Protein Folding
- Page ID
- 496
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Introduction and Protein Structure
Proteins have several layers of structure each of which is important in the process of protein folding. The first most basic level of this structure is the sequence of amino acids themselves.1 The sequencing is important because it will determine the types of interactions seen in the protein as it is folding. A novel sequence-based method based on the assumption that protein-protein interactions are more related to amino acids at the surface than those at the core.2 This study shows that not only is the amino acids that are in a protein important but also the order in which they are sequenced. The interactions of the amino acids will determine what the secondary and tertiary structure of the protein will be.
The next layer in protein structure is the secondary structure. The secondary structure includes architectural structures that extend in one dimension.1 Secondary structure includes α-Helixes and β-sheets ( Figure \(\PageIndex{1}\)). The α-helices, the most common secondary structure in proteins, the peptide –CO–NH–groups in the backbone form chains held together by NH ̄OC hydrogen bonds.”3 The α-helices form the backbone of proteins and help to aid in the folding process. The β-sheets form in two distinct ways. They are able to form in both parallel β-pleated sheets and anti parallel β-pleated sheets.1 When the α-helix or β-sheet is formed, the excluded volumes generated by the backbone and side chains overlap, leading to an increase in the total volume available to the translational displacement of water molecules.4 This is important because it leads to a more thermodynamically stable conformation and leads to less strain on the protein as a whole and thus are aided by the conformation.
The tertiary structure is the next layer in protein structure. This takes the α-Helixes and β-sheets and allows them to fold into a three dimensional structure.1 Most proteins take on a globular structure once folded. The description of globular protein structures as an ensemble of contiguous closed loops or tightened end fragments reveals fold elements crucial for the formation of stable structures and for navigating the very process of protein folding.5 The globular proteins generally have a hydrophobic core surrounded by a hydrophilic outer layer. These interactions are important because they lead to the global structure and help create channels and binding sites for enzymes.

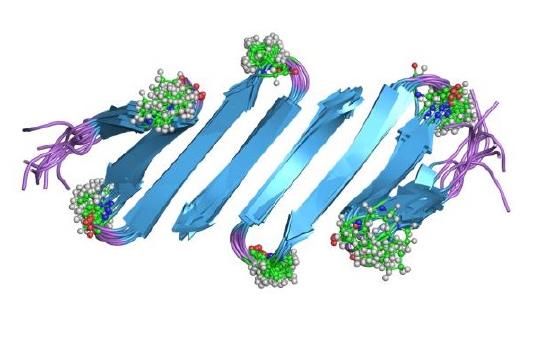
The last layer of protein structure is the quaternary structure. The folding transition and the functional transitions between useful states are encoded in the linear sequence of amino acids, and a long- term goal of structural biology is to be able to predict both the structure and function of molecules from the information in the sequence.6 The Subunit organization is the last level of structure in protein molecules.1 The organization of the subunits is important because that determines the types of interactions that can form and dictates its use in the body.
Protein Folding
Proteins are folded and held together by several forms of molecular interactions. The molecular interactions include the thermodynamic stability of the complex, the hydrophobic interactions and the disulfide bonds formed in the proteins. The figure below (Figure \(\PageIndex{2}\)) is an example of protein folding.
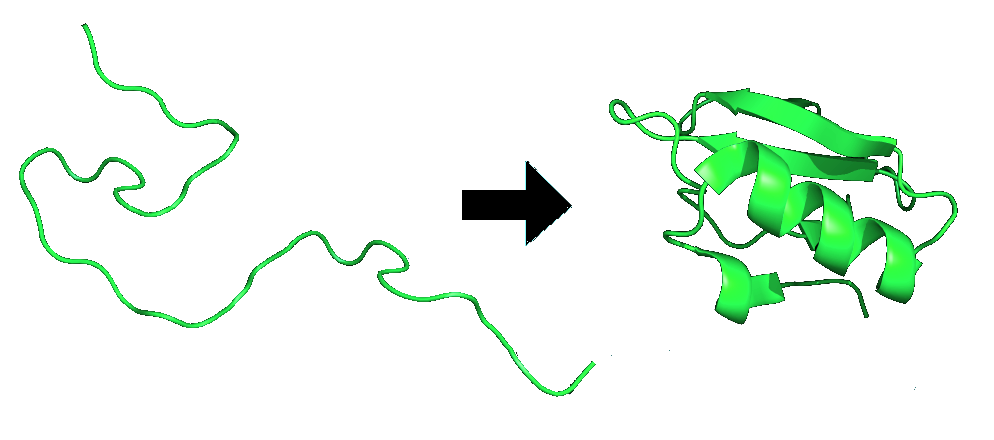
The biggest factor in a proteins ability to fold is the thermodynamics of the structure. The interaction scheme includes the short-range propensity to form extended conformations, residue-dependent long-range contact potentials, and orientation-dependent hydrogen bonds.7 The thermodynamics are a main stabilizing force within a protein because if it is not in the lowest energy conformation it will continue to move and adjust until it finds its most stable state. The use of energy diagrams and maps are key in finding out when the protein is in the most stable form possible.
The next type of interaction in protein folding is the hydrophobic interactions within the protein. The framework model and the hydrophobic collapse model represent two canonical descriptions of the protein folding process. The first places primary reliance on the short-range interactions of secondary structure and the second assigns greater importance to the long-range interactions of tertiary structure.6 These hydrophobic interactions have an impact not just on the primary structure but then lead to changes seen in the secondary and tertiary structure as well. Globular proteins acquire distinct compact native con- formations in water as a result of the hydrophobic effect.7 When a protein has been folded in the correct way it usually exists with the hydrophobic core as a result of being hydrated by waters in the system around it which is important because it creates a charged core to the protein and can lead to the creation of channels within the protein. The hydrophobic interactions are found to affect time correlation functions in the vicinity of the native state even though they have no impact on same time characteristics of the structure fluctuations around the native state.7 The hydrophobic interactions are shown to have an impact on the protein even after it has found the most stable conformation in how the proteins can interact with each other as well as folding themselves.

Another type of interaction seen when the protein is folding is the disulfide linkages that form in the protein (Figure \(\PageIndex{3}\)). The disulfide bond, a sulfur- sulfur chemical bond that results from an oxidative process that links nonadjacent (in most cases) cysteine’s of a protein.9 These are a major way that proteins get into their folded form. The types of disulfide bonds are cysteine-cysteine linkage is a stable part of their final folded structure and those in which pairs of cysteines alternate between the reduced and oxidized states.9 The more common is the linkages that cause the protein to fold together and link back on itself compared to the cysteines that are changing oxidation states because the bonds between cysteines once created are fairly stable.
Misfunctions
Proteins can miss function for several reasons. When a protein is miss folded it can lead to denaturation of the protein. Denaturation is the loss of protein structure and function.1 The miss folding does not always lead to complete lack of function but only partial loss of functionality. The miss functioning of proteins can sometimes lead to diseases in the human body.
Alzheimer's Disease
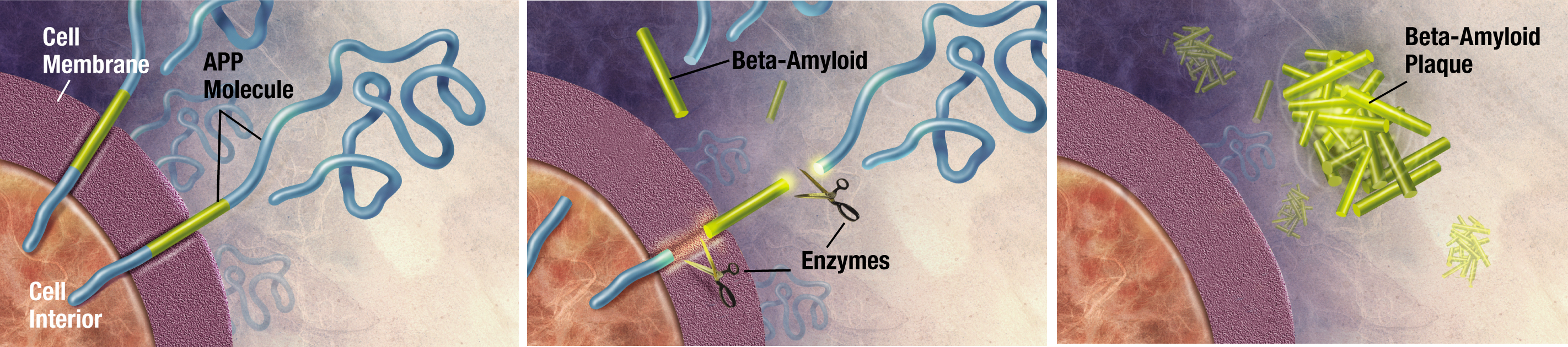
Alzheimer's Disease (AD) is a neurological degenerative disease that affects around 5 million Americans, including nearly half of those who are age 85 or older.10 The predominant risk factors of AD are age, family history, and heredity. Alzheimer’s disease typically results in memory loss, confusion of time and place, misplacing places, and changes in mood and behavior.11 AD results in dense plaques in the brain that are comprised of fibrillar β-amyloid proteins with a well-orders β-sheet secondary structure.12 These plaques visually look like voids in the brain matter (see figure 5) and are directly connected to the deterioration of thought processes. It has been determined that AD is a protein misfolding disease, where the misfolded protein is directly related to the formation of these plaques in the brain.13
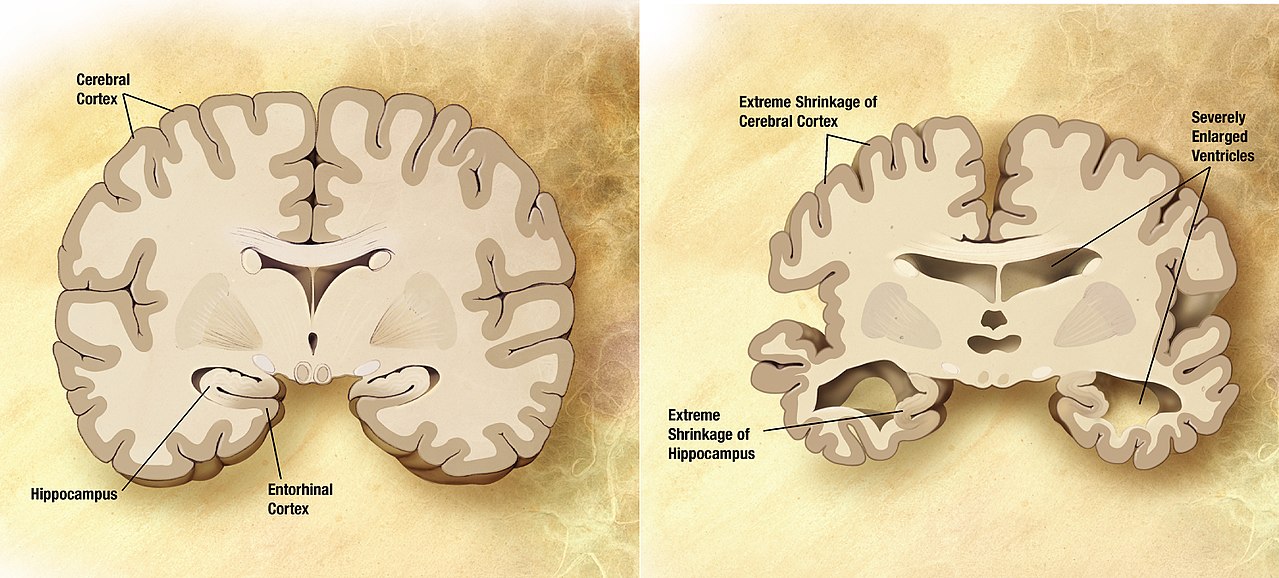
It is yet to be fully understood what exactly causes this protein misfolding to begin, but several theories point to oxidative stress in the brain to be the initiating factor. This oxidation results in damage to the phospholipids in the brain, which has been found to result in a faster accumulation of amyloid β-proteins.14
Cystic Fibrosis
Cystic Fibrosis (CF) is a chronic disease that affects 30,000 Americans. The typical affects of CF is a production of thick, sticky mucus that clogs the lungs and leads to life-threatening lung infection, and obstructs the pancreas preventing proper food processing.15 CF is caused by protein misfolding. This misfolding then results in some change in the protein known as cystic fibrosis transmembrane conductance regulator (CFTR), which can result in this potentially fatal disease.16 In approximately 70% of CF cases, a deletion of phenylalanine at position 508 in the CFTR is deleted. This deletion of Phe508 seems to be directly connected to the formation of CF.17 The protein misfolding that results in CF occurs prior to birth, but it is not entirely clear as to why.
Resources
- Garrett, R.H., and Grisham, C.M. Biochemistry fourth edition; Brooks/Cole. Australia, 2010. p. 93-95, 135, 143, 160.
- Chang, D.T., Syu Y., Lin, P., Predicting the protein-protein interactions using primary structures with predicted protein surface. BCM Bioinformatics. Jan 2010
- Bodis, P., Schwartz, E., Koepf, M. Cornelissen, J. Rowan A.E., Roeland J. Nolte, M. and Woutersen S. Vibrational self-trapping in beta-sheet structures observed with femtosecond nonlinear infrared spectroscopy. The Journal Of Chemical Physics 131, 2009
- Yasuda, S., Yoshidome, T. Oshima, H. Kodama R., Harano, Y. and Kinoshita, M. Effects of side-chain packing on the formation of secondary structures in protein folding. The Journal of Chemical Physics 132, 2010
- Papandreoul, N., Berezovsky I.N., Lopes, A., Eliopoulos, E., and Chomilier, J. Universal positions in globular proteins from observation to simulation. Eur. J. Biochem. 271, 4762–4768 (2004)
- Hausratha, A.C. A Kinetic theory of tertiary contact formation coupled to the helix-coil transition in polypeptides. The Journal of Chemical Physics. 2006
- Cieplaka M. and Niewieczerza S. Hydrodynamic interactions in protein folding. Journal of Chemical Physics. 130 2009
- Gront, D. Kolinskia, A. and Skolnick, J. A new combination of replica exchange Monte Carlo and histogram analysis for protein folding and thermodynamics. Journal of Chemical Physics. Vol. 115 2001
- Kadokura, H., Katzen F. and Beckwith, J. Protein disulfide bond formation in prokatyotes. Annual Review Biochem 2003.
- Alzheimer's Disease. Centers for Disease Control and Prevention. 2010, http://www.cdc.gov/aging/aginginfo/alzheimers.htm.
- Alzheimer's Association. 2011, www.alz.org.
- Glenner, G. G.; Wong, C. W. Alzheimer's disease: initial report of the purification and characterization of a novel cerebrovascular amyloid protein. Biochem. Biophys. Res. Commun.1984, 120, 885-890.
- Hashimoto, M.; Rockenstein, E.; Crews, L.; Mashliah, E. Role of Protein Aggregation in Mitochondrial Dysfunction and Neurodegeneration in Alzheimer’s and Parkinson’s Diseases. Neuromolecular Med. 2003, 4, 21-36
- The Cystic Fibrosis Foundation. 2011, http://www.cff.org/AboutCF.
- Cheung, J. C. and Deber, C. M. Misfolding of the Cystic Fibrosis Transmembrane Conductance Regulator and Disease. Biochemistry 2008, 47, 1465-1473.
- Koppaka, V.; Axelsen, P. Accelerated Accumulation of Amyloid β Proteins on Oxidatively Damaged Lipid Membranes. Biochemistry 2000, 39, 10011-10016.
- Riordan, J.M. et al. Identification of the cysticfibrosisgene: cloning and characterization of complementary DNA. Science 1989, 245, 1066-1073.

