14.5: Stage 1 - Digestion of Proteins
- Page ID
- 521938
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)- List the steps of protein digestion.
Protein digestion begins in the stomach (Figure \(\PageIndex{1}\)), where the action of gastric juice hydrolyzes about 10% of the peptide bonds. Gastric juice is a mixture of water (more than 99%), inorganic ions, hydrochloric acid, and various enzymes and other proteins.

The hydrochloric acid (HCl) in gastric juice is secreted by glands in the stomach lining. The pH of freshly secreted gastric juice is about 1.0, but the contents of the stomach may raise the pH to between 1.5 and 2.5. HCl(aq) helps to denature food proteins; that is, it unfolds the protein molecules to expose their chains to more efficient enzyme action.

Hydrolysis of Proteins
Proteases (digestive enzymes) are involved in digesting long protein chains into shorter fragments by splitting the peptide bonds that link amino acid residues. Some detach the terminal amino acids from the protein chain (exopeptidases, such as aminopeptidases, carboxypeptidase A); others attack internal peptide bonds of a protein (endopeptidases, such as trypsin, chymotrypsin, pepsin, papain, elastase).

The principal digestive enzyme precursor of gastric juice is pepsinogen, an inactive enzyme produced in cells located in the stomach wall. When food enters the stomach after a period of fasting, pepsinogen is converted to its active form—pepsin—in a series of steps initiated by the drop in pH. Pepsin catalyzes the hydrolysis of peptide linkages within protein molecules. It has a fairly broad specificity but acts preferentially on linkages involving the aromatic amino acids tryptophan, tyrosine, and phenylalanine, as well as methionine and leucine.
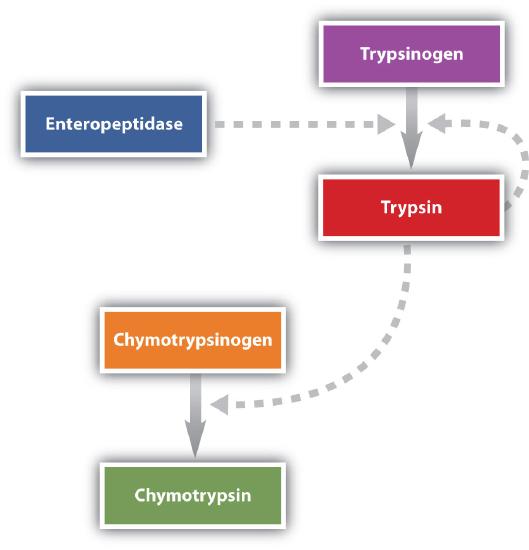
Protein digestion is completed in the small intestine. Pancreatic juice, carried from the pancreas via the pancreatic duct, contains inactive enzymes such as trypsinogen and chymotrypsinogen. They are activated in the small intestine as follows (Figure \(\PageIndex{5}\)): The intestinal mucosal cells secrete the proteolytic enzyme enteropeptidase, which converts trypsinogen to trypsin; trypsin then activates chymotrypsinogen to chymotrypsin (and also completes the activation of trypsinogen). Both of these active enzymes catalyze the hydrolysis of peptide bonds in protein chains. Chymotrypsin preferentially attacks peptide bonds involving the carboxyl groups of the aromatic amino acids (phenylalanine, tryptophan, and tyrosine). Trypsin attacks peptide bonds involving the carboxyl groups of the basic amino acids (lysine and arginine). Pancreatic juice also contains procarboxypeptidase, which is cleaved by trypsin to carboxypeptidase. The latter is an enzyme that catalyzes the hydrolysis of peptide linkages at the free carboxyl end of the peptide chain, resulting in the stepwise liberation of free amino acids from the carboxyl end of the polypeptide.
Aminopeptidases in the intestinal juice remove amino acids from the N-terminal end of peptides and proteins possessing a free amino group. Figure \(\PageIndex{3}\) illustrates the specificity of these protein-digesting enzymes. The amino acids that are released by protein digestion are absorbed across the intestinal wall into the circulatory system, where they can be used for protein synthesis.
Cleavage Sites
Here is a short list of proteases and the cleavage site in the protein substrate:
- Thrombin – Cleaves preferentially between the Arg and Gly residues. Off-target cleavage can occur at non-specific sites, normally from contaminating proteases. To ensure maximal protein integrity, the enzyme reagent must be very pure.
- Trypsin – Cleaves peptides on the C-terminal side of lysine and arginine amino acid residues. If a proline is on the carboxyl side of the cleavage site, the cleavage will not occur.
- Chymotrypsin – Cleaves at the C-terminus of the aromatic amino acids such as phenylalanine, tryptophan, and tyrosine. The specificity for aromatic side chains is because of the protein’s hydrophobic pocket.
- Lys-N (Endoproteinase) – Endoproteinase LysN cleaves at the amino-terminus of lysine residues (which includes methylated lysines).
- Carboxypeptidase B – hydrolyzes at the C-terminus of arginine or lysine.
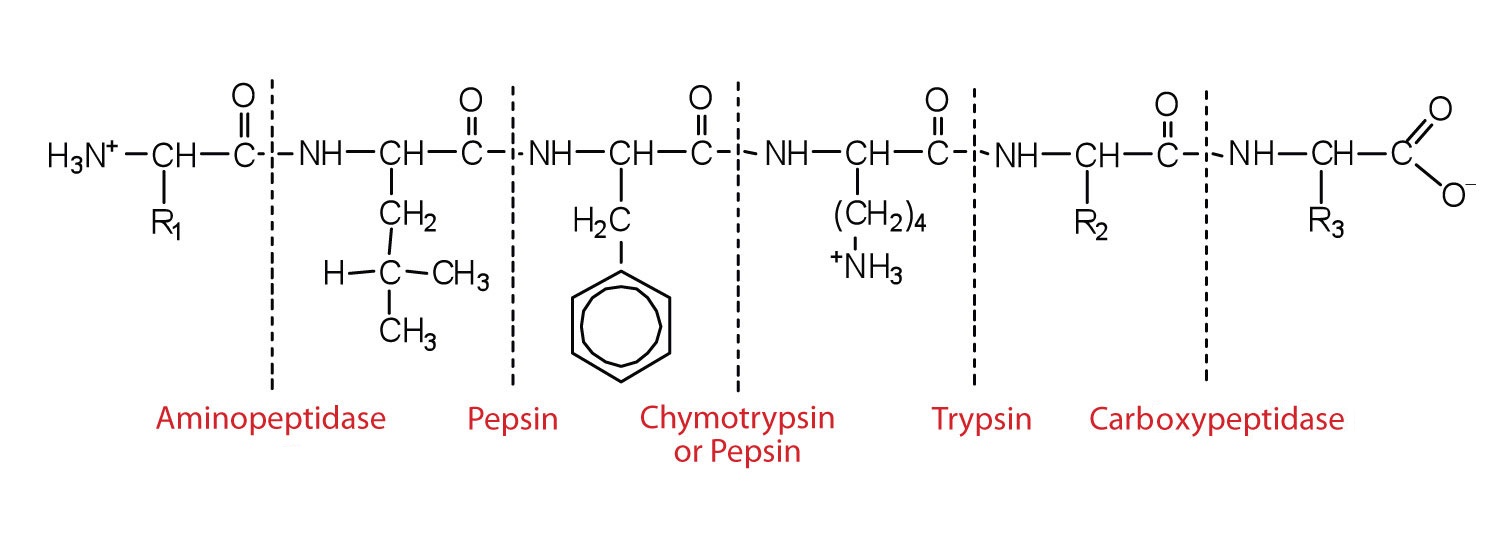
This diagram illustrates where in a peptide the different peptidases we have discussed would catalyze the hydrolysis of the peptide bonds.
Let’s use oxytocin (made of nine amino acids) as an example to illustrate the use of partial hydrolysis as a simple example of how a structure can be pieced together from fragments. Acid-catalyzed partial hydrolysis of oxytocin (after cleavage of the disulfide bridge) gives a mixture that includes the following peptides:
Ile-Gln-Asn-Cys; Gln-Asn-Cys-Pro; Pro-Leu-Gly; Cys-Tyr-Ile-Gln-Asn; and Cys-Pro-Leu-Gly.
Solution
When we match the overlapping regions of these fragments, the complete sequence of oxytocin appears:
Cys-Tyr-Ile-Gln-Asn
Ile-Gln-Asn-Cys
Gln-Asn-Cys-Pro
Cys-Pro-Leu-Gly
Pro-Leu-Gly
Complete structure: Cys-Tyr-Ile-Gln-Asn-Cys-Pro-Leu-Gly
Show where trypsin and chymotrypsin would cleave the following peptide.
Tyr-Ile-Gln-Arg-Leu-Gly-Phe-Lys-Asn-Trp-Phe-Gly-Ala-Lys-Gly-Arg-Gln-Gln
Solution
Trypsin cleaves peptides on the C-terminal side of lysine and arginine amino acid residues. If a proline is on the carboxyl side of the cleavage site, the cleavage will not occur. It will provide the following fragments:
Tyr-Ile-Gln-Arg-Leu-Gly-Phe-Lys; Asn-Trp-Phe-Gly-Ala-Lys-Gly-Arg; Gln-Gln
Chymotrypsin cleaves at the C-terminus of the aromatic amino acids such as phenylalanine, tryptophan, and tyrosine. It will provide the following fragments:
Tyr; Ile-Gln-Arg-Leu-Gly-Phe; Lys-Asn-Trp; Phe-Gly-Ala-Lys-Gly-Arg-Gln-Gln
Summary
During digestion, the peptide bonds in proteins undergo hydrolysis, breaking down into amino acids. The digestion process of proteins begins in the stomach and continues in the small intestine.


