6.6: Covalent Modification
- Page ID
- 189958
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Reversible inhibitors are extremely important in regulating enzyme activity. They can turn enzymes on or off, acting as activators or inhibitors, respectively. In addition, enzymes can be regulated via covalent modification or post-translational modification. That means that, after the enzyme has been assembled in the cell, its structure can be modified further by adding special groups to specific locations. In the case of regulation, these groups are added reversibly. Even though the group is added covalently -- it is bonded to the protein -- a reaction path exists for the removal of the group again.
There are three examples of these modifications that we will look at here. In each case, the behaviour of the protein is modified because of changes in the intermolecular attractions within the protein (or between the protein and another molecule).
Phosphorylation is a very common modification. In phosphorylation, a phosphate group is attached to an amino acid side chain. The most commonly phosphorylated side chain is a serine. Tyrosine is often phosphorylated, too. Those sites are preferred because of the strong P-O bond formed during the reaction.
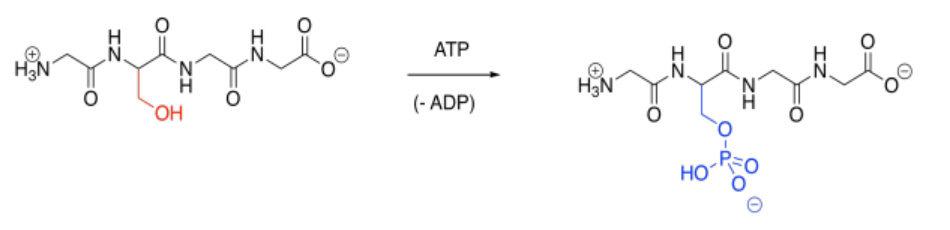
Phosphorylation is typically carried out under the control of another enzyme called a kinase. That's right, one enzyme will bind another, tying a phosphate group onto it before releasing it again. A phosphate group can be removed again via another enzyme called a phosphatase. The fact that these modifications are carried out by specific enzymes helps to explain their specificity. A particular kinase may bind to its target protein in a well-defined position, phosphorylating it only at that position (although some kinases may be less selective).
Of course, the key result of phosphorylation is that a neutral serine is suddenly masked by an anionic phosphate group. That negative charge alters intermlecular attractions quite starkly, because suddenly attractive and repulsive forces pop up where there were none before.
Acetylation is also quite common. Acetylation is the addition of an acetyl or ethanoyl group; usually, the group is added to a lysine. The forward reaction is driven by the strong amide bond that results.
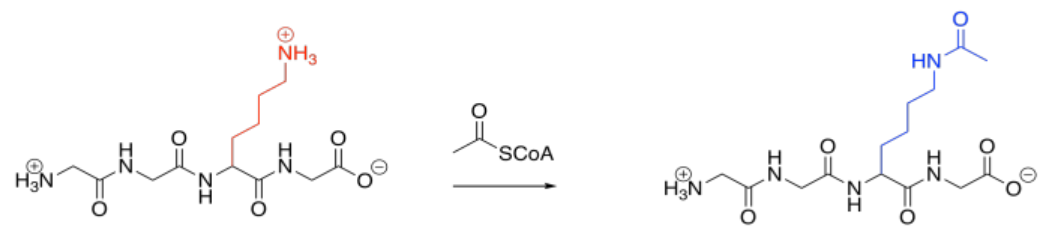
Acetylation is carried out by an acetylase or an acyltransferase. Like phosphatases, these are enzymes that bind their target proteins in order to modify their structures. Because lysines are normally positively charged at biological pH, acetylation results in the sudden disappearance of charged species because the are masked with neutral acyl groups. Intermolecular attractions can be dramatically affected as a result.
Prenylation is the addition of a hydrocarbon side chain, most often to a cysteine side chain. Sulfur is a particularly good nucleophile for carbon chains of this sort, enhancing selectivity for cysteine. As in the other cases, however, the reaction is carried out under the control of an enzyme (a geranyltrasnferase, for example, or a farnesyltransferase), and so other side chains might be targeted, instead.
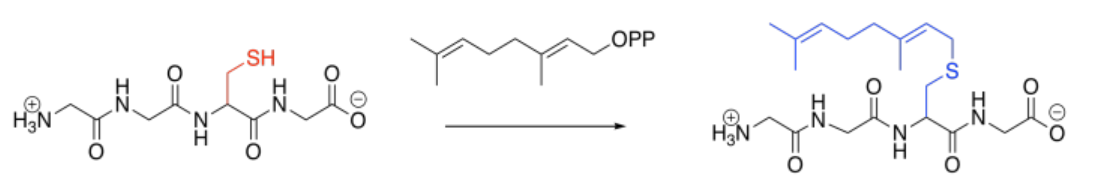
Those enzyme types highlight two common groups that are added in these cases: the geranyl group and the related farnesyl group. These groups are structurally related, and are both in the terpenoid family of natural products. Terpenes are based on five-carbon building blocks, which you can easily trace out in the geranyl and farnesyl structures.
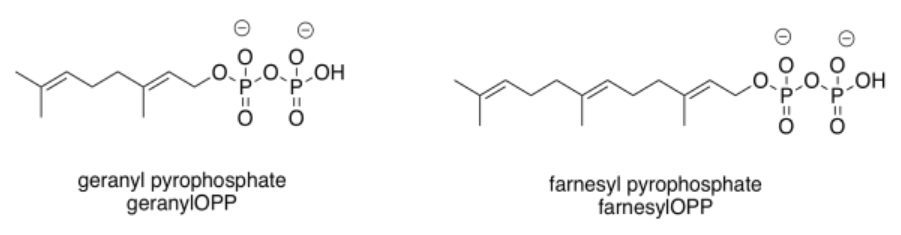
Changes in intermolecular attractions are more subtle in these cases, because prenylation is not accompanied by a full change in the charge of a side chain as it was in the other cases. Instead, addition of this group enhances the hydrophobicity of that part of the protein. A hydrophobic group is one that is completely incapable of interacting with water molecules, lacking ions, dipoles, or hydrogen bonding sites. As a result, it tends to get displaced or pushed out of the way by surrounding water molecules. That "pushing away" by water has the net effect of piling hydrophobic groups together.
What we tend to get, then, is hydrophobic groups sticking to each other, even though their attraction for each other (London forces) is actually pretty weak. Of course, because London forces are relatively weak, the amount of London forces, or the relative area of a molecule that can interact with another via London forces, becomes very important. Geranyl and farnesyl groups are moderately long, and so they exert significant amounts of London forces. As a result, two prenylated parts of a protein might stick together, or one newly prenylated part might huddle together with some hydrophobic side chains, such as valines, leucines, and isoleucines. Also, prenylation sometimes causes a protein that was formerly more water-soluble to migrate over to the cell membrane, where it could interact with the lipid bilayer. So, prenylation, despite causing seemingly modest changes in intermolecular attractions, can elicit dramatic changes in behaviour in the protein.
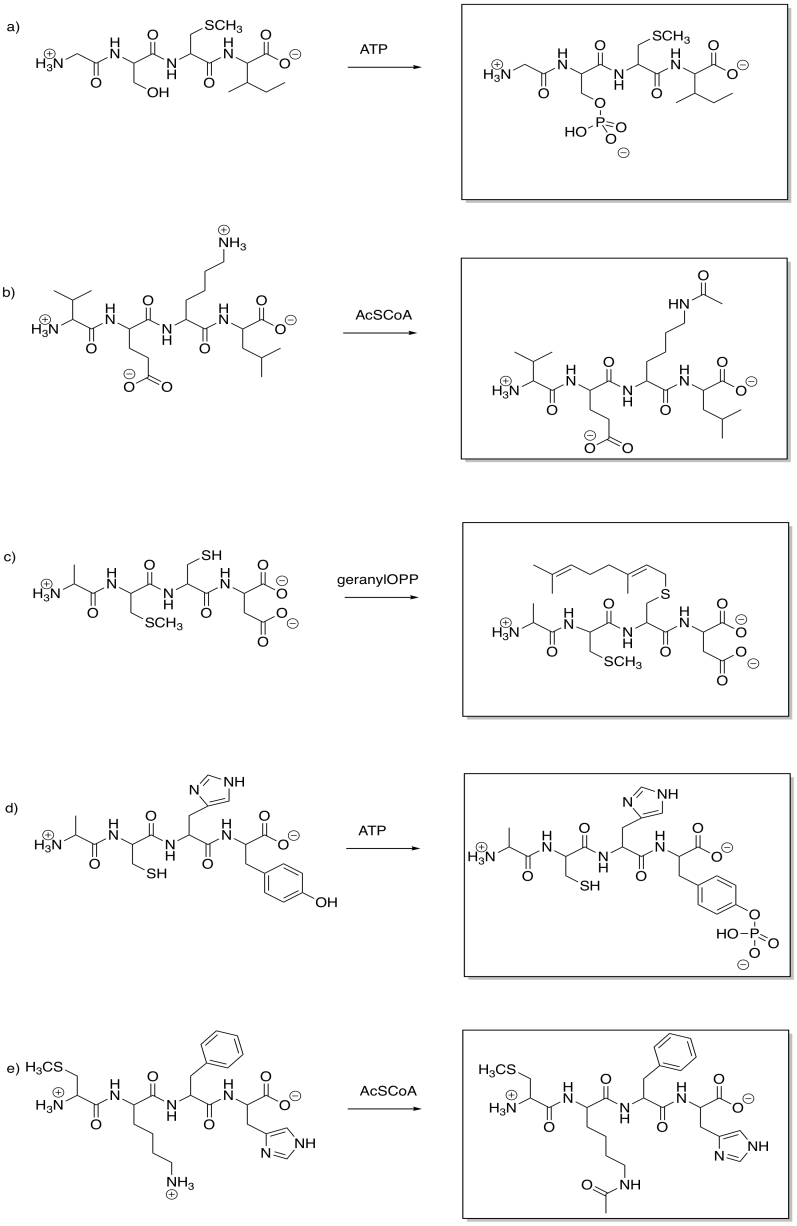
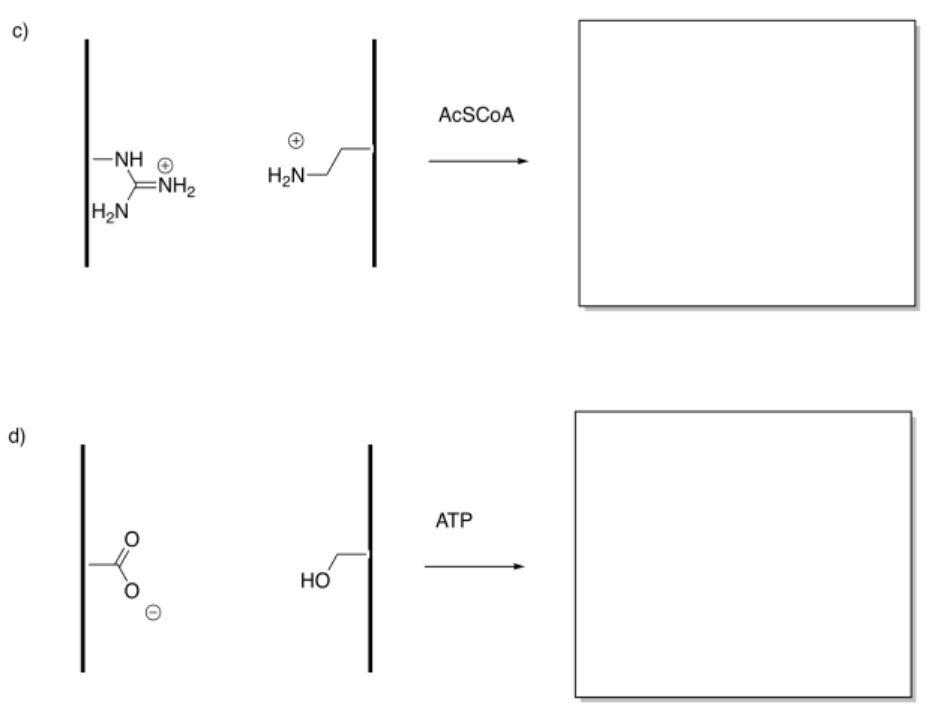
Show the products of the following modifications.
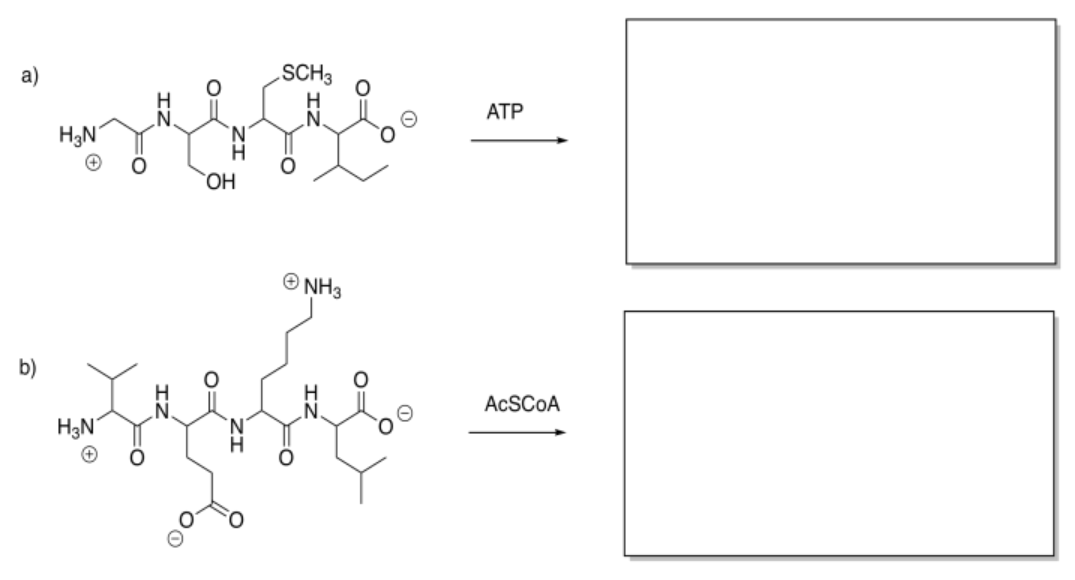
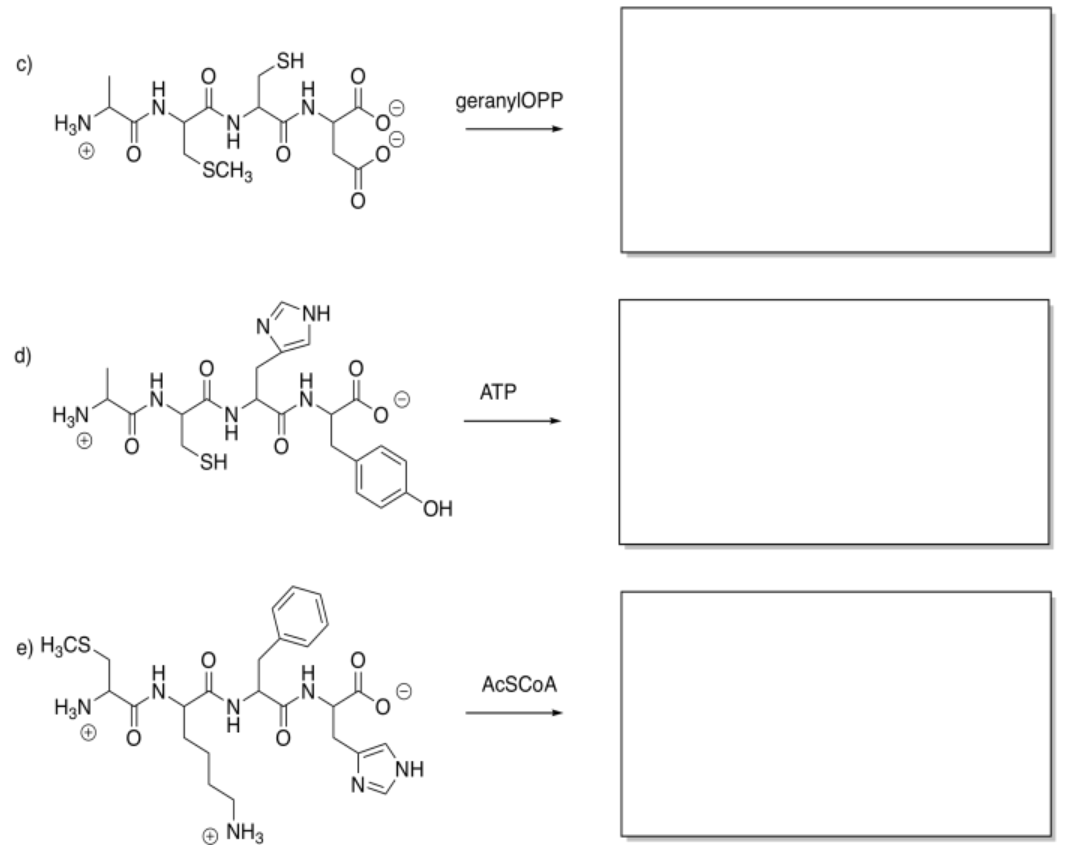
Answer-
Indicate whether attraction between the strands would increase or decrease after modification.
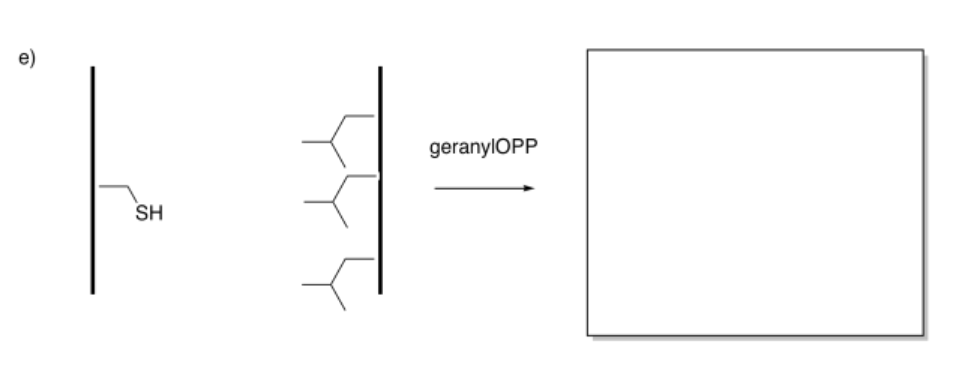
Answer-

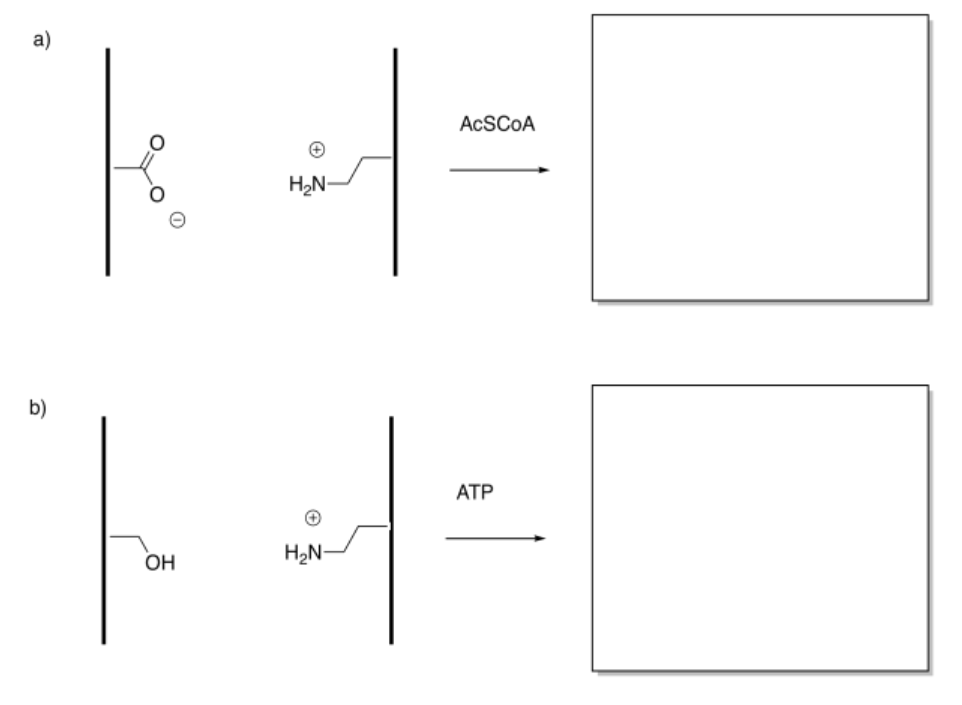
In the following drawings, the object's interaction with one of the surfaces will change upon modification. This change in interaction will result in a net shift of the object to the left or to the right. The shift may result because of attractive or repulsive forces. Indicate the direction of shift in each case.
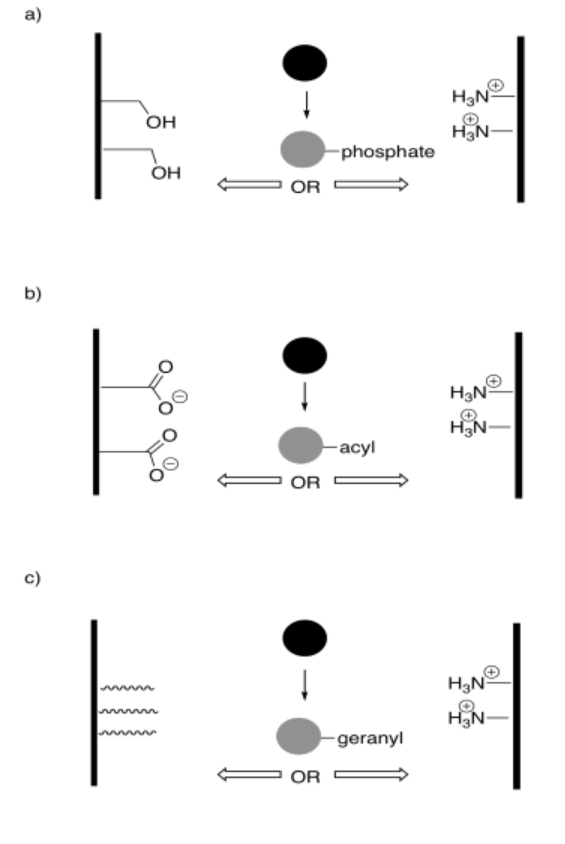
Answer-
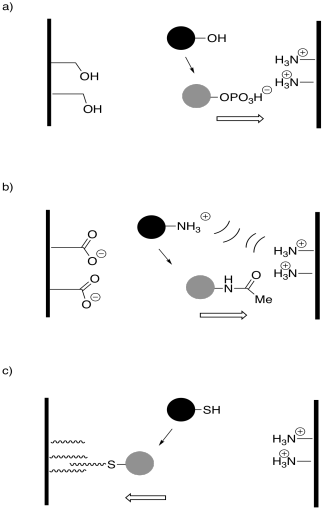

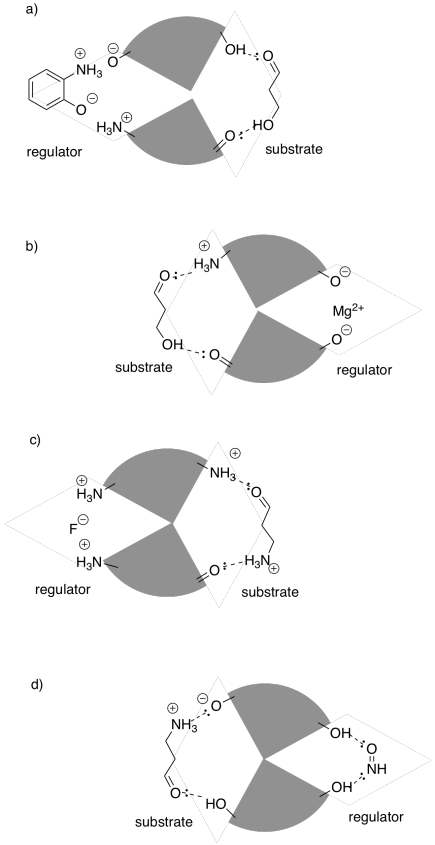
In the following drawings, the active site is closed. Show how the regulator will bind, causing the active site to open, and also show how the substrate will bind.
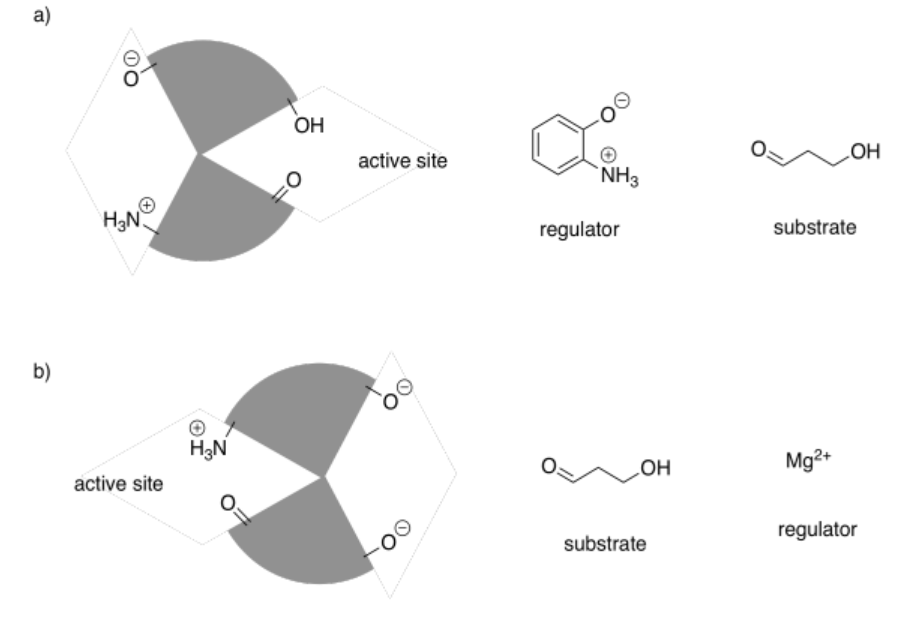
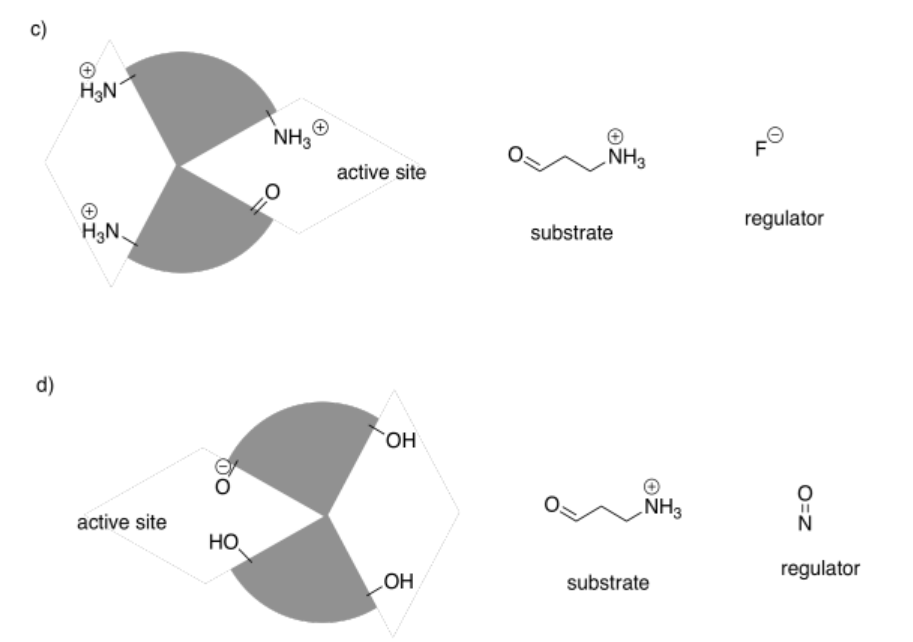
Answer-
The following drawings depict an allosteric site that will be modified on the left and a binding site on the right. Imagine the middle bar is a see-saw that can tilt left or right. In each case, decide whether the modification will lead to tighter or to looser binding of the substrate.
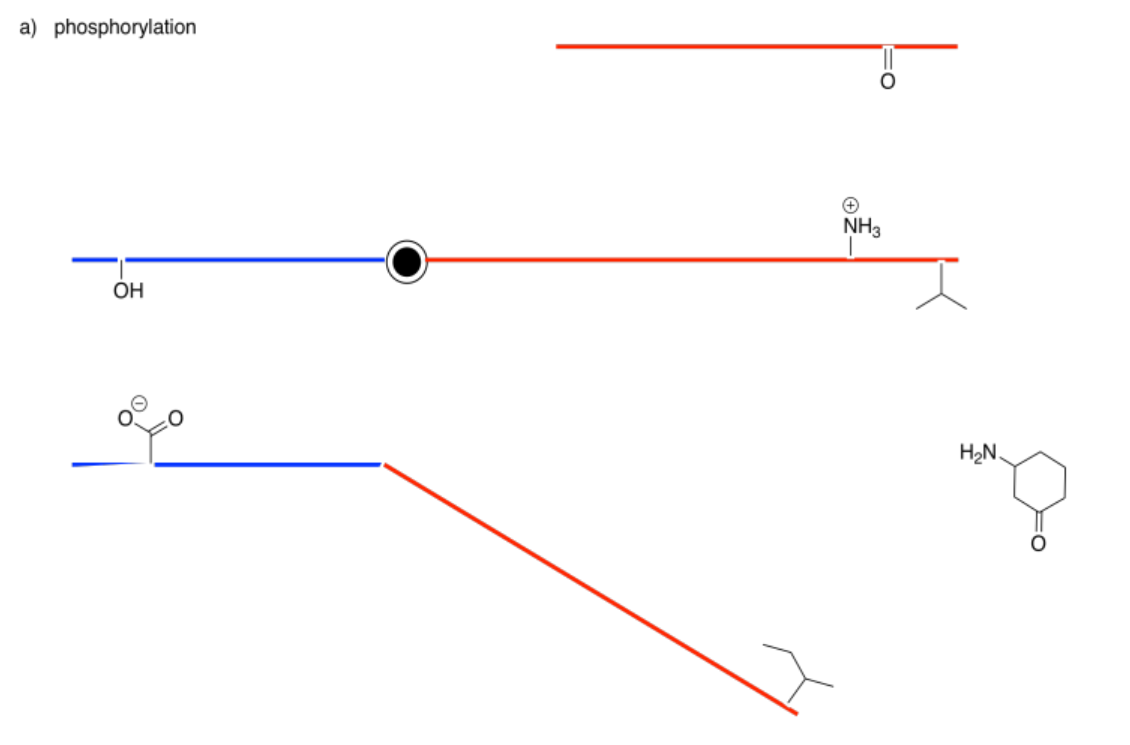
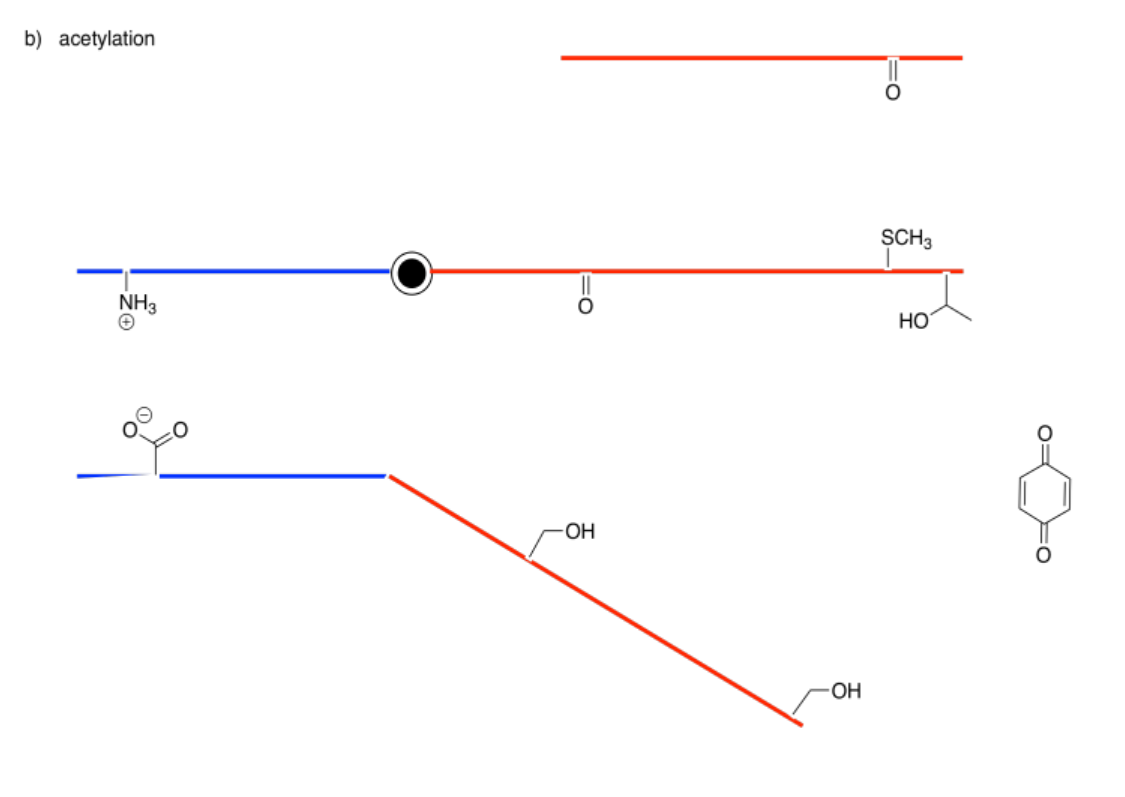
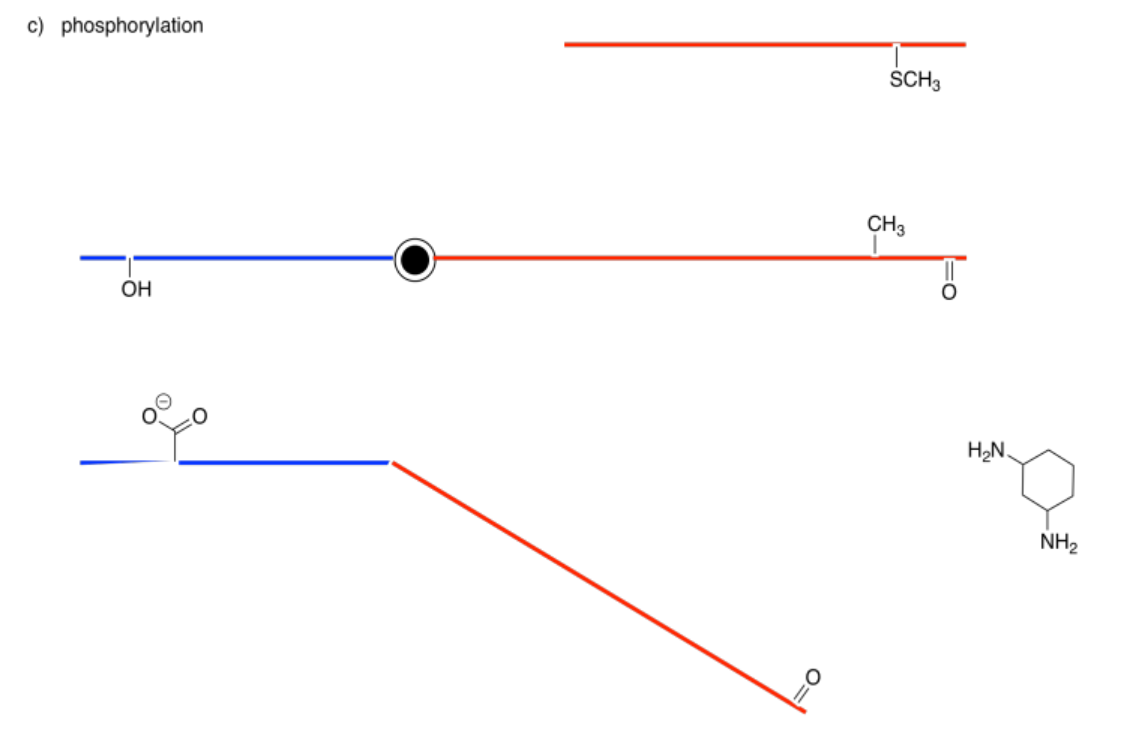
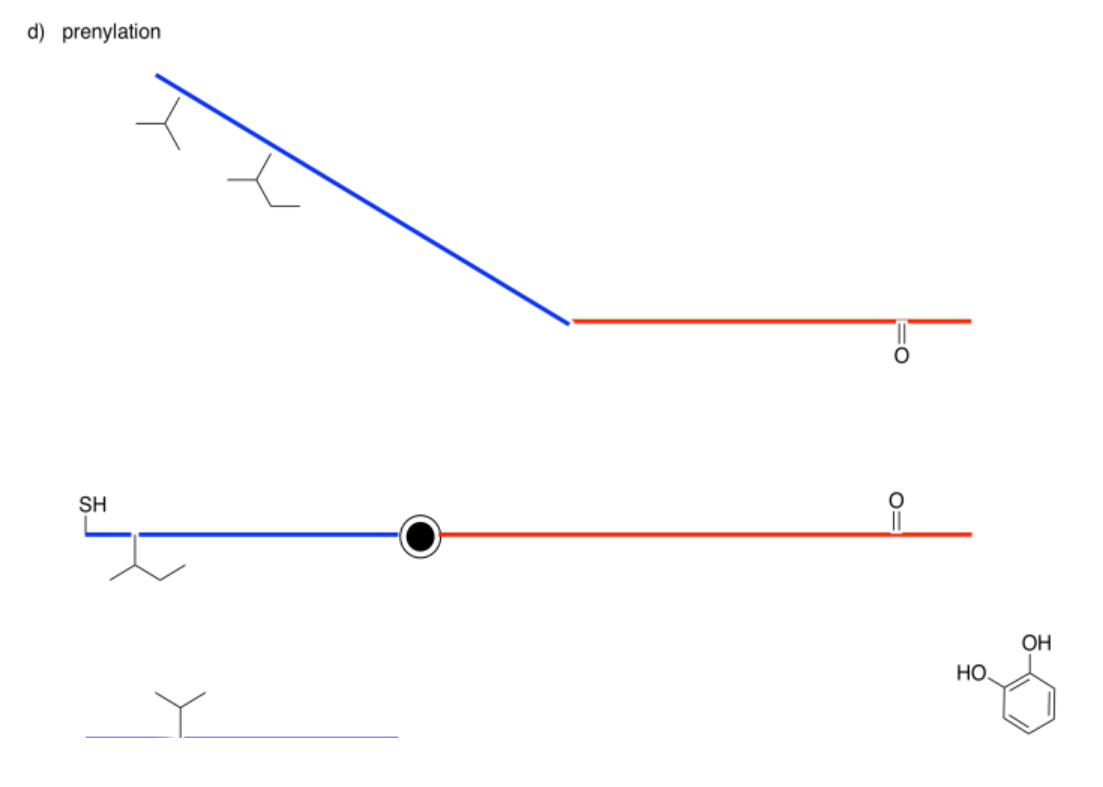
Answer-



