4.5: Genes and Genomes
- Page ID
- 347428
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Source: BiochemFFA_7_1.pdf. The entire textbook is available for free from the authors at http://biochem.science.oregonstate.edu/content/biochemistry-free-and-easy

Introduction
For many years, scientists wondered about the nature of the information that directed the activities of cells. What kind of molecules carried the information, and how was the information passed on from one generation to the next? Key experiments, done between the 1920s and the 1950s, established convincingly that this genetic information was carried by DNA. In 1953, with the elucidation of the structure of DNA, it was possible to begin investigating how this information is passed on, and how it is used.
Genomes
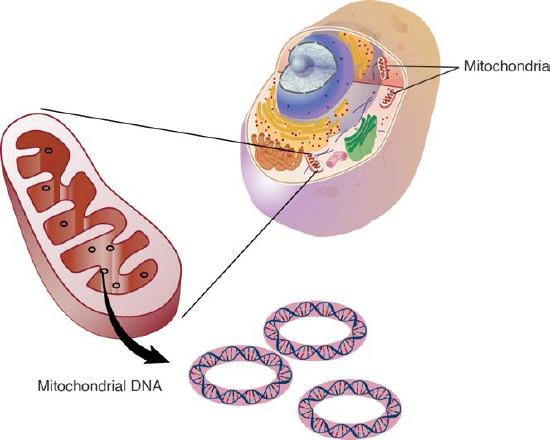
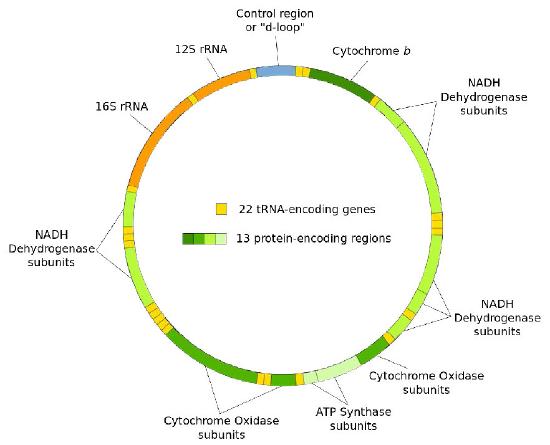
We use the word “genome” to describe all of the genetic material of the cell. That is, a genome is the entire sequence of nucleotides in the DNA that is in all of the chromosomes of a cell. When we use the term genome without further qualification, we are generally referring to the chromosomes in the nucleus of a eukaryotic cell. As you know, eukaryotic cells have organelles like mitochondria and chloroplasts that have their own DNA (Figure 7.1 & 7.2). These are referred to as the mitochondrial or chloroplast genomes to distinguish them from the nuclear genome.
Starting in the 1980s, scientists began to determine the complete sequence of the genomes of many organisms, in the hope of better understanding how the DNA sequence specifies cellular functions. Today, the complete genome sequences have been determined for thousands of species from all domains of life, and many more are in the process of being worked out by groups of scientists across the world.
Global genome initiative
The Global Genome Initiative, a collaborative effort to sequence at least one species from each of the 9,500 described invertebrate, vertebrate, and plant families is one of many such ventures. The information from these various efforts is collected in enormous online repositories, so that it is freely available to scientists. As the sequence databases compile ever more information, the fields of computational biology and bioinformatics have arisen, to analyze and organize the data in a way that helps biologists understand what the information in DNA means in the cellular context.
Genes
It has been known for many years that phenotypic traits are controlled by specific regions of the DNA that were termed “genes”. Thus, DNA was envisioned as a long string of nucleotides, in which certain regions, the genes, were separated by non-coding regions that were simply referred to as intergenic sequences (inter=between; genic=of genes). Early experiments in molecular biology suggested a simple relationship between the DNA sequence of a gene and its product, and led scientists to believe that each gene carried the information for a single protein. Changes, or mutations in the base sequence of a gene would be reflected in changes in the gene product, which in turn, would manifest itself in the phenotype or observable trait. This simple picture, while still useful, has been modified by subsequent discoveries that demonstrated that the use of genetic information by cells is somewhat more complicated. Our definition of a gene is also evolving to take new knowledge into consideration.
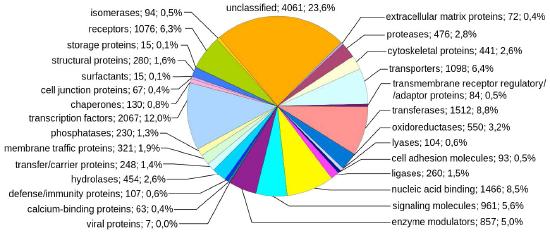
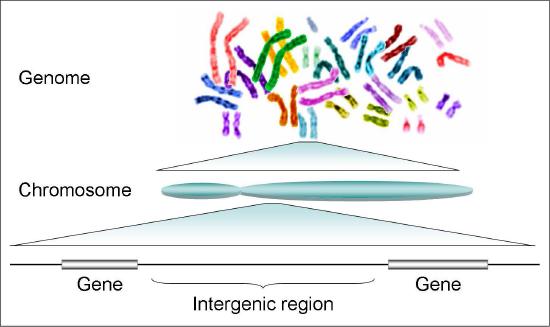
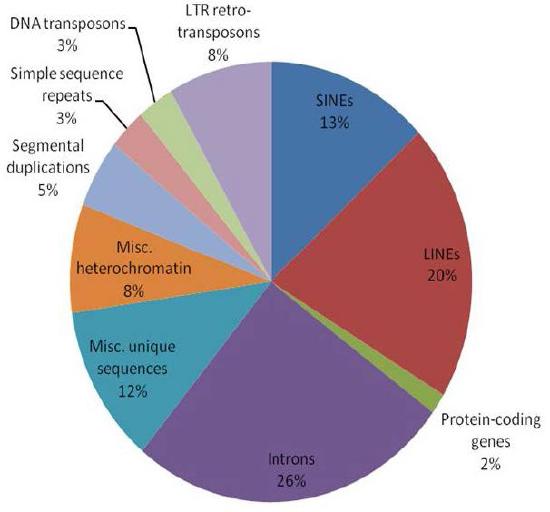
Figure 7.4 - Human genes sorted by class
Matters of size
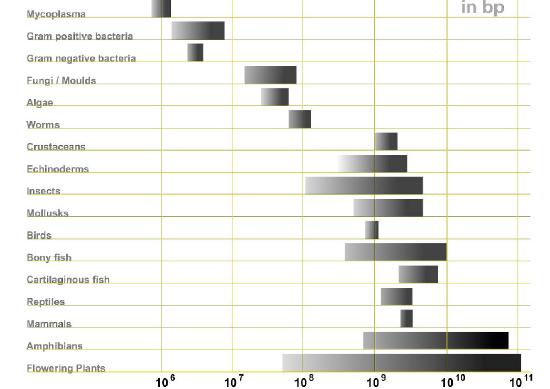
A common-sense assumption about genomes would be that if genes specify proteins, then the more proteins an organism made, the more genes it would need to have, and thus, the larger its genome would be. Comparison of various genomes shows, surprisingly, that there is not necessarily a direct relationship between the complexity of an organism and the size of its genome (Figure 7.5). To understand how this could be true, it is necessary to recognize that while genes are made up of DNA, all DNA does not consist of genes (for purposes of our discussion, we define a gene as a section of DNA that encodes an RNA or protein product). In the human genome, less than 2% of the total DNA seems to be the sort of coding sequence that directs the synthesis of proteins. For many years, non-coding DNA in genomes was believed to be useless, and was described as “junk DNA” although it was perplexing that there seemed to be so much “useless” sequence. Recent discoveries have, however, demonstrated that much of this so-called junk DNA may play important roles in evolution, as well as in regulation of gene expression.
Introns
So, what is all the non-coding DNA doing there? We know that even coding regions in our DNA are interrupted by non-coding sequences called introns. This is true of most eukaryotic genomes. An examination of genes in eukaryotes shows that non-coding intron sequences can be much longer than the coding sections of the gene, or exons. Most exons are relatively small, and code for fewer than a hundred amino acids, while introns can vary in size from several hundred base-pairs to many kilobase-pairs (thousands of base-pairs) in length. For many genes in humans, there is much more of intron sequence than coding (a.k.a. exon) sequence. Intron sequences account for roughly a quarter of the genome in humans.
Other non-coding sequences
What other kinds of non-coding sequences are there? One function for some DNA sequences that do not encode RNA or proteins is in specifying when and to what extent a gene is used, or expressed. Such regions of DNA are called regulatory regions and each gene has one or more regulatory sequences that control its expression. However, regulatory sequences do not account for all the rest of the DNA in our genomes, either.
Transposable sequences
Surprisingly, almost half of the human genome appears to consist of several kinds of repetitive sequences. Many of the repetitive sequences are known to be transposable elements (transposons), sections of DNA that can move around within the genome. Sometimes referred to as “jumping genes” these transposable elements can move from one chromosomal location to another, either through a simple “cut and paste” mechanism that cuts the sequence out of one region of the DNA and inserts it into another location, or through a process called retrotransposition involving an RNA intermediate.
LINES & SINES
There are millions of copies of each of two major classes of such transposable elements, the LINEs (Long Interspersed Elements) and SINEs (Short Interspersed Elements) in our genomes.
LINEs and SINEs are both a kind of transposable element called retrotransposons, sequences that are copied into RNA, then reverse transcribed back into DNA before being inserted into new locations. This movement is typically not sequence specific, meaning that the transposons can be inserted randomly in the genome, in many cases within coding regions. As might be expected, this can disrupt the function of the gene. Transposons may also insert within regulatory regions, and change the expression of the genes they control. As a major cause of mutation in genomes, transposons play an important role in evolution.
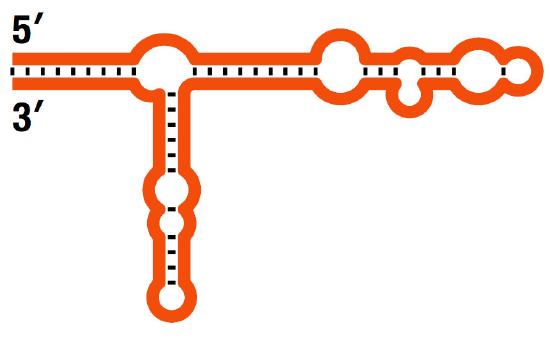
Finally, recent findings have shown that much of the genome is transcribed into RNAs, even though only about 2% encodes proteins. What are the RNAs that do not encode proteins? Ribosomal RNAs (Figure 7.7) and transfer RNAs, together with the small nuclear RNAs that function in splicing, account for some of these non-translated transcripts, but not all. The remaining RNAs are regulatory RNAs, small molecules that play an important role in regulating gene expression. As we understand more about genomes, it is becoming evident that the so-called “junk” DNA is anything but.