5.4.2: Base Pairing in DNA - The Watson-Crick Model
- Last updated
- Save as PDF
- Page ID
- 413283
Objectives
After completing this section, you should be able, given the necessary Kekulé structures, to show how hydrogen bonding can occur between thymine and adenine, and between guanine and cytosine; and to explain the significance of such interactions to the primary and secondary structures of DNA.
Study Notes
Watson and Crick received the Nobel Prize in 1962 for elucidating the structure of DNA and proposing the mechanism for gene reproduction. Their work rested heavily on X-ray crystallographic work done on RNA and DNA by Franklin and Wilkins. Wilkins shared the Nobel Prize with Watson and Crick, but Franklin had been dead four years at the time of the award (you cannot be awarded the Nobel Prize posthumously).
The history of Watson and Crick’s proposed DNA model is controversial and a travesty of scientific ethics. Rosalind Franklin was deeply involved in the determination of the structure of DNA, and had collected numerous diffraction patterns. Watson attended a departmental colloquium at King’s College given by Franklin, and came into possession of an internal progress report she had written. Both departmental colloquia and progress reports are merely methods of discussion between colleagues; works presented in these fora are not considered by scientists to be “published” works, and therefore are not in the public domain. Watson and Crick not only were aware of Franklin’s work, but used her unpublished data, presented in confidence within her own college.
The final blow came about a year after the colloquium. Watson visited Wilkins at King’s College, and Wilkins inexplicably handed over Franklin’s diffraction photographs without her consent. Had Franklin’s work not been secretly taken from her, she might quite possibly have solved the DNA structure before Watson and Crick, who at the time did not yet have their own photographs. This is truly one of the sadder episodes of questionable scientific ethics and discovery that I have ever encountered.
References
Kass-Simon, G., and P. Farnes. Women of Science: Righting the Record. Bloomington, IN: Indiana University Press, 1990.
Maddox, B. Rosalind Franklin: The Dark Lady of DNA. New York: HarperCollins, 2002.
Intermolecular Forces in Nucleic Acids
The nucleic acids RNA and DNA are involved in the storage and expression of genetic information in a cell. Both are polymers of monomeric nucleotides. DNA exists in the cell as double-stranded helices while RNA typically is a single-stranded molecule which can fold in 3D space to form complex secondary (double-stranded helices) and tertiary structures in a fashion similar to proteins. The complex 3D structures formed by RNA allow it to perform functions other than simple genetic information storage, such as catalysis. Hence most scientists believe that RNA preceded both DNA and proteins in evolution as it can both store genetic information and catalyze chemical reactions.
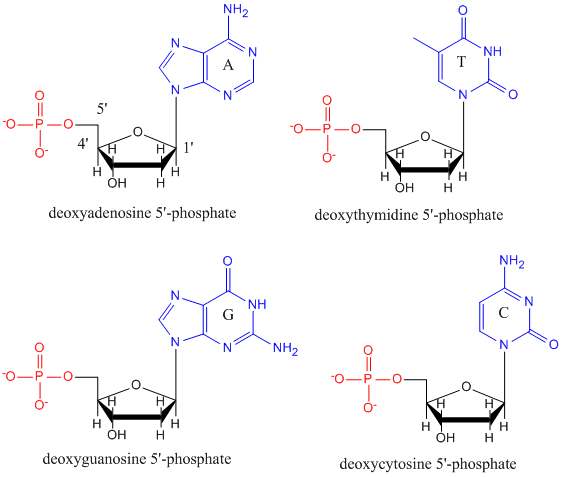
DNA
DNA is a polymer, consisting of monomers call deoxynucleotides. The monomer contains a simple sugar (deoxyribose, shown in black above), a phosphate group (in red), and a cyclic organic R group (in blue) that is analogous to the side chain of an amino acid. Only four bases are used in DNA (in contrast to the 20 different side chains in proteins) which we will abbreviate, for simplicity, as A, G, C and T. They are bases since they contain amine groups that can accept protons. The polymer consists of a sugar - phosphate - sugar - phosphate backbone, with one base attached to each sugar molecule. As with proteins, the DNA backbone is polar but also charged. It is a polyanion. The bases, analogous to the side chains of amino acids, are predominately polar. Given the charged nature of the backbone, you might expect that DNA does not fold to a compact globular (spherical) shape, even if positively charged cations like Mg bind to and stabilize the charge on the polymer. Instead, DNA exists usually as a double-stranded (ds) structure with the sugar-phosphate backbones of the two different strands running in opposite directions (5'-3' and the other 3'-5'). The strands are held together by hydrogen bonds between bases on complementary strands. Hence like proteins, DNA has secondary structure but in this case, the hydrogen bonds are not within the backbone but between the "side chain" bases on opposing strands. It is actually a misnomer to call dsDNA a molecule, since it really consists of two different, complementary strands held together by hydrogen bonds. A structure of ds-DNA showing the opposite polarity of the strands is shown below.
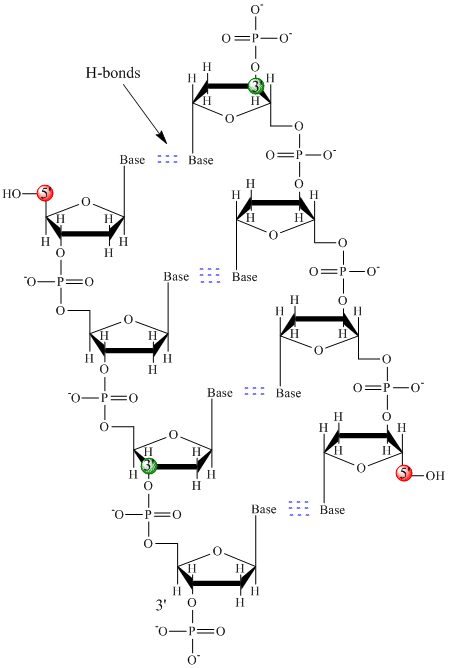
In 1950, Erwin Chargaff of Columbia University showed that the molar amount of adenine (A) in DNA was always equal to that of thymine (T). Similarly, he showed that the molar amount of guanine (G) was the same as that of cytosine (C). Chargaff's findings clearly indicate that some type of heterocyclic amine base pairing exists in the DNA structure. In double stranded DNA, the guanine (G) base on one strand can form three H-bonds with a cytosine (C) base on another strand (this is called a GC base pair). The thymine (T) base on one strand can form two H-bonds with an adenine (A) base on the other strand (this is called an AT base pair). Double-stranded DNA has a regular geometric structure with a fixed distance between the two backbones. This requires the bases pairs to consists of one base with a two-ring (bicyclic) structure (these bases are called purines) and one with a single ring structure (these bases are called pyrimidines). Hence a G and A or a T and C are not possible base pair partners.
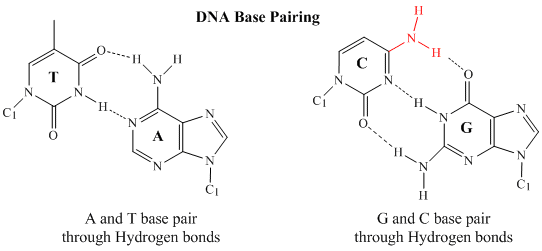
Secondary Structure of DNA
The three-dimensional structure of DNA was the subject of an intensive research effort in the late 1940s to early 1950s. DNA exists as a double-stranded molecule that twists around its axis to form a helical structure,stabilized through Watson-Crick hydrogen bonding between purines and pyrimidines, and through pi-pi stacking interactions among the bases arranged in structure. helical column. Each strand is a complement to the other; the nucleotides on one strand hydrogen-bond with complementary nucleotides on the opposite strand—that is, side-by-side with the 5′ end of one chain next to the 3′ end of the other. The purine and pyrimidine bases face the inside of the helix, with guanine always opposite cytosine and adenine always opposite thymine. The double helical "twist" occurs because of the angular geometry of each bonded nucleotide.
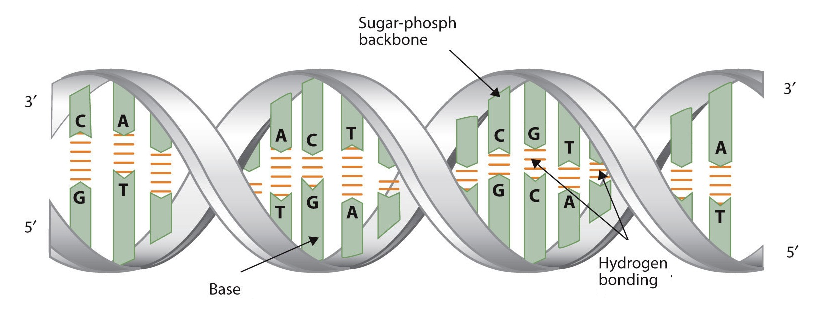
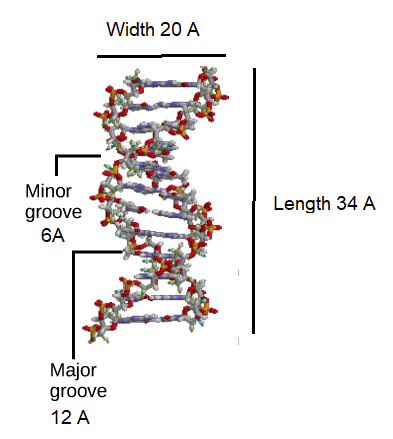
Initial work revealed the DNA polymer had a regular repeating pattern X-ray diffraction data shows that a repeating helical pattern is 20 Angstrom units wide and occurs every 34 Angstrom units with 10 nucleotide subunits per turn. Each subunit occupies 3.4 Angstrom units which is the same amount of space occupied by a single nucleotide unit. The helix is Under most conditions, the two strands are slightly offset, which creates a 12 Angstrom major groove on one face of the double helix, and a 6 Angstrom minor groove on the other. The overall DNA polymer varies in length (number of sugar-phosphate units connected), base composition (how many of each set of bases) and sequence (the order of the bases in the backbone).
What do we mean when we say information is encoded in the DNA molecule? An organism’s DNA can be compared to a book containing directions for assembling a model airplane or for knitting a sweater. Letters of the alphabet are arranged into words, and these words direct the individual to perform certain operations with specific materials. If all the directions are followed correctly, a model airplane or sweater is produced.
In DNA, the particular sequences of nucleotides along the chains encode the directions for building an organism. Just as saw means one thing in English and was means another, the sequence of bases CGT means one thing, and TGC means something different. Although there are only four letters—the four nucleotides—in the genetic code of DNA, their sequencing along the DNA strands can vary so widely that information storage is essentially unlimited.
Deoxyribonucleic acid (DNA) stores genetic information, while ribonucleic acid (RNA) is responsible for transmitting or expressing genetic information by directing the synthesis of thousands of proteins found in living organisms. But how do the nucleic acids perform these functions?
Three processes are required:
- Replication, in which new copies of DNA are made.
- Transcription, in which a segment of DNA is used to produce RNA.
- Translation, in which the information in RNA is translated into a protein sequence.
Exercise \(\PageIndex{1}\)
For this short DNA segment,
- Identify the 5′ end and the 3′ end of the molecule.
- Circle the atoms that comprise the backbone of the nucleic acid chain.
- Write the nucleotide sequence of this DNA segment.
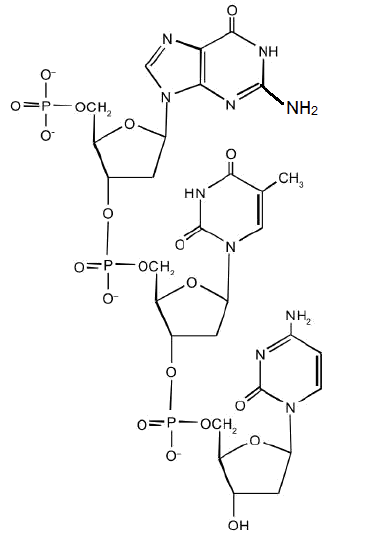
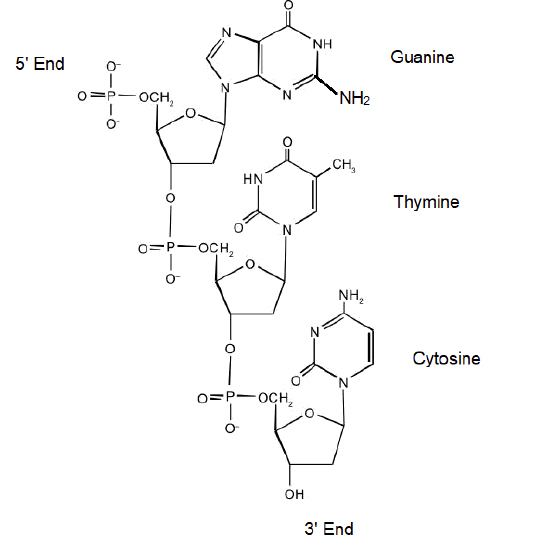
- Answer
-
Exercise \(\PageIndex{2}\)
Which nitrogenous base in DNA pairs with each listed nitrogenous base?
- Cytosine
- Adenine
- Guanine
- Thymine
- Answer
-
- Guanine
- Thymine
- Cytosine
- Adenine
Exercise \(\PageIndex{3}\)
How many hydrogen bonds can form between the two strands in the short DNA segment shown below?
5′ ATGCGACTA 3′ 3′ TACGCTGAT 5′
- Answer
-
22 (2 between each AT base pair and 3 between each GC base pair).
Exercise \(\PageIndex{4}\)
A segment of one strand from a DNA molecule has the sequence 5′-TCCATGAGTTGA-3′. What is the sequence of nucleotides in the opposite, or complementary, DNA chain?
- Answer
-
Knowing that the two strands are antiparallel and that T base pairs with A, while C base pairs with G, the sequence of the complementary strand will be 3′-AGGTACTCAACT-5′ (can also be written as TCAACTCATGGA).

