16.8: Observing the progress of hydrogenation and dehydrogenation reactions by UV spectroscopy
- Page ID
- 1016
To study any enzyme-catalyzed reaction, a researcher must have available some sort of test, or assay, in order to observe the reaction's progress and measure its rate. In many cases, an assay simply involves running the reaction for a specified length of time, then isolating and quantifying the product using a separation technique such as high pressure liquid chromatography (HPLC). This type of assay can be extremely time-consuming, however, so it is to the researcher's great advantage if a more convenient assay can be found.
Redox reactions in which a nicotinamide coenzyme participates as a hydride donor or acceptor are generally quite convenient to assay. In fact, the progress of these reactions can usually be observed in real time, meaning that the researcher doesn't need to stop the reaction in order to see how far it has progressed. NADPH and NADH have distinctive UV absorbance bands centered at 340 nm, with a strong molar absorbtivity of 6290 M-1 cm-1. The oxidized coenzymes NADP+ and NAD+ do not absorb at this wavelength (see the spectra in section 4.3D). Therefore, the course of a hydrogenation reaction in which NAD(P)H is converted to NAD(P)+ can be observed in real time if it is run in a quartz cuvette in a UV spectrometer. By observing the decrease in absorbance at 340 nm, the researcher can calculate how many moles of NAD(P)H has been oxidized at any given time point, and this number is equal to the number of moles of organic substrate that has been reduced. Likewise, a dehydrogenase reaction can be followed in real time by monitoring the increase in absorbance at 340 nm as NAD(P)+ is converted to NAD(P)H.
Example 16.8.1
- How many moles of substrate have been oxidized?
- What is the solution concentration of NADP+?
- The enzyme has a mass of 25 kilodaltons (25,000 g/mol).You added 5 mL of a 2 ng/mL solution of pure enzyme to start the reaction. How many reactions does each enzyme molecule catalyze, on average, per second? (This number is referred to by biochemists as the 'turnover number').
In many cases, it is also possible to monitor a reaction in real time using a UV assay, even if the reaction of interest is not a NAD(P)H or NAD(P)+ - dependent reduction or oxidation. The strategy used here is a called a coupled assay. Let's use the enzyme transaldolase (section 13.3D) as an example. To run the transaldolase reaction, a researcher would combine the two substrates erythrose-4-phosphate (E4P) and fructose-6-phosphate (F6P) in a suitable buffer, and then start the reaction by adding transaldolase enzyme. Unfortunately, the reaction taking place is invisible - how far it has progressed after 5 minutes, for example, could not be determined without stopping the reaction and somehow isolating and quantifying one of the two products. In the coupled UV assay strategy, the researcher also adds two other enzymes - triose phosphate isomerase (section 13.2A) and glycerol phosphate dehydrogenase (section 16.4D), along with NADH. If these enzymes are added in large excess, any glyceraldehyde phosphate (GAP) produced by the transaldolase reaction will be immediately isomerized to dihydroxyacetone phosphate (DHAP) by the isomerase enzyme and then reduced to (R)-glycerol phosphate ((R)-GP), with the concurrent oxidation of NADH to NAD+ .
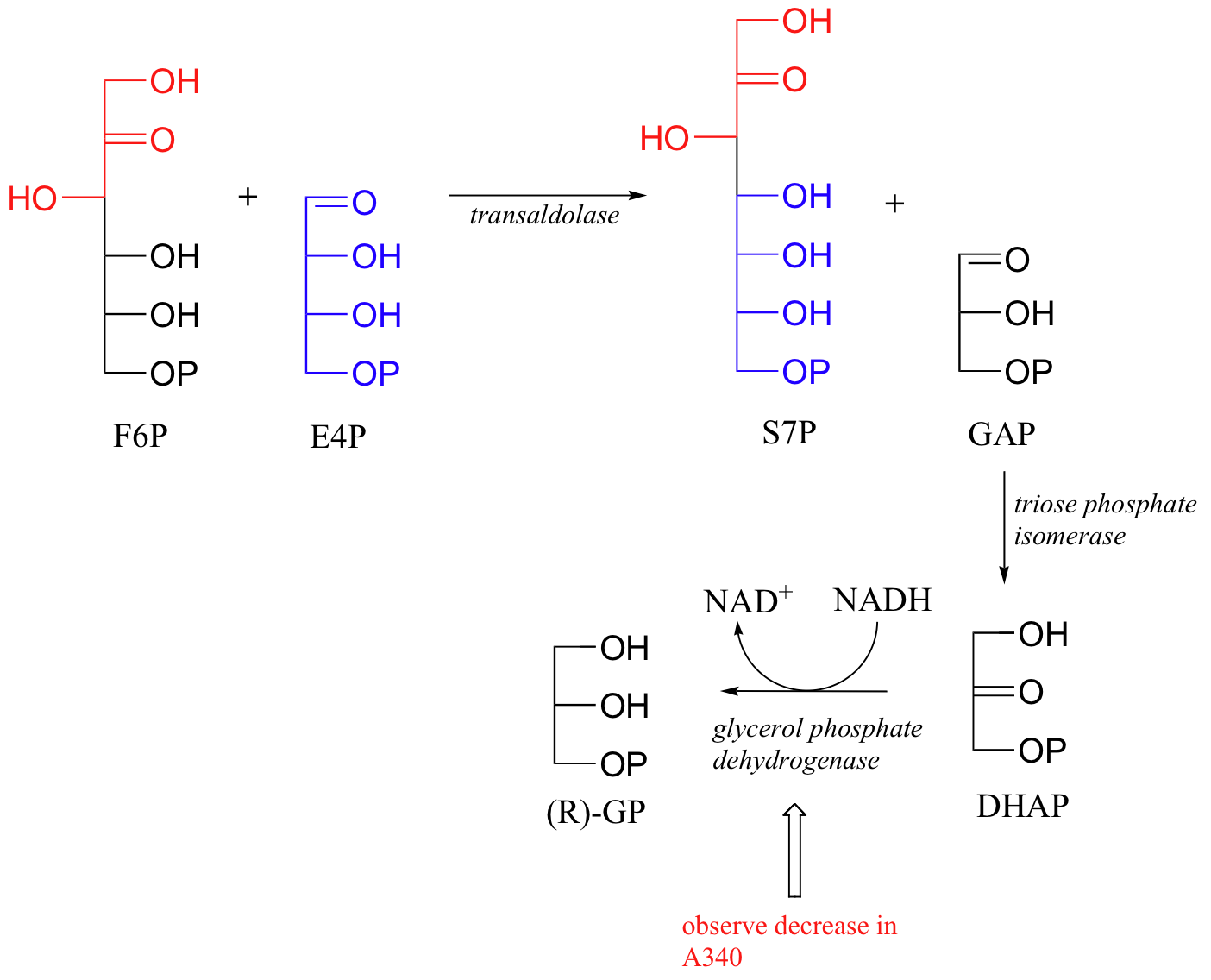
The production of GAP by transaldolase, therefore, can be indirectly (but accurately) monitored in real time by 'coupling' its production to NADH oxidation and the resulting observable decrease in A340. (The Enzymes, vol 7, 3rd ed. Editor: P.D. Boyer. Academic Press, New York. pp. 259-280).


