7.10: Transport Across Cell Membranes
- Page ID
- 517609
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Source: BiochemFFA_3_2.pdf. The entire textbook is available for free from the authors at http://biochem.science.oregonstate.edu/content/biochemistry-free-and-easy
Introduction
Ions generally cross cell membranes via ion channels.
See the electrochemical discussion of ion transport for more energetics details.
Selective passive ion channels
Ion channels are pore-forming membrane proteins in the membranes of all cells that regulate movement of selected ions across a membrane (Figures \(\PageIndex{1}\) & \(\PageIndex{2}\)). They help to establish the resting membrane potential and to affect action potentials and other electrical signals. They are very important in the process of nerve transmission. Ion channels control the flow of ions across secretory and epithelial cells, and consequently help to regulate cell volume by affecting osmotic pressure.
Ion channels are essential features of almost all cells, functioning as selective “tunnels” that restrict movement through them to ions with specific characteristics (typically size). The size of the opening is very narrow (usually one or two atoms wide) and is able to select even against ions that are too small.
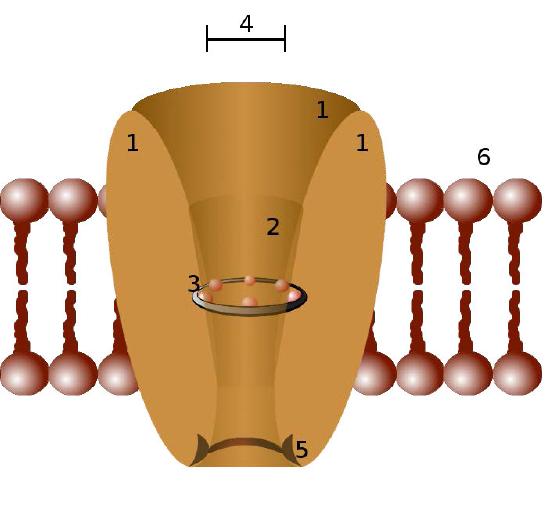
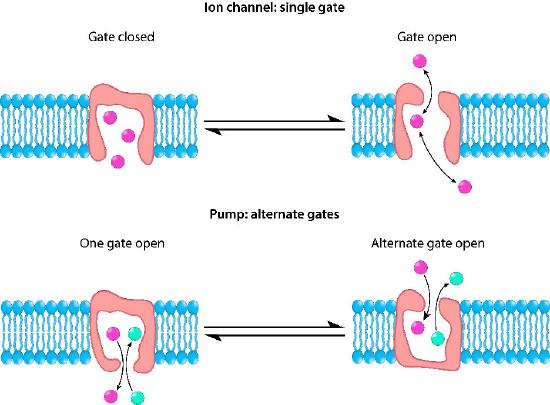
Control mechanisms
Ion channels are controlled by mechanisms that include voltage, ligands, light, temperature, and mechanical deformation (stretch activated). Ligand-gated ion channels (LGICs) are transmembrane proteins which open to selectively allow ions such as Na+, K+, Ca++, or Cl− to pass through the membrane in response to the binding of a ligand messenger.
Sound waves cause mechanical deformation of hair cells in the ear. This results in the opening of ion channels and initiation of a nerve signal to the brain.
Sodium ion channels in the tongue for sugar receptors open in response to binding of sucrose, allowing sodium concentration in the nerve cell to increase and initiate a nerve signal to the brain. In this case, the default for the gate is to be closed and it opens in response to binding of a ligand (sucrose).
In light sensing cells of the eye, calcium gates are open by default, but stimulation by light causes them to close, triggering a series of events that result in a signal being sent the brain about the perception of light. Thus, in this case, the stimulus (light) causes an open channel to close.
Moving the other direction, nerve signals originating in the brain travel to muscle tissue and through a complicated set of exchanges, result in the opening of calcium gates of muscle cells, increasing the concentration of calcium and stimulating muscular contraction.
Voltage gated channels are essential for transmission of nerve signals.
Ion movement through channels
The ability of ion channels to select against ions too large is intuitive - the size of the opening in the ion channel simply isn’t big enough for a larger ion to fit through the opening. Potassium, for example, passes through sodium channels rarely because the opening is too small.
Potassium channels that are selective for potassium ions must be big enough to allow potassium to enter, but if size were the only selection means, then sodium ions would also readily pass through potassium channels, since sodium ions (0.95 Å) are smaller than potassium ions (1.33 Å). In order for potassium channels to select against sodium ions and favor potassium ions, other considerations come into play.
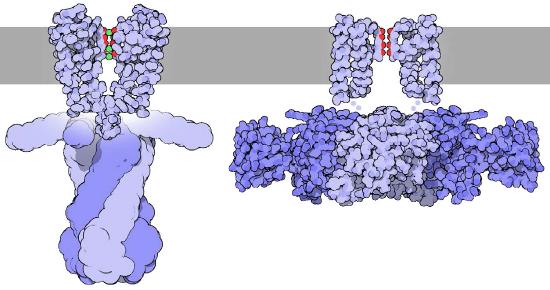
Hydration shell
To understand this unique selectivity, it is important to understand how ions move through channels. Before an ion can pass through a channel, it must first be dissociated from (stripped of) the water molecules in its hydration shell - water molecules surrounding ions in aqueous solutions (Figure \(\PageIndex{4}\)). This process requires an input of energy.
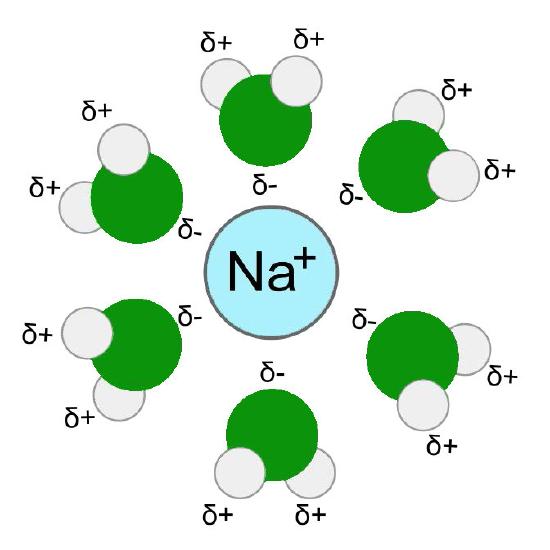
Comparable to enzymes
Just as enzymes lower the activation energy of enzymatic reactions and thus allow them to more readily occur, so too do channel proteins lower the energy requirements for a molecule to traverse a lipid bilayer. In the absence of the channel protein, the dehydration energy is mostly prohibitive for most polar molecules to occur, so very few make it across the lipid bilayer without the channel protein. This is why ion channel/transport proteins are so important to the cell.
After the water has been stripped, the ion can pass through the channel and when it arrives at the other side of the channel, the diffusing ion becomes rehydrated, thus regaining the energy that was required initially to strip away the water molecules from the ion.
Selectivity of the potassium channel
The potassium channel (Figure \(\PageIndex{5}\)) uses the dimensions of the potassium ion precisely to shepherd it through the channel. The sodium ion, which has different dimensions has a more difficult time making it through the channel despite its smaller size. The reason this is rooted in the energy required for dehydration.
For potassium ions, after the water has been stripped off, precisely positioned carbonyl groups along the channel help to stabilize the ion as it moves. The sodium ion, on the other hand is too small and does not make efficient connections with carbonyl groups and thus has a more difficult path. Because of this, the energy difference between dehydration and rehydration of a sodium ion in a potassium channel is more energetically unfavorable than the same process for a potassium ion. The lower effective activation energy for dehydration of the potassium ion in the channel leads to the selectivity.
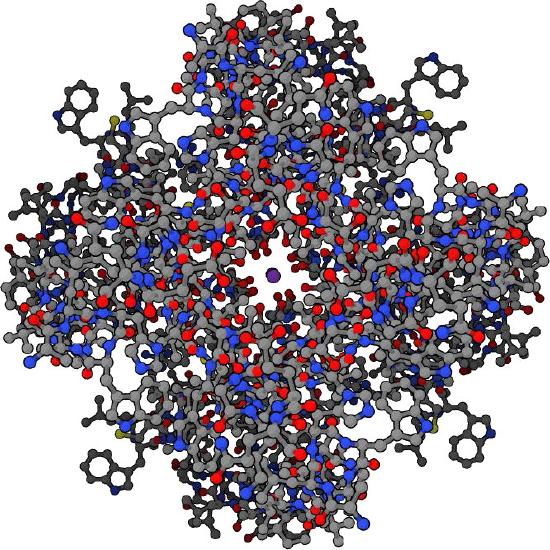
Ions only flow down the concentration gradient through these types of channels. Because they are often unidirectional they will not contribute to ion flow in the opposite direction even if that is down the concentration gradient.
Active transport of ions
During active transport, at least one molecule is being moved against a concentration gradient. A common, but not exclusive, energy source is ATP (see Na+/K+ ATPase), but other energy sources are also employed. For example, the sodium-glucose transporter uses a sodium gradient as a force for actively transporting glucose into a cell. Thus, it is important to know that not all active transport uses ATP energy.
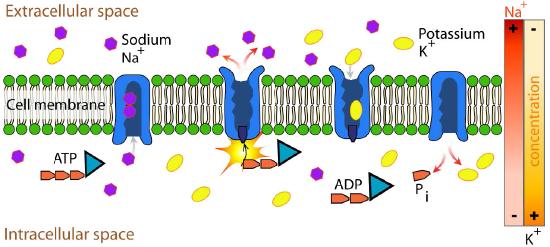
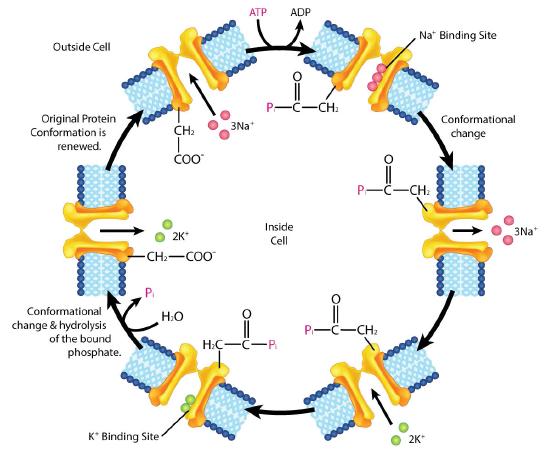
Example: Na+/K+ ATPase
An important integral membrane transport protein is the Na+/K+ ATPase antiport (Figures \(\PageIndex{6}\) and \(\PageIndex{7}\)), which moves three sodium ions out of the cell and two potassium ions into the cell with each cycle of action. In each case, the movement of ions is against the concentration gradient. Since three positive charges are moved out for each two positive charges moved in, the system is electrogenic.
The protein uses the energy released during ATP hydrolysis to create ion gradients that are important both in maintaining cellular osmotic pressure and (in nerve cells) for creating the sodium and potassium gradients necessary for signal transmission. Failure of the system to function results in swelling of the cell due to movement of water into the cell through osmotic pressure. The transporter expends about one fifth of the ATP energy of animal cells.
The cycle of action occurs as follows:
- Pump binds ATP followed by binding of 3 Na+ ions from cytoplasm of cell
- ATP hydrolysis results in phosphorylation of aspartate residue of pump. ADP is released
- Phosphorylated pump undergoes conformational change to expose Na+ ions to exterior of cell. Na+ ions are released.
- Pump binds 2 extracellular K+ ions.
- Pump dephosphorylates causing it to expose K+ ions to cytoplasm as pump returns to original shape.
- Pump binds 3 Na+ ions, binds ATP and releases 2 K+ ions to restart process
The Na+/K+ ATPase is classified as a P-type ATPase. This category of pump is notable for having a phosphorylated aspartate intermediate and is present across the biological kingdoms - bacteria, archaeans, and eukaryotes.



