Section 1A. What is a protein?
- Page ID
- 81224
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
Proteins are biological macromolecules consisting of long chains of amino acids. A shorter chain of amino acids is called a peptide. Proteins and peptides are biological polymers formed from amino acid monomers. Each amino acid is composed of an amine group, a side chain (R), and a carboxylic acid functionality (Figure 1).
Figure 1. An amino acid.
There are 22 naturally occurring amino acids, 20 of which are encoded by the genome (Figure 2). These amino acids are usually grouped according to the character of their side chains, which may be acidic, basic, neutral, or hydrophobic. In a protein, portions of the linear sequence of amino acids may take on a secondary structure (such as a helix or a sheet) based on intermolecular forces between amino acid residues. The full-length protein will form a tertiary structure or overall shape, and the structure of the protein is intimately linked to the protein’s function.
Figure 2. Table of naturally occurring amino acids sorted by side chain and with their three-letter and one-letter codes. (Reproduced under a Creative Commons license from Dancojocari.)
1. In Figure 1 above, circle the amine group, draw a triangle around the side chain, and draw a square around the carboxylic acid group of the amino acid.
A. 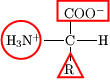
The linear chains of amino acids that form proteins are connected by amide (or peptide) bonds that form by a dehydration reaction between two amino acids (Figure 3). By convention, we write and draw peptides from the N-terminus (the side with the free amine group) to the C-terminus (the side with the free carboxylic acid).
Figure 3. Formation of a peptide bond between two amino acids.
2. In Figure 3 above, label the N-terminus and the C-terminus of the dipeptide product. Circle the amide bond.
A. 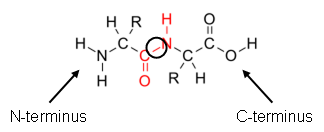
While many properties of peptides and proteins can be measured to provide useful information, we will focus on measurements of peptide and protein masses using mass spectrometry. Mass spectrometry measures the mass and charge of molecules in the gas phase. As a result, the mass of individual molecules is important. The monoisotopic mass of a molecule is the mass of that compound when it is composed of the most abundant isotope of each atom in the molecule. For example, if we calculate the monoisotopic mass of CO2, we will use the masses of carbon-12 and oxygen-16 since these are the most abundant isotopes of these elements. This gives us a monoisotopic mass of
12.000 amu + 2 (15.995 amu) = 43.99 amu.
This is different than the molar mass we would calculate using average atomic weights from the periodic table; on the periodic table, we find a molar mass for carbon of 12.011 amu. This represents the weighted average of all carbon atoms, which includes 98.9% carbon-12 and 1.1% carbon-13. Table 1 shows the monoisotopic mass of each amino acid and the mass of that amino acid as a residue in a protein or peptide chain.
| Amino Acid | Single-Letter Code | Residue MW (amu) | Amino Acid MW (amu) |
|---|---|---|---|
| glycine | G | 57.02 | 75.03 |
| alanine | A | 71.04 | 89.05 |
| serine | S | 87.03 | 105.04 |
| proline | P | 97.05 | 115.06 |
| valine | V | 99.07 | 117.08 |
| threonine | T | 101.05 | 119.06 |
| cysteine | C | 103.01 | 121.02 |
| isoleucine | I | 113.08 | 131.09 |
| leucine | L | 113.08 | 131.09 |
| asparagine | N | 114.04 | 132.05 |
| aspartic acid | D | 115.03 | 133.04 |
| glutamine | Q | 128.06 | 146.07 |
| lysine | K | 128.09 | 146.11 |
| glutamic acid | E | 129.04 | 147.05 |
| methionine | M | 131.04 | 149.05 |
| histidine | H | 137.06 | 155.07 |
| phenylalanine | F | 147.07 | 165.08 |
| arginine | R | 156.10 | 174.11 |
| tyrosine | Y | 163.06 | 181.07 |
| tryptophan | W | 186.08 | 204.09 |
1. Consider the data in Table 1. By what value do the residue molecular weight (MW) and the amino acid MW differ? Why is the MW of an amino acid residue in a peptide chain different from the mass of the full amino acid?
A. The mass of the amino acid residue is 18 amu lighter than the mass of the amino acid because the formation of peptide bonds at the N- and C-termini results in the loss of water from the amino acid.
2. Draw the structure for the tetrapeptide G, C, L, W. Refer to Figure 2 for the structure of amino acid side chains.
A.
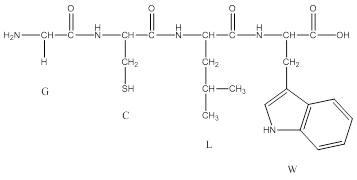
3. Calculate the monoisotopic molecular weight of the tetrapeptide using the data in Table 1.
A. When computing a peptide or protein molecular weight, sum up all residue masses and add 18.02 (for the H and OH at the termini):
57.02+103.01+113.08+186.08+18.01 = 477.20 amu
Note: In protein mass spectrometry, the Da is a common unit. It is the same as amu.
4. Calculate the molecular weight of the tetrapeptide using molar mass information from the periodic table. Why is this molecular weight different from the monoisotopic mass you calculate in question 3? Which mass is the mass measured in mass spectrometry?
A. Glycine is C2H5NO2. Cysteine is C3H7NO2S. Leucine is C6H13NO2. Tryptophan is C11H12N2O2.
This is a total of 22 carbons, 37 hydrogens, 5 nitrogens, 8 oxygens, and 1 sulfur atom. For the peptide, we must subtract the equivalent of 3 water molecules representing dehydryation to form the peptide bonds (i.e., total atom count should be less 6 hydrogens and 3 oxygens). This gives us 22 C, 31 H, 5 N, 5 O, and 1 S.
22 (12.011) + 31 (1.008) + 5 (14.007) + 5 (15.999) + 1 (32.06) = 477.58 Da.
The periodic table gives the weighted average molar masses of each element. This is the average mass of a population of GCLW peptides. No single GCLW peptide actually has this mass, and it is slightly heavier than the monoisotopic mass since the most abundant isotopes of H, C, N, and O are the lightest isotopes. The monoisotopic mass is the mass of a GCLW peptide composed of atoms that are the most abundant isotope of each element in the peptide. This will be the most intense peak on the mass spectrum.


