In-Class Exercises – Chromatograms and Mass Spectra
- Page ID
- 120134
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Learning Objectives
After completing these exercises, you will be able to…
(1) Explain the relationship between a total ion chromatogram (TIC), an extracted (or selected) ion chromatogram (XIC), and a tandem (MS/MS) mass spectrum.
(2) Interpret GC-MS and LC-MS/MS data and predict/explain simple fragmentation patterns
Exercise 1
Within your group of three, assign and distribute a different color (RED, BLUE, YELLOW) “Duplo” molecule to each person.
In the table below, record the number of individual blocks that make up each color. These will represent the molecular masses of your group’s three molecules.
|
Molecular Mass |
|
|
RED |
|
|
BLUE |
|
|
YELLOW |
- Predict the elution order of the three molecules following GC chromatographic separation. Assume that RED has the lowest boiling point and YELLOW has the highest.
Elution order: 1st_________________ 2nd________________ 3rd_______________
- What logic could you use to predict how each Duplo molecule would break apart (“fragment”) if dropped on the ground?
- How would this same logic apply to the fragmentation of actual molecules during GC-MS? (In other words, why do molecules break apart in certain places and not in others?)
- Fill in the table below with your predictions for the “masses” (# of blocks) of the likely fragments of each molecule. [Note: each molecule does not necessarily need to form five different mass fragments.]
|
Fragment 1 |
Fragment 2 |
Fragment 3 |
Fragment 4 |
Fragment 5 |
||
|
Molecular Mass |
Mass |
Mass |
Mass |
Mass |
Mass |
|
|
RED |
||||||
|
BLUE |
||||||
|
YELLOW |
Once your group has agreed on an elution order and predicted fragment sizes, bring your intact molecules up to the front of the class and place them in the correct order. If another group has a different elution order than you, discuss your reasoning and come to a consensus.
Now let’s look at how each molecule fragments in the mass spectrometer.
- Will every fragment be detected?
- “Each fragment is formed by the breakdown of the next largest fragment.”
- Describe how you would plot the total ion chromatogram (TIC) for this example and explain which ions contribute to the TIC signal.
- Describe how you would plot an extracted ion chromatogram (XIC) for this example.
Exercise 2
(Adapted from Bullen, Fitch, Kelly, and Larive, 2013, Environmental Analysis – Lake Nakuru Flamingos: Pesticides, Analytical Sciences Digital Library, Online.)
Mass spectrometry is widely used in analytical chemistry, forensics, bioanalysis, and environmental analysis. Some advantages of mass spectrometry include its sensitivity and low detection limits. Both qualitative and quantitative results can be obtained using a mass spectrometer. However, mass spectrometers are generally more costly than electrochemical and spectroscopic instruments.
A total ion chromatogram (TIC) was collected by GC-MS for a standard solution containing a mixture of 20 organochlorine pesticides:
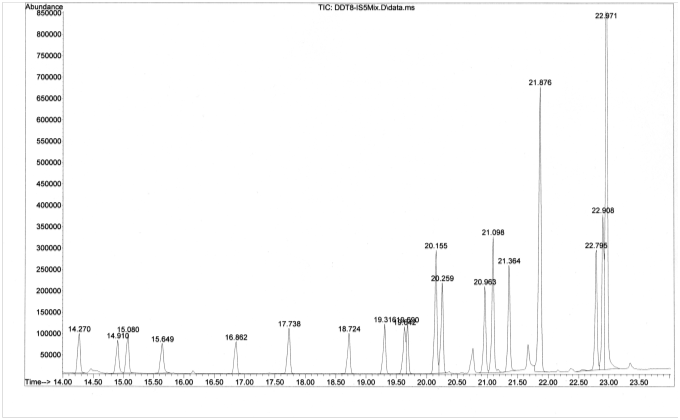
Consider the chromatographic peak at 21.876 min corresponding to the insecticide, DDT.
- List three factors responsible for the broadness of the peak.
- What does the y-axis label “Abundance” refer to?
- What has to happen for a DDT molecule to create a detector response?
- Explain why neutral molecules are not detected in MS.
To help confirm the identity of the peak at 21.876 min as DDT, it would be common to compare a mass spectrum (m/z vs. abundance) from the middle portion of the peak to a reference spectrum for DDT, taken under similar ionization conditions. A useful source for reference EI mass spectra is the NIST Chemistry WebBook.
The NIST mass spectrum (GC-MS; EI; 70eV) of DDT (MW = 354.5 g/mol) is given below.
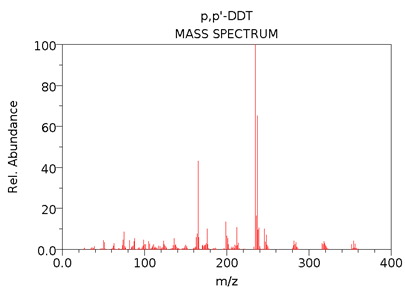
- Predict how the mass spectrum of DDT-d8 would differ from the mass spectrum above. (DDT-d8 is an isotopically labeled DDT in which 8 hydrogen atoms have been replaced by 8 deuterium atoms.)
We often use internal standards to correct for instrument variability and losses during sample processing. Ideally, the internal standard is an isotopically labeled version of the analyte of interest. For example, if you were analyzing a water sample for DDT concentration, you might inject a known amount of DDT-d8 at the time of collection, then prepare your sample (e.g., extract, concentrate, clean up) for injection on the GC-MS.
- Consider the TIC for a water sample extract that contained DDT as well as the internal standard DDT-d8 that you added. Is there a problem? If so, how would you solve it?
Exercise 3
In 2010, the Deep Water Horizon oil spill released 5 million barrels of crude oil into the bottom waters of the Gulf of Mexico. One strategy to minimize surface slicks was to release chemical dispersants (i.e., surfactants) into the water by plane or into the deep water near the wellhead, 5000 feet below the surface. This was the first time that dispersants were released into deep water on a large scale (2.1 million gallons of dispersants), so there was considerable uncertainty about what would happen to the dispersant chemicals over time. The following figure was taken from a paper titled “Fate of Dispersants Associated with the Deepwater Horizon Oil Spill” by Elizabeth Kujawinski et al. (2011).
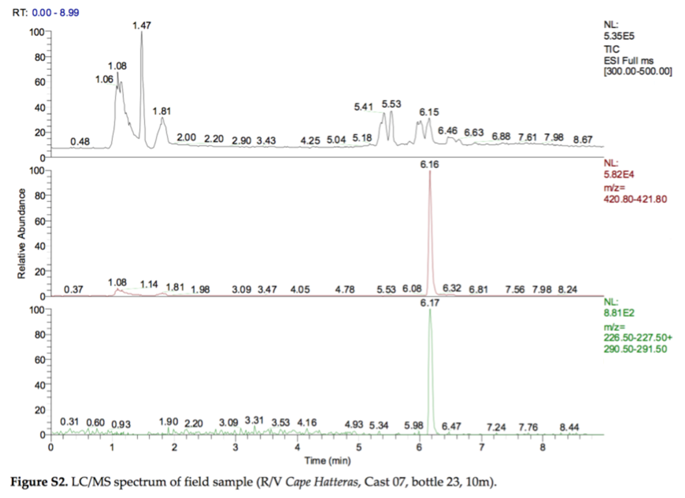
- Have each person in your group pick a panel in the figure, take a couple minutes to study and annotate your panel (referring to other panels as needed), then take turns describing your panel to the other members of your group. In your explanations be sure to include a full description of the type of data plotted, the axes, and the meaning of the information on the right side of your panel. [Bonus: can you find the error in the figure caption?]
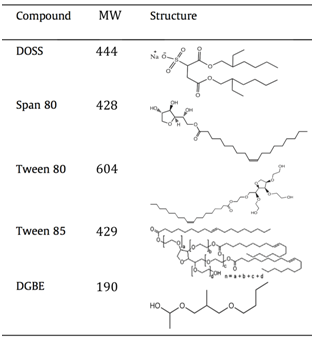
- The table above (modified from Glover et al., 2014, Chemosphere 111: 596-602) lists some of the components of Corexit, a dispersant mixture used during the Deep Water Horizon oil spill. Assuming the LC/MS analysis by Kujawinski et al. utilized electrospray ionization in negative mode, which component did they detect in their field sample? Explain your reasoning.
- How do you think Kujawinski et al. quantified the amount of DOSS in their field sample?
In fact, the data from Figure S2 were collected using a liquid chromatograph coupled to an ion trap mass spectrometer operated in MS/MS mode. In this mode, the mass spectrometer uses the mass analyzer (an ion trap) to isolate ions of one particular m/z, fragment them, then sort/detect the resulting fragment m/z values. In this way Kujawinski et al. were able to create an MS/MS spectrum (not shown), which plots the m/z values for the fragments of the selected ion. The bottom panel in Figure S2 shows the XIC for two characteristic fragments (m/z 227 and m/z 291) of the molecular ion m/z 421.
- Describe a situation in which it would be advantageous to use an MS/MS (“tandem”) instrument? Explain your reasoning.


