4: My First Package
- Page ID
- 434896
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)In this activity we are going to make a package (ChemPack1) that contains two modules, ChemName and ChemInformatics. ChemName will use the random module to randomly return the name of a chemical from a text file. The second, ChemInformatics will obtain the molar mass from PubChem for that chemical. We will then write a script (MainScript.py) that asks the user to input the mass of a randomly generated chemical name and returns the moles within that mass.
Directory Structure
First create a folder for your activity, we will call it Chem_Package_Tutorial and within create a python program called mainScript.py and a folder called ChemPack1. The ChemPack1 folder will contain a __init__.py file, which tells python that it is a package.
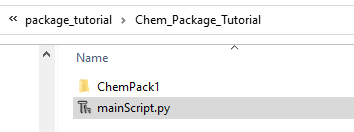 Figure \(\PageIndex{1}\): Copy and Paste Caption here. (Copyright; author via source)
Figure \(\PageIndex{1}\): Copy and Paste Caption here. (Copyright; author via source)Within ChemPack1 you will create two subfolders (Cheminformatics and ChemName), each of which in turn contain a __init__.py file, and so they are "subpackages" within the ChemPack1 package. The "objects" you are creating are actually in these packages.
 Figure \(\PageIndex{1}\): Copy and Paste Caption here. (Copyright; author via source)
Figure \(\PageIndex{1}\): Copy and Paste Caption here. (Copyright; author via source)
ChemName
ChemName is a package because it has an __init__.py file and it contains the program chemName.py that uses the random module to generate the random named chemical from the file chemName.txt. So you need to add this file to your package (folder). We have attached a sample file here (you need to right click, "save link as", and save it in the ChemName folder).
 Figure \(\PageIndex{1}\): Copy and Paste Caption here. (Copyright; author via source)
Figure \(\PageIndex{1}\): Copy and Paste Caption here. (Copyright; author via source)
chemName.py
chemName is a program that defines a function called getName().
import random
import os
path = os.path.abspath(__file__)#get absolute path
fileName='\\chemName.txt' #name of the .txt file
sPath = path.split('\\') #split the string
rPath = sPath[:-1] #remove the last item (name of the .py file)
jPath = '\\'.join(rPath) #join the list back into a string
newPath = jPath+fileName #join the path and the file name
x = int(random.randrange(0, 314, 1)) #generate a random number from 0-314
def getName():
with open (newPath, 'r' ) as fp:
chem=fp.readlines()[x]
chemName = chem.rstrip('\n')
fp.close()
return chemName
Because this is a package, this function is really a method, and can be called from another program using
chemName.getName()
Run the program chemName.py in Python. Nothing appears to happen, why?
- Answer
-
The function getName() gets a random name from the file chemnamt.txt and returns it, but does nothing else.
add two lines to the bottom of chemName.py that causes it to print the name of a randomly generated chemical (NOTE: comment out the lines after completing this activity)
- Answer
-
You can do this in several ways, but you need to call the function and then print the output. You can do this in two steps where in the first step you assign a variable name to the returned value, and then in the second step print that variable name, or you could embed the call statement into a print statement.
Cheminformatics.py
This sub-package (as denoted by the __init__.py file) contains the program ChemProperties.py This provides you to functions, one that gets a chemical property from the name of a molecule from the PubChem API. The second that uses the first to get the chemical property of molar mass to get the molar mass from PubChem and then determine the moles in a given mass of a sample
import requests
def getProperty(name, chemProperty):
uri = ("https://pubchem.ncbi.nlm.nih.gov/rest/pug/compound/name/"
+ str(name) +"/property/"+ str(chemProperty) + "/txt")
#print (uri)
res = requests.get(uri)
chemProperty=float(res.text.rstrip())
#moles = mass/molar_mass
return chemProperty
def calcMoles(name, mass):
molarMass=getProperty(name, 'molecularweight')
moles = int(mass)/int(molarMass)
return moles
What three parameters do you need to define variables for to send as arguments (inputs) to the above functions?
Solution
name, chemProperty and mass
To the bottom of the above script write some code that asks for the mass of barium and using f strings prints out
The molar mass of barium is x g/mol and y g has z mole.
So if you input 4.0 g it prints
The molar mass of barium is 137.33 g/mol and 4.0 g has 0.0291 mole.
POUND OUT CODE BEFORE MOVING ON TO NEXT TASK
Solution
You need to set up 3 variables
name=barium
chemProperty = molecular weight
and you need an input statement to assign the value to the variable mass. Now, you can creat two new variables that you embed into the f string, or you can embed your functions into the f string.
mainScript.py
Place mainScript.py in the folder next to the ChemPack1 package (figure \(\PageIndex{1}\)
from ChemPack1.ChemInformatics import ChemProperties
from ChemPack1.ChemName import chemName
randomChemName = str(chemName.getName())
print ('Your Chemical is ' + randomChemName)
sample_weight = float(input("Weight of the {} sample in units of g: ".format(randomChemName)))
moles = ChemProperties.calcMoles(randomChemName, sample_weight)
print ('{}g {} = {:.3f}moles'.format(sample_weight, randomChemName, moles))
Contributors
The code for this activity was developed by Elena Waters

