4.4: Interpreting Electron Ionization Mass Spectra
- Page ID
- 401621
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Different ionization methods will produce different types of mass spectra. This page describes the interpretation of the mass spectra derived from electron ionization mass spectrometry.
In order for mass spectrometry to be helpful the ions that are studied must originate from a pure source of the sample of interest, for to consider ions that are unrelated to the compound could be worse than having no mass spectral data at all. It is essential to ensure that a sample contains no material that will produce unexpected ions that could interfere with interpretation. Thus, the combination of gas chromatography and mass spectrometry (GCMS) is often the method of choice. The gas chromatograph can separate the compound of interest from other potentially interfering compounds and present it to the mass spectrometer in purified form over a time interval sufficient to obtain the mass spectrum.
Per unit quantity of sample, mass spectrometry perhaps delivers more information concerning a molecule than does any other spectroscopic technique. Although mass spectrometry rarely solves a problem "single-handedly", it often provides the critical complimentary data that finally "crack" an identification problem. For a complete identification of a sample compound 0.1 to 1 micrograms of the sample may be required. If all that is needed is detection and verification of the presence of an already-identified compound the sample requirements are less severe (sometimes requiring only femtomole quantities).
Although mass spectrometry has been used successfully in the determination of many compounds (usually having a molecular weight of less than 1,000), one consistent obstacle is whether a compound of interest is amenable to vapor phase analysis. However, techniques such as field desorption (where a sample molecule is desorbed from the surface of a probe or electrode as an ion when it enters a high electric field), and chemical derivatization to increase thermal stability and volatility, extend the applicability of mass spectrometry to increasing numbers and types of compounds.
The formation of Molecular Ions (\(\ce{M^{+}}\))
When the vaporized organic sample passes into the ionization chamber of a mass spectrometer, it is bombarded by a stream of electrons. These electrons have a high enough energy to knock an electron off an organic molecule to form a positive ion. This ion is called the molecular ion - or sometimes the parent ion. The molecular ion is often given the symbol \(\ce{M^{+}}\) or \(\ce{M^{\cdot +} }\)- the dot in this second version represents the fact that somewhere in the ion there will be a single unpaired electron. That's one half of what was originally a pair of electrons - the other half is the electron which was removed in the ionization process.
The most important ion sought in the mass spectrum is the molecular ion (the ionized, intact molecule) because it is a direct indicator of the molecular weight of the compound. While the molecular weight is usually sufficient to confirm the presence of a given compound, electron ionization mass spectrometry can give even more information from the fragmentation of ions. Since some compounds can have molecular weights that are nearly identical, it is much better to compare the major peaks in the experimental spectrum to those in a reference spectrum of the authentic compound.
Fragmentation
During the process of ionization considerable excess energy may be transferred to the molecular ion, which depending on its stability (related to its structure) may decompose into various fragment ions. The resulting fragment ions define subsets of atoms that may be related to functional groups or structural components of the original molecule. The array of fragment ions represented by peaks in a mass spectrum is often called a fragmentation pattern. Careful study of the fragmentation pattern is especially important when the mass spectrum is to be used to elucidate molecular structure or to distinguish between two or more structural isomers.
While some fragmentation of the sample molecules is necessary for structure evaluation, many compounds are so effectively fragmented by EI that no appreciable population of molecular ions survives to give an indication of the molecular weight in the mass spectrum. Chemical ionization (CI), involving ionization of the sample molecule by collisions with ions of a reagent gas (for example, the ions produced by electron ionization of methane), is a more "gentle" ionization technique that often produces abundant protonated molecular ions. The resultant spectrum provides a good indication of molecule weight but may have insufficient fragmentation for functional group and structural evaluation.
The simplest case is that a molecular ion breaks into two parts - one of which is another positive ion, and the other is an uncharged free radical.
\[M^{\cdot +} \rightarrow X^+ + Y^{\cdot}\]
The uncharged free radical won't produce a line on the mass spectrum. Only charged particles will be accelerated, deflected and detected by the mass spectrometer. These uncharged particles will simply get lost in the machine - eventually, they get removed by the vacuum pump. The ion, X+, will travel through the mass spectrometer just like any other positive ion - and will produce a line on the stick diagram. All sorts of fragmentations of the original molecular ion are possible - and that means that you will get a whole host of lines in the mass spectrum. For example, the mass spectrum of pentane looks like this:
Each line represents a different fragment produced when the molecular ion breaks up.
In the stick diagram showing the mass spectrum of pentane, the line produced by the heaviest ion passing through the machine (at m/z = 72) is due to the molecular ion. The tallest line in the stick diagram (in this case at m/z = 43) is called the base peak. This is usually given an arbitrary height of 100, and the height of everything else is measured relative to this. The base peak is the tallest peak because it represents the commonest fragment ion to be formed - either because there are several ways in which it could be produced during fragmentation of the parent ion, or because it is a particularly stable ion.
Interpreting fragmentation patterns
The following will demonstrate the interpretation of fragmentation patterns using examples of simple orgaminc molucules.
Example \(\PageIndex{1}\): Mass Spectrum of Pentane
Let's have another look at the mass spectrum for pentane:
What causes the line at m/z = 57?
Solution
How many carbon atoms are there in this ion? There can't be 5 because 5 x 12 = 60. What about 4? 4 x 12 = 48. That leaves 9 to make up a total of 57. How about C4H9+ then?
C4H9+ would be [CH3CH2CH2CH2]+, and this would be produced by the following fragmentation:
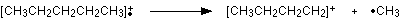
The methyl radical produced will simply get lost in the machine.
The line at m/z = 43 can be worked out similarly. If you play around with the numbers, you will find that this corresponds to a break producing a 3-carbon ion:
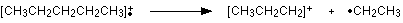
The line at m/z = 29 is typical of an ethyl ion, [CH3CH2]+:
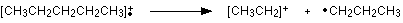
The other lines in the mass spectrum are more difficult to explain. For example, lines with m/z values 1 or 2 less than one of the easy lines are often due to loss of one or more hydrogen atoms during the fragmentation process.
Example \(\PageIndex{2}\): Pentan-3-one
This time the base peak (the tallest peak - and so the commonest fragment ion) is at m/z = 57. But this isn't produced by the same ion as the same m/z value peak in pentane.
If you remember, the m/z = 57 peak in pentane was produced by [CH3CH2CH2CH2]+. If you look at the structure of pentan-3-one, it's impossible to get that particular fragment from it.
Work along the molecule mentally chopping bits off until you come up with something that adds up to 57. With a small amount of patience, you'll eventually find [CH3CH2CO]+ - which is produced by this fragmentation:
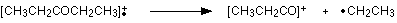
You would get exactly the same products whichever side of the CO group you split the molecular ion. The m/z = 29 peak is produced by the ethyl ion - which once again could be formed by splitting the molecular ion either side of the CO group.
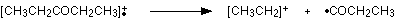
Peak heights and the stability of ions
The more stable an ion is, the more likely it is to form. The more of a particular sort of ion that's formed, the higher its peak height will be. We'll look at two common examples of this.
Examples involving carbocations (carbonium ions)
Summarizing the most important conclusion from the page on carbocations:
Order of stability of carbocations
primary < secondary < tertiary
Applying the logic of this to fragmentation patterns, it means that a split which produces a secondary carbocation is going to be more successful than one producing a primary one. A split producing a tertiary carbocation will be more successful still. Let's look at the mass spectrum of 2-methylbutane. 2-methylbutane is an isomer of pentane - isomers are molecules with the same molecular formula, but a different spatial arrangement of the atoms.
Look first at the very strong peak at m/z = 43. This is caused by a different ion than the corresponding peak in the pentane mass spectrum. This peak in 2-methylbutane is caused by:
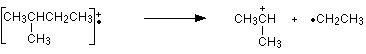
The ion formed is a secondary carbocation - it has two alkyl groups attached to the carbon with the positive charge. As such, it is relatively stable. The peak at m/z = 57 is much taller than the corresponding line in pentane. Again a secondary carbocation is formed - this time, by:
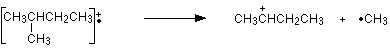
You would get the same ion, of course, if the left-hand CH3 group broke off instead of the bottom one as we've drawn it. In these two spectra, this is probably the most dramatic example of the extra stability of a secondary carbocation.
Examples involving acylium ions, [RCO]+
Ions with the positive charge on the carbon of a carbonyl group, C=O, are also relatively stable. This is fairly clearly seen in the mass spectra of ketones like pentan-3-one.
The base peak, at m/z=57, is due to the [CH3CH2CO]+ ion. We've already discussed the fragmentation that produces this.
Using mass spectra to distinguish between compounds
Suppose you had to suggest a way of distinguishing between pentan-2-one and pentan-3-one using their mass spectra.
| pentan-2-one | CH3COCH2CH2CH3 | |
| pentan-3-one | CH3CH2COCH2CH3 |
Each of these is likely to split to produce ions with a positive charge on the CO group. In the pentan-2-one case, there are two different ions like this:
- [CH3CO]+
- [COCH2CH2CH3]+
That would give you strong lines at m/z = 43 and 71.
With pentan-3-one, you would only get one ion of this kind:
- [CH3CH2CO]+
In that case, you would get a strong line at 57. You don't need to worry about the other lines in the spectra - the 43, 57 and 71 lines give you plenty of difference between the two. The 43 and 71 lines are missing from the pentan-3-one spectrum, and the 57 line is missing from the pentan-2-one one.
The two mass spectra look like this:
Computer matching of mass spectra
As you've seen, the mass spectrum of even very similar organic compounds will be quite different because of the different fragmentations that can occur. Provided you have a computer data base of mass spectra, any unknown spectrum can be computer analysed and simply matched against the data base.


