1.3: Mass Spectrometry additional details
- Page ID
- 358959
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Terms for mass spectra interpretation
- Molecular ion (M.+): If the molecular ion appears, it will be the highest mass in an EI spectrum (except for isotope peaks discussed below). This peak will represent the molecular weight of the compound. Its appearance depends on the stability of the compound. Double bonds, cyclic structures and aromatic rings stabilize the molecular ion and increase the probability of its appearance.
- Reference Spectra: Mass spectral patterns are reproducible. The mass spectra of many compounds have been published and may be used to identify unknowns. Instrument computers generally contain spectral libraries which can be searched for matches.
- Fragmentation: General rules of fragmentation exist and are helpful to predict or interpret the fragmentation pattern produced by a compound. Functional groups and overall structure determine how some portions of molecules will resist fragmenting, while other portions will fragment easily. A detailed discussion of those rules is beyond the scope of this introduction, and further information may be found in your organic textbook or in mass spectrometry reference books. A few brief examples by functional group are described (see Fragmentation Patterns).
- Isotopes: Isotopes occur in compounds analyzed by mass spectrometry in the same abundances that they occur in nature. A few of the isotopes commonly encountered in the analyses of organic compounds are below along with an example of how they can aid in peak identification.
Relative Isotope Abundance of Common Elements
| Element | Isotope | Relative Abundance |
Isotope | Relative Abundance |
Isotope | Relative Abundance |
|---|---|---|---|---|---|---|
| Carbon | 12C | 100 | 13C | 1.11 | ||
| Hydrogen | 1H | 100 | 2H | 0.016 | ||
| Nitrogen | 14N | 100 | 15N | 0.38 | ||
| Oxygen | 16O | 100 | 17O | 0.04 | 18O | 0.20 |
| Sulfur | 32S | 100 | 33S | 0.78 | 34S | 4.40 |
| Chlorine | 35Cl | 100 | 37Cl | 32.5 | ||
| Bromine | 79Br | 100 | 81Br | 98.0 |
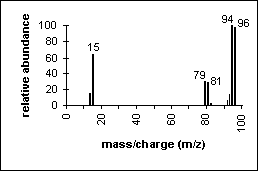
Methyl Bromide: An example of how isotopes can aid in peak identification.
 |
|
The ratio of peaks containing 79Br and its isotope 81Br (100/98) confirms the presence of bromine in the compound.
Other ionization methods
An array of ionization methods and mass analyzers are available to meet the needs of many types of chemical analysis. A few are listed here with a highlight of their usefulness.
Sample introduction/ionization methods
| Ionization method |
Typical Analytes |
Sample Introduction |
Mass Range |
Method Highlights |
|---|---|---|---|---|
| Electron Impact (EI) | Relatively small volatile |
GC or liquid/solid probe |
to 1,000 Daltons |
Hard method versatile provides structure info |
| Chemical Ionization (CI) | Relatively small volatile |
GC or liquid/solid probe |
to 1,000 Daltons |
Soft method molecular ion peak [M+H]+ |
| Electrospray (ESI) | Peptides Proteins nonvolatile |
Liquid Chromatography or syringe |
to 200,000 Daltons |
Soft method ions often multiply charged |
| Fast Atom Bombardment (FAB) | Carbohydrates Organometallics Peptides nonvolatile |
Sample mixed in viscous matrix |
to 6,000 Daltons |
Soft method but harder than ESI or MALDI |
| Matrix Assisted Laser Desorption (MALDI) | Peptides Proteins Nucleotides |
Sample mixed in solid matrix |
to 500,000 Daltons |
Soft method very high mass |
Outside Links
- A library of spectra can be found in the NIST WebBook, a data collection of the National Institute of Standards and Technology.
- Useful tools such as an exact mass calculator and a spectrum generator can be found in the MS Tools section of Scientific Instrument Services webpage.
- The JEOL Mass Spectrometry website contains tutorials, reference data and links to other sites.
- More general information and tutorials can be found in Scimedia, an educational resource.
- At the University of Arizona, the Wysocki Research Group studies surface-induced dissociation (SID) tandem mass spectrometry.
- Many more interesting and useful links can be found by following the site links in the above references.
Contributors and Attributions
Dr. Linda Breci, Associate Director Arizona Proteomics Consortium University of Arizona