13.3: Primary structure- peptides
- Page ID
- 435684
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
42
Harper College Chemistry Department
Amino acids and peptide bonds
Proteins are long chains of small ‘building block’ molecules called amino acids. There are twenty different amino acids generally found in proteins, all of which are based on the common structural framework shown below.
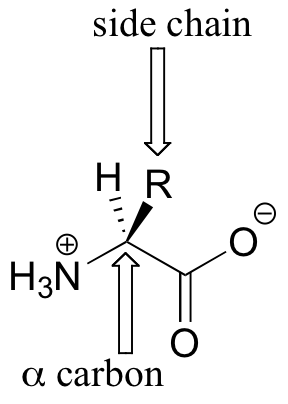
Amino acids contain an amino group on one side and a carboxylate group on the other, with a substituted carbon, referred to as the ‘alpha-carbon’, in the middle. It is the substituent on the alpha-carbon – designated ‘R’ in the general structure here, and often referred to as the amino acid ‘side chain’ – that distinguishes the 20 different amino acids. In all but one of the twenty, the alpha-carbon is a chiral center (in glycine, the ‘R’ group is simply a hydrogen, and thus the alpha-carbon is achiral). Virtually all of the amino acids found in proteins have the stereochemistry shown in the general figure above, and are referred to as L-amino acids (‘L’ stand for levorotary, the property of rotating plane polarized light in a counter-clockwise direction). Each amino acid is designated by both a three-letter and a single-letter abbreviation: the amino acid alanine, for example, is designated ‘Ala’ or just ‘A’.
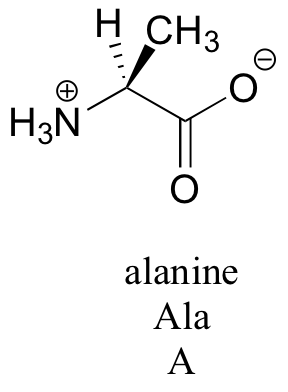
The side chains of amino acids contain a variety of functional groups (see table 5). Some like alanine have only alkyl functionality, while others have alcohol, thiol, sulfide, or amide groups. Four amino acids have aromatic groups on their side chains: phenylalanine has a phenyl group, tyrosine a phenol, tryptophan an indole, and histidine an imidazole. Many of the side chains are ‘ionizable’, capable of acting as acids or bases depending on their protonation state. This property makes them extremely important in enzyme catalysis. Aspartate and glutamate, for example, both have carboxylate groups and are deprotonated at pH 7. Lysine and arginine have amine and imino groups, respectively, and are protonated (and positively charged) at pH 7. Histidine, with its imidazole group, is mainly protonated and positively charged at pH 7. The phenol group of tyrosine is also capable of donating a proton. We will have more to say later about the pKa values of the different side chains in chapter 7. As you progress in your study of organic chemistry, it will become very helpful to familiarize yourself with the structures of all 20 amino acids.
Proteins are formed when amino acids are linked together by ‘peptide bonds’ to form long chains (a peptide bond is really just a specific type of amide functional group). When they are part of a protein chain, individual amino acids are often referred to as ‘residues’. The figure below shows a ‘dipeptide’ – two amino acids linked together by a peptide bond.
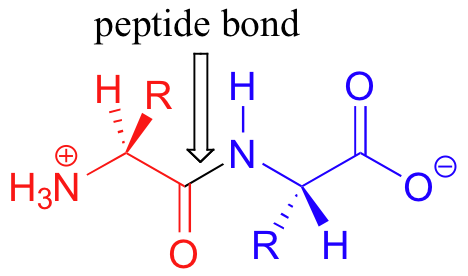
Many protein chains are several hundred amino acids long. The ‘backbone’, or ‘main chain’ of a protein refers to the repeating nitrogen – alpha-carbon – carbonyl pattern: in other words, everything but the side chain. As we have discussed previously, the peptide bond has significant double-bond character due to resonance, and therefore there is a considerable barrier to rotation. Other sigma bonds in proteins, however, have more rotational flexibility.

