5.13: Peptide and Protein Synthesis
- Page ID
- 199679
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Peptides and proteins are very important in biology. As a result, synthesis of these molecules has become very important, allowing for the laboratory study of model compounds that can give us insight into how proteins work, as well as pharmaceutically important compounds.
Insulin and glucagon are two important peptides (or small proteins) that regulate blood sugar. Insulin signals that blood sugar levels are high and that the body should begin storing this excess sugar. Glucagon signals that blood sugar levels are low and so the body may need to access its long-term energy stores. Both compounds are important in medicine.
Glucagon has a relatively simple structure, for a protein. It is a string of just 29 amino acids, connected together in the following order:
His-Ser-Gln-Gly-Thr-Phe-Thr-Ser-Asp-Tyr-Ser-Lys-Tyr-Leu-Asp-Ser-Arg-Arg-Ala-Gln-Asp-Phe-Val-Gln-Trp-Leu-Met-Asn-Thr.
The structure is composed of readily-available starting materials. The amino acids are simple units that connect together to form a chain. Conceptually, it seems easy to put two amino acids together to make a side chain. When two amino acids come together, they lose a molecule of water together, and the remaining pieces are able to bond to each other to make the dipeptide. It ought to be just as easy to add a third, and so on.
Structurally, amide or peptide bonds are very stable and resistant to carboxyl substitution. That stability makes optimal structures from which to construct proteins. Despite being composed of very long chains of linked amino acids, proteins actually have some limits on their conformational flexibility (their "floppiness"). That allows proteins to more reliably hold a particular shape. The shape of proteins is crucial to their function as enzymes and other
Both the stability and the structural rigidity of peptides arises from the nature of the peptide bond. The pi donation that hinders nucleophiles from substituting at the carbonyl is pronounced enough that it can be considered to form an additional bond. Thus, peptides behave as though they contain C=N bonds rather than C-N bonds. X-ray structure determinations show that the peptide nitrogens in proteins are trigonal planar, not pyramidal. In addition, many peptides exhibit cis-trans isomerism. For every peptide bond, two different isomers can occur, depending on whether a substituent attached to nitrogen is on the same side of the C=N bond as the carbonyl oxygen or the opposite side.
The great stability of these structures does not mean they are easy to make. Part of the difficulty stems from the fact that amino acids are difunctional. In order to form long chain structures, amino acids must be able to react twice: once with an amine, to grow in one direction, and once with a carboxylic acid to grow in the other direction. In other words, an amino acid contains both a nucleophile and an electrophile.
Suppose we were to try to make the dipeptide, ala-phe. This peptide contains an alanine connected to a phenylalanine through a peptide bond. The peptide bond is formed between the carboxylic acid of alanine and the amine of phenylalanine.
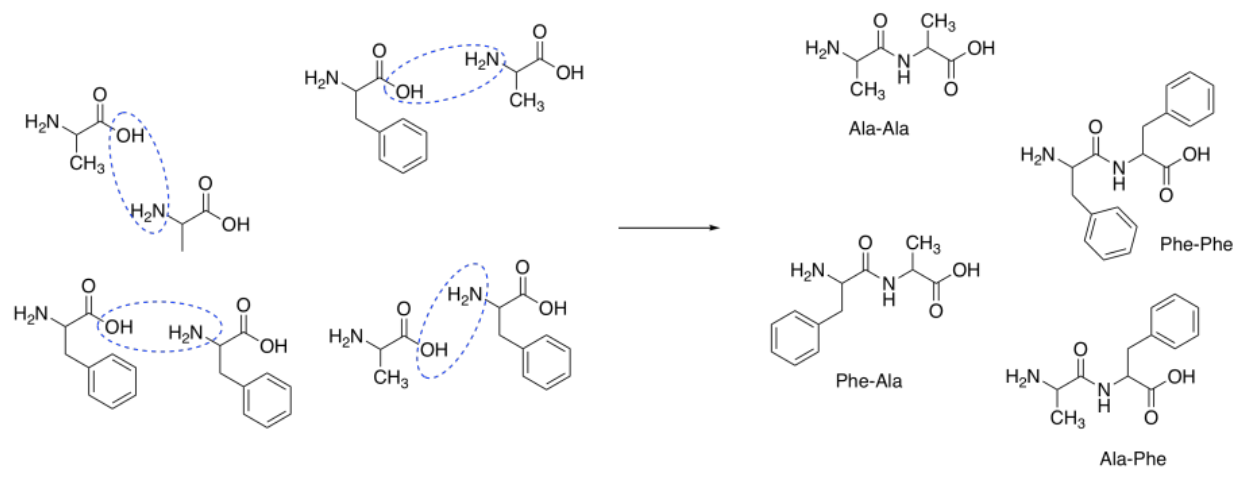
Assuming the amino acids do react together to form the peptide, combining these two reactants would likely produce a mixture of four dipeptides:
Ala-Phe Ala-Ala Phe-Phe Phe-Ala
In other words, peptide formation from amino acids is non-selective.
Exercise \(\PageIndex{1}\)
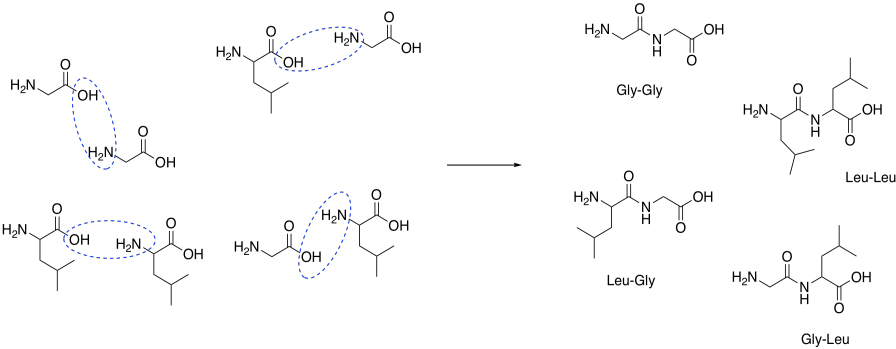
Draw structures for the four peptides formed by combining glycine and leucine.
- Answer
-
Exercise \(\PageIndex{2}\)
What tripeptides would be produced by mixing Ala, Gly and Val?
- Answer
-
That combination would give thw following dipeptides:
ala-ala ala-gly ala-val
gly-gly gly-ala gly-val
val-val val-ala val-gly
Of course, we might also get tripeptides, such as ala-ala-gly-val, and so on.
Exercise \(\PageIndex{3}\)
Simply combining these peptides might not result in any peptide formation at all. Why not?
- Answer
-
Carboxylic acids usually require activation before they can act as nucleophiles. That problem is actually complicated here because the carboxylic acid is in equilibrium with a carboxylate salt (read further on the page).
An additional complication in peptide synthesis is that amines and carboxylic acids do not really exist together. Instead, a proton is transferred from the carboxylic acid to the amine, forming a salt. The carboxylate is no longer very electrophilic, due to its negative charge. Because of its positive charge, the ammonium ion is no longer very nucleophilic.
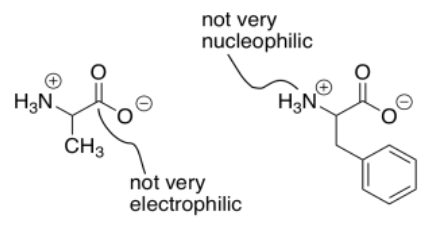
As a result, there are actually two distinct problems in peptide synthesis. There is a selectivity problem, because each amino acid has a nucleophilic part and an electrophilic part. There is no way to ask one compound to react only using its electrophile and another compound to react only using its nucleophile. There is also a reactivity problem: the carboxyl group in this case is a terrible electrophile, and the amine is a terrible nucleophile.
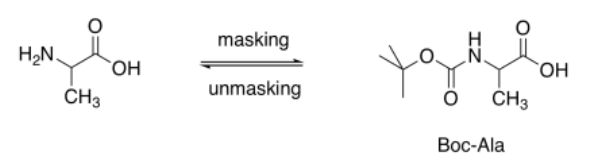
In laboratory syntheses, a number of techniques have been used to make peptide synthesis selective. Most frequently, protecting groups are used. A protecting group "masks" one of the two functional groups on an amino acid, but leaves the other one open. If one amino acid has its amine protected, it can only react via its carboxylic acid. If the other amino acid has its carboxylic acid protected, it can only react via its amino group. Only one combination will result.
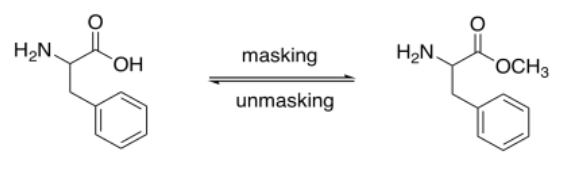
The key to protecting groups is that the reaction used to mask one of the functional groups must be reversible. You must be able to take the protecting group back off when it is no longer needed.
Carboxylic acids are normally protected as esters. Esters can be removed via acid- or base-catalyzed hydrolysis (as can amides, but esters are more reactive, being farther up the ski hill).
Amines are normally protected as amides. However, we need to be able to remove specific amides her: the ones that mask the amines, not the ones that we have formed to link two amino acids together. As a result, in peptide synthesis, amines are usually protected as carbamates. Carbamates can be cleaved more easily than amides.
Exercise \(\PageIndex{4}\)
Propose a reason for the relatively higher reactivity of carbamates compared to amides.
- Answer
-
The relative reactivity of carboxyloids results from a balance between sigma electron withdrawing effects and pi-donation. An electronegative atom attached to a carbonyl tends to withdraw electron density, making the carbonyl even more positive and electrophilic. On the other hand, pi-donation from a neighbouring atom with a lone pair actually lowers electrophilicity by forming a stable, conjugated system.
In a carbamate, an additional electronegative atom is added to the carbonyl: it has an oxygen as well as a nitrogen adjacent to the C=O group. That atom draws electron density away from the carbonyl, making it more electrophilic. However, pi-donation from the additional oxygen does not result in a more stable conjugated system. The maximum pi system is still just three atoms long; it either involves conjugation of the O-C=O or the N-C=O unit. It does not, for example, lead to an even more stable conjugated system that is four atoms long.
As a result, the added oxygen probably contributes more to the electron-withdrawing effect than it does to stabilisation of the pi system.
Exercise \(\PageIndex{5}\)
Fill in the blanks in the following peptide synthesis.
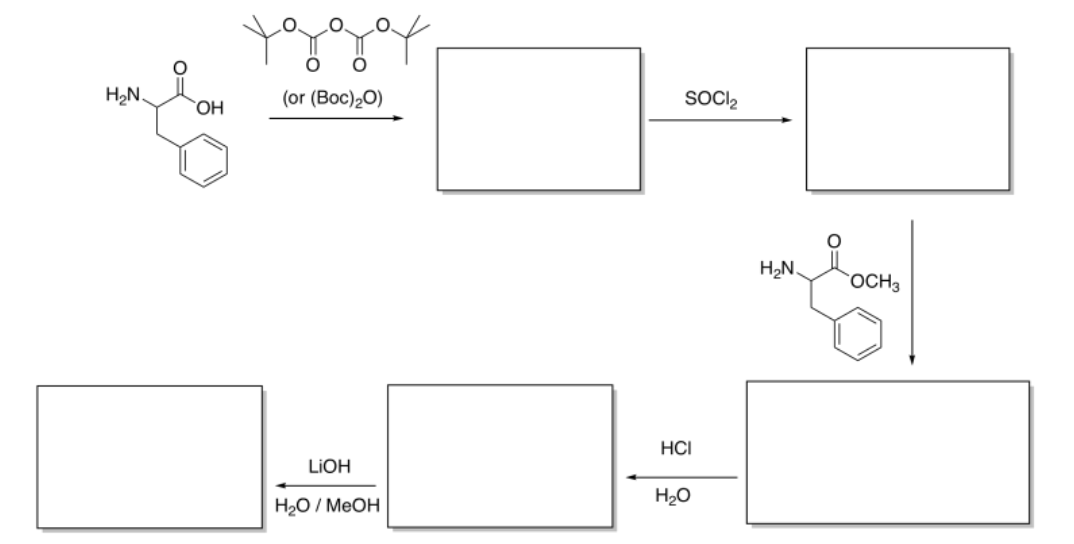
Answer-
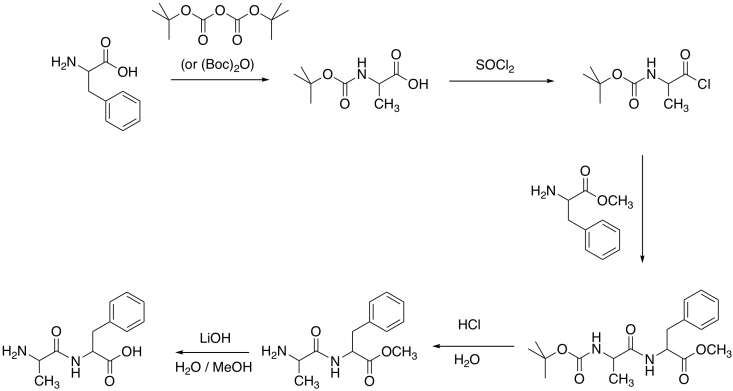
To get around the problem of low electrophilicity of the carboxylic acid, a number of coupling agents have been developed. A coupling agent can temporarily convert the carboxylate anion into a more reactive electrophile. To do so, it exploits the nucleophilicity of the carboxylate anion. After donating to the coupling agent, the carbonyl compound becomes more electrophilic.
Thionyl chloride (SOCl2) can accomplish this goal, of course. It converts relatively non-electrophilic carboxylic acids into much more electrophilic acid chlorides. Thionyl chloride can be a little harsh, however, so chemists have sought to develop milder conditions that can knit two amino acids together without the necessity of forming a reactive acid chloride.
Some of the most commonly-used coupling agents for peptide synthesis are carbodiimides. These compounds contain an electrophilic N=C=N unit to act as an initial electrophile. Diisopropylcarbodiimide (DIC) and dicyclohexylacarbodiimide (DCC) were some of the earliest and simplest examples of these compounds developed for peptide synthesis.
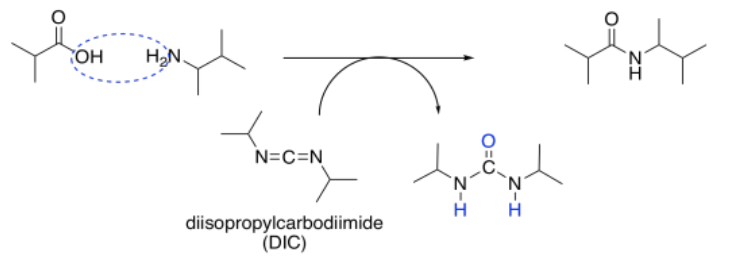
Once a carboxylate has donated to the electrophilic carbon of the carbodiimide, a better leaving group is formed. The adjacent nitrogen atoms act as basic sites, picking up a proton from the carboxylic acid on one amino acid and from the ammonium ion intermediate formed by the other amino acid.
Exercise \(\PageIndex{6}\)
Propose an advantage of EDCI as a coupling agent for amino acids.
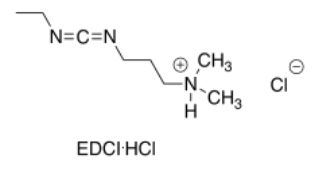
Answer-
The more polar hydrochloride salt of EDCI is often used, as pictured. This more polar compound dissolves well in polar solvents, such as water, that also dissolve amino acids.
One can also imagine the amino group in EDCI acting as a site for catalysis, shuttling protons from one place to another.


