15: Biomaterials- Protein Structure Determination by FTIR Spectroscopy
- Page ID
- 73007
CHEM 174 Physical Chemistry Laboratory II
Biomaterials: Protein Structure Determination by FTIR Spectroscopy
Introduction
The goal of this research is to investigate the effects of environmental stressors on proteins. Proteins are among the most important structural components of living cells and they are biological polymers composed of amino acids. An amino acid has both an amine group (-NH2) and a carboxyl group (-COOH). A generic representation of an amino acid and the simplest of the series, glycine, are shown below.
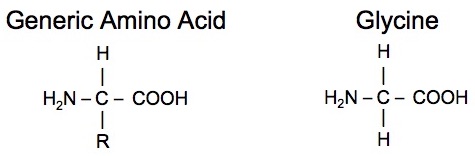
There are many different possibilities for the R-group and there are a total of 20 amino acids that are commonly found in proteins. Note that except for glycine, the central carbon is a chiral-center as in for example the amino acid phenylaniline.
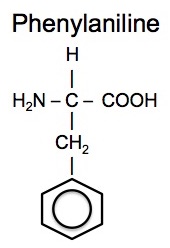
Amino acids polymerize by reactions of the amine group of one amino acid with the carboxyl acid group of another.
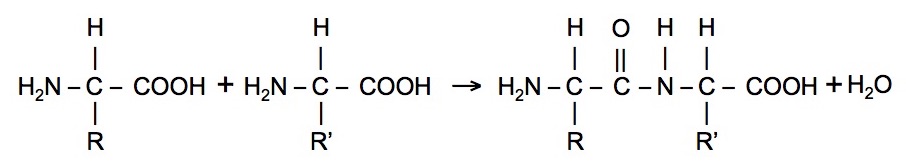
This process is repeated many times to construct a protein.
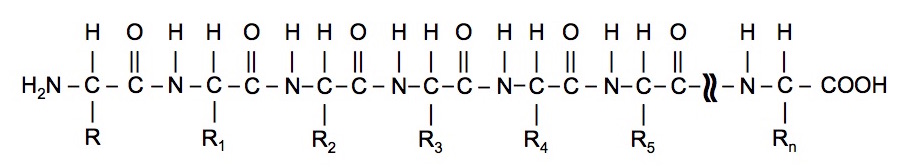
The properties of the protein are determined by the nature of the R groups and their interactions through forces such as hydrogen bonding.
Molecules vibrate, rotate and stretch at characteristic frequencies when they absorb infrared radiation. These frequencies are determined by structure of the molecule. We have used infrared spectroscopy to examine protein structure and the effects toxic compounds on proteins.
Procedure
Fourier transform infrared spectroscopy will be used in a total internal reflectance mode in this study. An advantage of total internal reflectance is that a solution containing a sample only needed to be placed on top of an infrared window. Problems with the large infrared absorption by water are avoided and no cells to contain the solutions are required.
Record an FT-IR spectrum of the simplest amino acid, glycine, to see the infrared absorption due to the amino and carboxyl acid groups. Record spectra of both glycine crystals and an aqueous-phase solution.
Next record infrared spectra of solid-phase gelatin and of a gelatin solution. Gelatin is composed of the protein collagen. Mix the gelatin solution with methanol; a white precipitate should result showing that the protein was denatured. Record infrared spectra of the precipitate and also record an FT-IR of methanol for comparison.
Summary of procedure:
Part A: Take FT-IR spectra of:
- Glycine in solid phase.
- Collagen protein in solid phase.
- Glycine in aqueous phase.
- Collagen in aqueous phase.
- Denature the collagen by adding a small amount of methanol. Take FT-IR spectra of the precipitate. Take an FT-IR of methanol
Some Sample IR Spectra
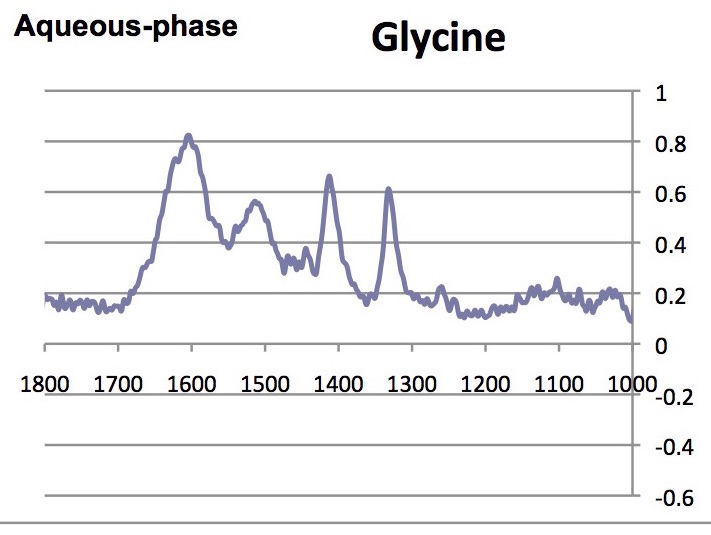
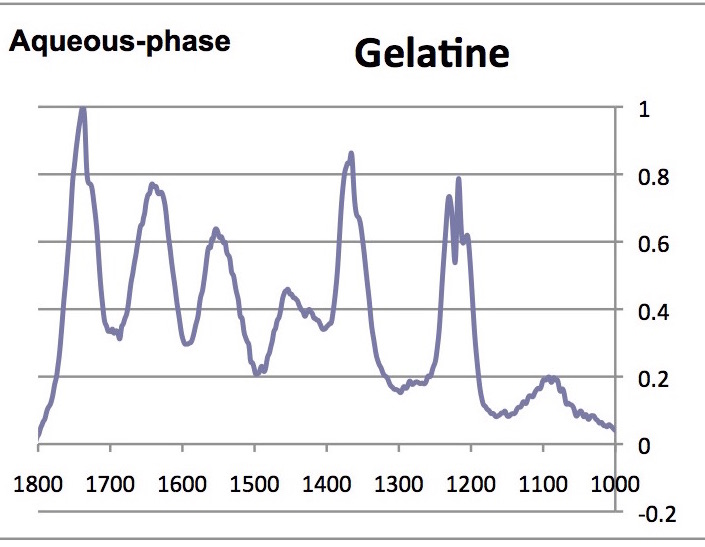
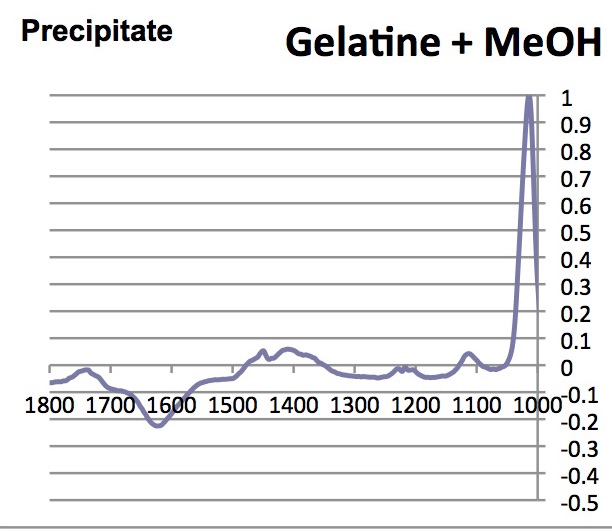
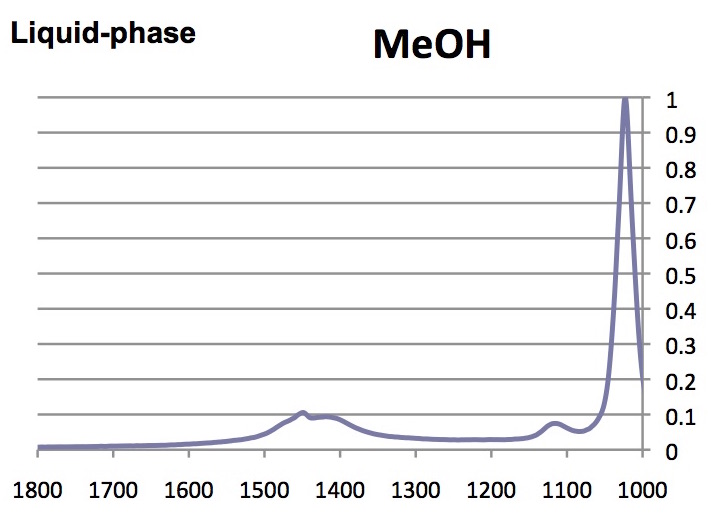
Note that the absorbance of the highest peak in the complete spectrum was normalized to 1.00; below only the range between 1000 and 1800 cm-1 are shown.
Report
- Use the research journal article format and discuss your results by answering the following questions in your report:
- What are the similarities and differences between the spectra of glycine, collagen and the denatured participate?
- Are there any differences between the solid and aqueous phase spectra?
- What can you say about the relationship between the protein’s functional groups and the absorbance peaks in the spectra? You may need to do some literature research to answer this question.
References
Nielson, D.L. and M.M. Cox, “Lehninger Principles of Biochemistry” 4ed., Freeman, 2005.
Alberts, B., A. Johnson, J. Lewis, M. Raff, K. Roberts and P. Walter, “Molecular Biology of The Cell” 5ed., Garland Science, 2008.

