4.3: Additional Data Retrieval Approaches in PubChem
- Page ID
- 170161
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
Classification Browser
The PubChem Classification Browser, which allows the user to navigate or search PubChem records associated to a hierarchical classification system of interest, is available via URL:
http://pubchem.ncbi.nlm.nih.gov/classification
The Classification Browser can also be accessed from the PubChem home page (through the “Services” menu at the top or the “Classification” icon on the right column of the page). Currently, the Classification Browser can retrieve records annotated with terms in the following classification systems:
- MeSH (Medical Subject Headings)
- ChEBI
- FDA Pharmacological Classification
- KEGG
- LIPID MAPS
- World Health Organization (WHO)’s Anatomical Therapeutic Chemical (ATC)
- World Intellectual Property Organization (WIPO)’s IPC (International Patent Classification)
The Classification Browser provides a powerful way to quickly and visually find a desired subset of PubChem records. The output can be displayed in Tree view or List view.
An important feature of the Classification Browser is that the Table of Contents presented on the Compound Summary is integrated into the Classification Browser, allowing users to quickly retrieve compounds with a particular type of information available. For example, the figure below shows how to retrieve all compounds with the boiling point information from PubChem.
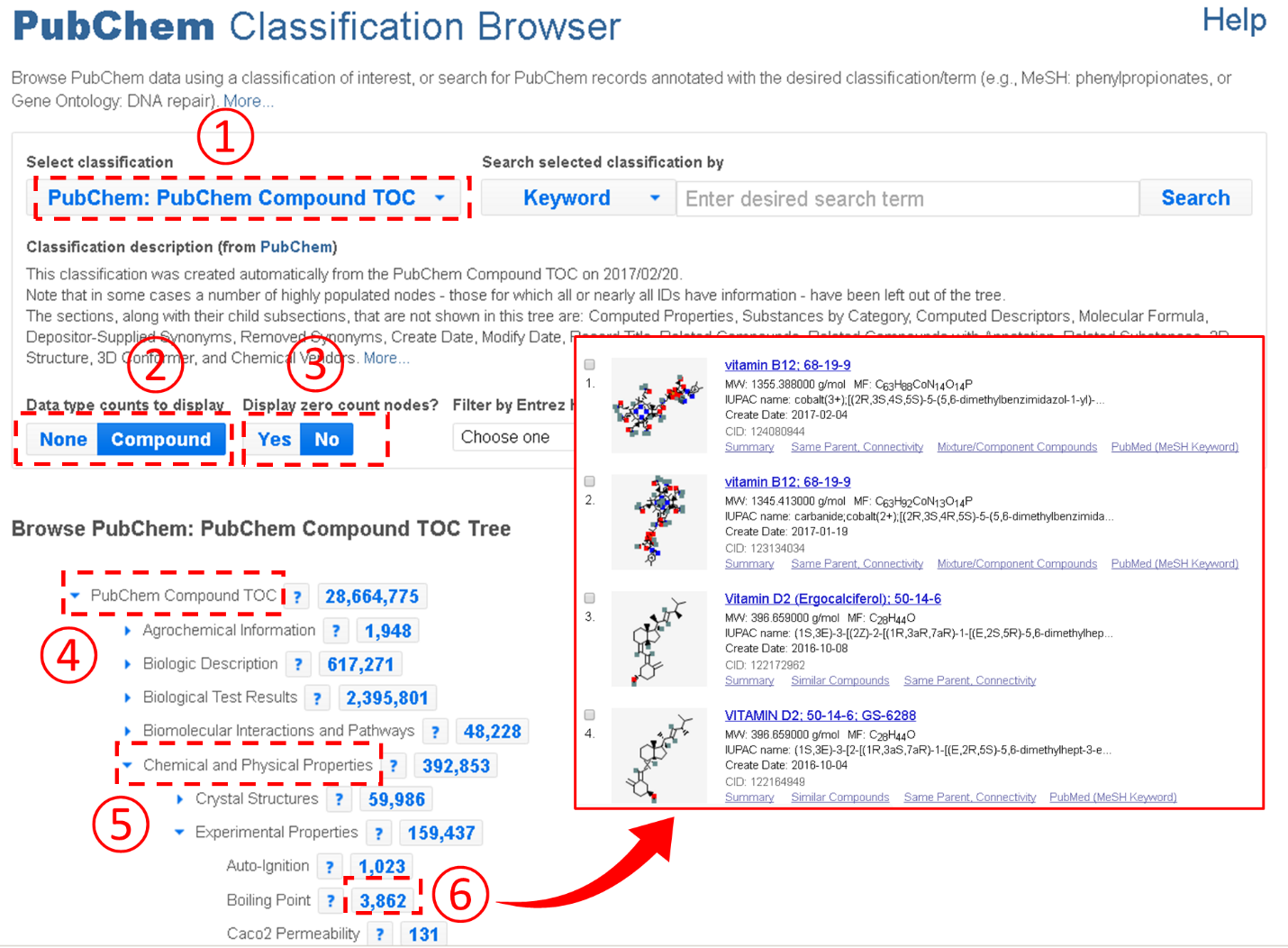
In the example above, users need to expand the Table of Contents tree to locate the boiling point node. However, this task may not be easy to some users who do not have prior knowledge about where the node that they want to find is located in the Table of Contents tree system. To assist these users, the Classification Brower supports a keyword search against the node names and descriptions of the classification trees. For example, the example below shows how to retrieve compounds with the CAS Registry number. Note that this task involves a search for the term “CAS”.
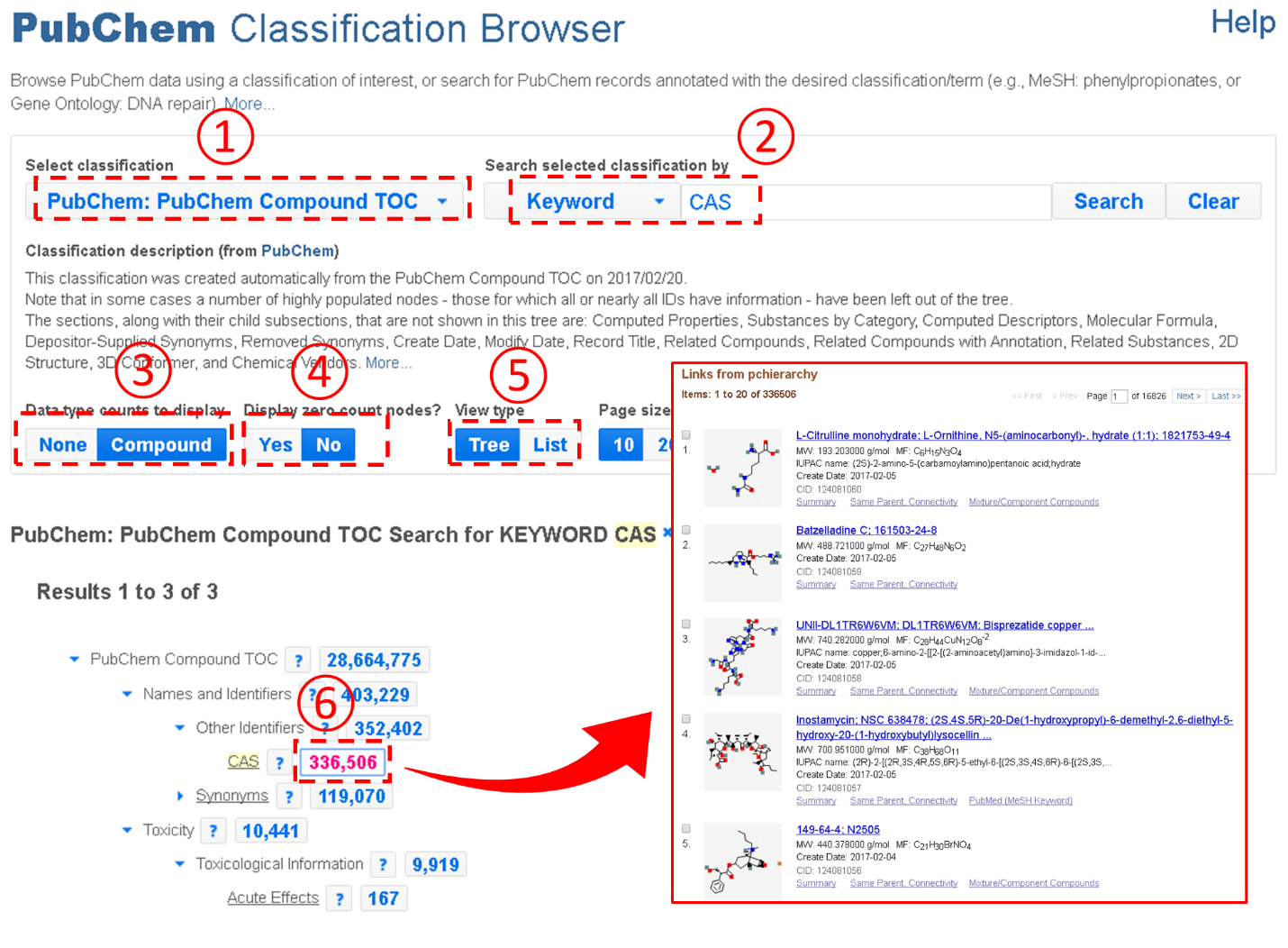
The Classification Browser also supports the PubChem BioAssay Classification Tree, providing an additional approach to browse, search, and access the BioAssay data. More detailed information on the Classification Browser is available at the URL:
http://pubchem.ncbi.nlm.nih.gov//classification/docs/classification_help.html
Identifier Exchange Service
The Identifier Exchange Service can be found at the following URL: http://pubchem.ncbi.nlm.nih.gov/idexchange
This service allows the user to convert one type of identifiers for a given set of chemical structures into a different type of identifiers for identical or similar chemical structures. Currently, it supports seven types of identifiers: CID, SID, InChI, InChIKey, SMILES, synonyms, Registry ID. When Registry ID is selected as an input or output identifier type, the DSN (Data Source Name) should also be provided.
The input identifier list may be provided using a string, a text file, or Entrez history. When a service request is submitted, it will be queued on PubChem servers. Once the actual task starts to run, the input identifiers will be converted into CIDs (called input CIDs) during the computation, and the CIDs (called output CIDs) that satisfy the condition specified by one of the following operation types will be retrieved:
- Same CID: Same CIDs as input CIDs.
- Same, Stereochemistry: CIDs that have same stereo centers as input CIDs.
- Same, Isotopes: CIDs that have the same isotopes as input CIDs.
- Same, Connectivity: CIDs that have the same connectivity as input CIDs.
- Same parent: CIDs that have the same parents as input CIDs.
- Same parent, Stereochemistry: CIDs that have the same stereo centers and parents as input CIDs.
- Same parent, Isotopes: CIDs that have the same isotopes and parents as input CIDs.
- Same parent, Connectivity: CIDs that have the same connectivity and parents as input CIDs.
- Similar 2D compounds: CIDs similar to the input CIDs in PubChem’s 2-D similarity.
- Similar 3D conformers: CIDs similar to the input CIDs in PubChem’s 3-D similarity.
These output CIDs are then converted into the identifier type specified by the user and written into a file or sent to Entrez history. In practice, the identifier exchange service may be used as a quick approach to search the PubChem Compound database using multiple queries, although this type of task may be performed programmatically (for example, using PUG-REST,1 which will be discussed in Module 7). A more detailed information is available at the URL:
http://pubchem.ncbi.nlm.nih.gov//idexchange/idexchange-help.html
The PubChem Data Sources page
As discussed in Section 3.4, the PubChem Data Sources page (https://pubchem.ncbi.nlm.nih.gov/sources/) helps users determine who provided what information. This page can be used to retrieve the data provided by a data depositor or to download the annotations collected from a data source. For example, the following figure illustrates how to download the boiling point data collected from DrugBank.2
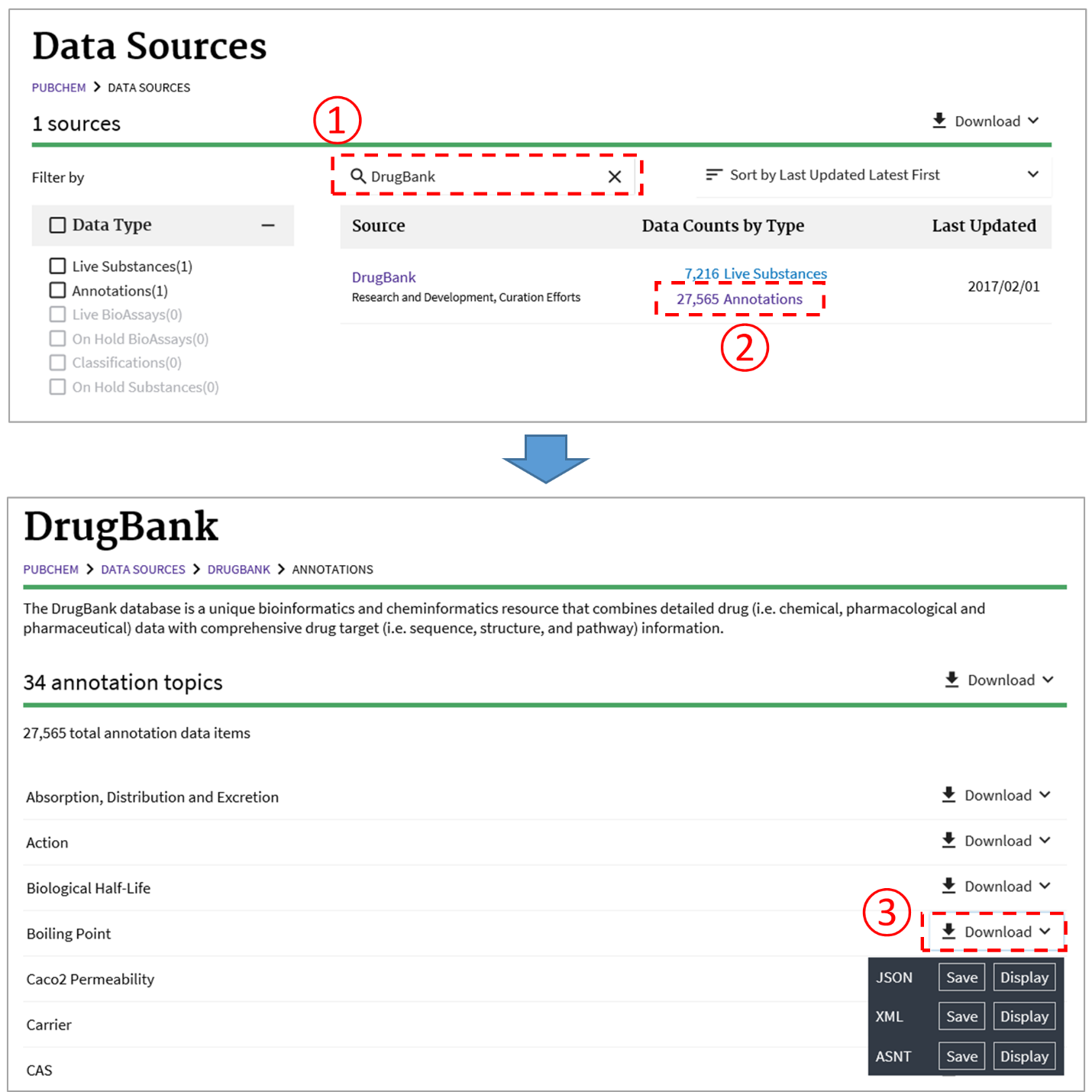
To obtain a particular kind of annotated information (e.g., boiling points) through the PubChem Data Sources page, one may need to know “in advance” which depositors provide that information. This can be done through a PUG-REST request1 (to be discussed in detail in Module 7). For example, the following PUG-REST request returns all data sources that provide the boiling point information for chemicals.
https://pubchem.ncbi.nlm.nih.gov/rest/pug/annotations/heading/boiling%20point/TXT
On the other hand, one may want to know what kind of information is provided by a given data source. This can also be done using a PUG-REST request:
https://pubchem.ncbi.nlm.nih.gov/rest/pug/annotations/sourcename/DrugBank/TXT
This example retrieves all types of annotations collected from DrugBank.

