an adapter is needed to convert the two-electron process of succinate oxidation to the one-electron process of iron reduction.
- The adapter is another flavenoid structure, FAD, which is very similar to the FMN used in Complex I.
After the electrons have arrived and are stored in the form of FADH2, they are passed along, one at a time, through a series of three FeS clusters. Just as in Complex I, these carriers are all bound to the proteins, so they are held in one place and do not move. The outer sphere mechanism by which the electrons are transferred can only operate over a short distance, from one electron carrier to the next.
Complex II also contains a structure we did not see in Complex I: a cytochrome. A cytochromes is a protein that contains a heme complex of iron. They look a little bit like hemoglobin, with which you are undoubtedly familiar by now. As in the FeS clusters, the cytochrome can access either the Fe(II) or the Fe(III) state. Like the FeS clusters, they just accept one electron at a time. We will find out that some cytochromes are mobile, but this one is not. It stays where it is and waits for a ubiquinone to bind nearby.
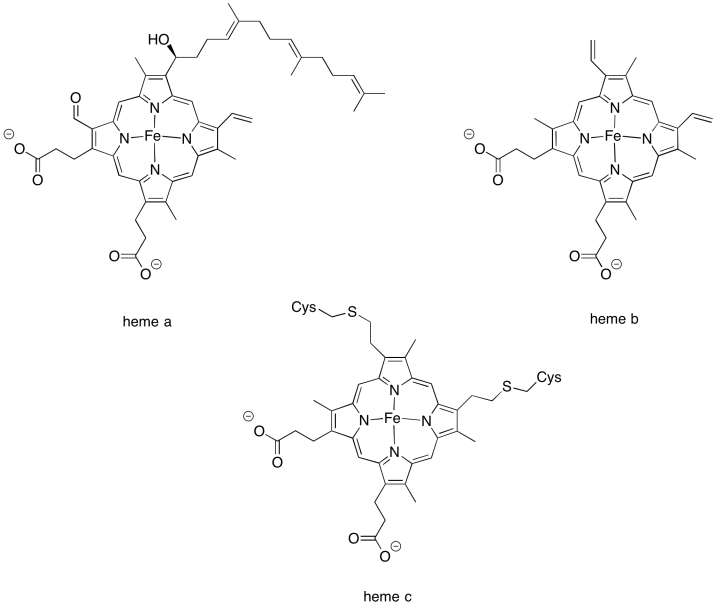
There are few different variations in the heme family that are commonly encountered in biology. Heme b is a common one and has the simplest structure. Heme a contains a hydrocarbon tail as well as a formyl group (-HC=O) attached to its central porphyrin ring. Heme c is covalently bound to the surrounding protein via cysteine residues.
Exercise \(\PageIndex{1}\)
Look again at the X-ray crystal structure of the ligand in Complex II. Which type of heme is present?
- Answer
-
Heme b.
Exercise \(\PageIndex{2}\)
The basic porphyrin structure at the centre of all the hemes is a macrocycle with the formula (C4H2NCH)4. Draw this structure.
- Answer
-
A porphyrin contains four pyrrole rings (five-membered, aromatic ring containing a nitrogen) arranged to form a 16-membered macrocycle.
The exact role of the cytochrome in Complex II is unclear. Although cytochromes form part of the electron transport chain in Complexes III and IV, it might not do so in Complex II. Another possible role is storage of an electron (which does arrive through the electron transport chain). When the ubiquinone binds nearby, the cytochrome may release its electron to the ubiquinone. A second electron would arrive directly from the nearby FeS cluster. That mechanism may allow for more rapid reduction of ubiquinone to ubiquinol, so that it spends less time in the semiquinone stage. As a result, the radical species would be less likely to move away before being fully reduced. Radical species can cause damage, so limiting the movement of radical species through the cell is important.
- The role of the heme group in Complex I is not clear.
- It has been suggested that the heme group may act like a capacitor. It may store one of the two electrons needed to reduce ubiquinone to ubiquinol.
- Storage of an extra electron could result in more rapid reduction of ubiquinone to ubiquinol.
- Rapid, two-electron reduction limits the lifetime of potentially damaging radical intermediates in the cell.
Once two protons and two electrons are delivered to ubiquinone, the resulting ubiquinol travels on to Complex III for the next step in oxidative phosphorylation.
Exercise \(\PageIndex{3}\)
Provide a mechanism for the reduction of FAD by succinate.
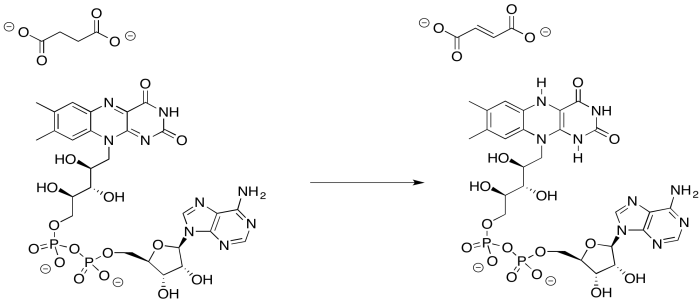
- Answer
-
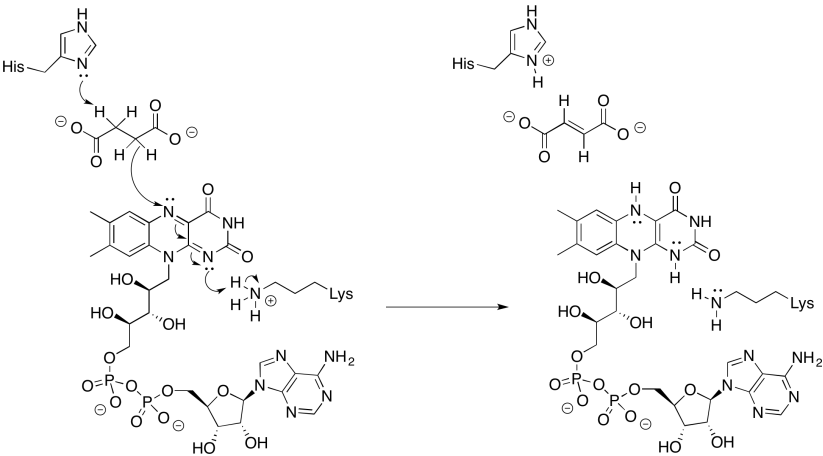
Exercise \(\PageIndex{4}\)
It's difficult to measure the reduction potential of an individual site within a protein. However, researchers have been able to estimate these values by measuring EPR spectra under various conditions. Assuming the reduction potentials below, draw a reaction progress diagram for transport of an electron all the way from succinate to ubiquinone.
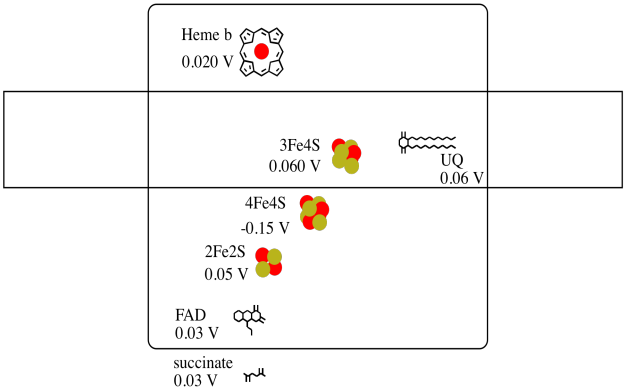
- Answer
-
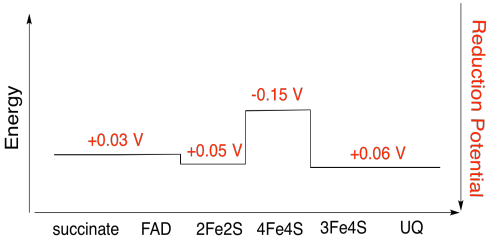
Exercise \(\PageIndex{5}\)
Using the values in the figure above, calculate the energy change when an electron is transferred from the 4Fe4S cluster to the 3Fe4S cluster.
- Answer
-
Assuming the reduction potentials are:
4Fe4S(ox) + e- → 4Fe4S(red) Eored = -0.15 V
3Fe4S(ox) + e- → 3Fe4S(red) Eored = 0.06 V
Then the potential difference for the reaction, ΔEo = 0.06 - (-0.15) V = 0.21 V.
The Faraday relation ΔG = - n F ΔEo gives
ΔG = - 1 x 96,485 J V-1 mol-1 x 0.21 V = 20,262 J mol-1 = 20 kJ mol-1
Exercise \(\PageIndex{6}\)
Assuming all iron is present as Fe(III), calculate the overall charges on:
a) 2Fe2S b) 3Fe4S c) 4Fe4S
- Answer a
-
a) Iron charges: 2 x Fe(III) = 6+
Ligand charges: 2 sulfides = 2 x 2- = 4- ; 4 cysteines = 4 x 1- = 4- ; total = 8-
Overall: 2-
- Answer b
-
b) Iron charges: 3 x Fe(III) = 9+
Ligand charges: 4 sulfides = 4 x 2- = 8- ; 3 cysteines = 3 x 1- = 3- ; total = 11-
Overall: 2-
- Answer c
-
c) Iron charges: 4 x Fe(III) = 12+
Ligand charges: 4 sulfides = 4 x 2- = 8- ; 4 cysteines = 4 x 1- = 4- ; total = 12-
Overall: 0
X-ray crystal structures: Zhou, Q., Zhai, Y., Lou, J., Liu, M., Pang, X., Sun, F. Thiabendazole inhibits ubiquinone reduction activity of mitochondrial respiratory complex II via a water molecule mediated binding feature. Protein Cell 2011, 2, 531-542. Images obtained via RCSB Protein Data Bank (3SFD).


