6.1: Serine proteases
- Page ID
- 165296
Serine proteases are just one type of endoproteases. However, they are extremely abundant in both prokaryotes and eukaryotes. Protease A, a chymotrypsin-like protease from Stremptomyces griseus, has a very different primary sequence than chymotrypsin, but its overall tertiary structure is quite similar to chymotrypsin, The positions of the catalytic triad amino acids in the primary sequences of the protein are very similar, indicating that the genes for the proteins diverged from a common precursor gene. In contrast, subtilisin, a serine protease from B. Subtilis, has both limited sequence and tertiary structure homology to chymotrypsin. However, when folded it also has a catalytic triad (Ser 221 - His 64 - Asp 32) similar to that of chymotrypsin (Ser 195 - His 57 - Asp 102).
Proteases have multiple functions, other than in digestion, including degrading old or misfolded proteins and activating precursor proteins (such as clotting proteases and proteases involved in programmed cell death). In general, the catalytic strategies used by four classes of enzymes: the serine proteases, carbonic anhydrases, restriction endonucleases, and nucleoside monophosphate (NMP) kinases. The first three classes of enzymes catalyze reactions that require the addition of water to a substrate. Proteases can also be integral membrane proteins, and carry out their activities in the hydrophobic environment of the membrane. For example, aberrant cleavage of the amyloid precursor protein by the membrane protease presenillin can lead to the development of Alzheimers.
Chymotrypsin
Consider the mechanism of catalysis of the enzyme known as chymotrypsin. Found in our digestive system, chymotrypsin’s catalytic action is cleaving peptide bonds in proteins and it uses the side chain of a serine in its mechanism of catalysis. Many other protein- cutting enzymes employ a very similar mechanism and they are known collectively as serine proteases. It acts fairly specifically, cutting not all peptide bonds, but only those that are adjacent to specific amino acids in the protein. One of the amino acids it cuts adjacent to is phenylalanine. The enzyme’s action occurs in two phases – a fast phase that occurs first and a slower phase that follows. The enzyme has a substrate binding site that includes a region of the enzyme known as the S1 pocket. Let us step through the mechanism by which chymotrypsin cuts adjacent to phenylalanine.
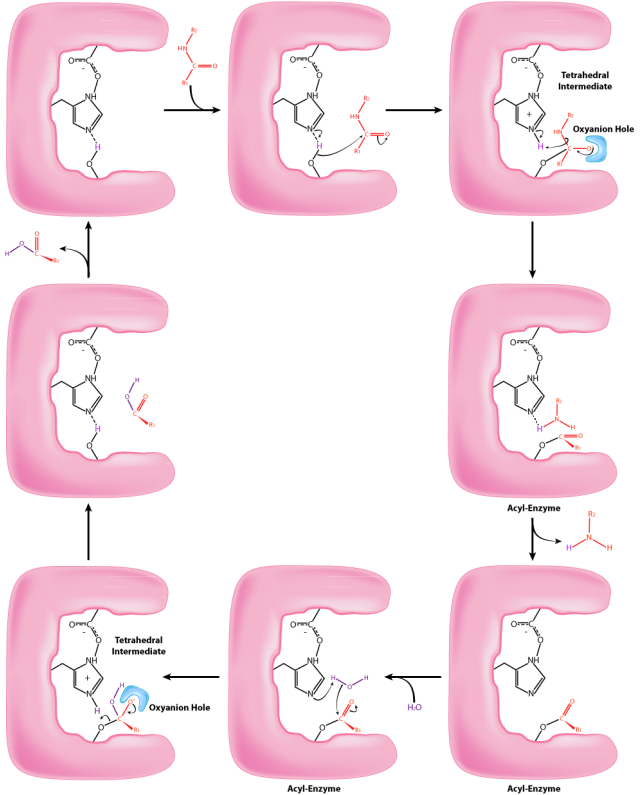 Figure 6.1.1: mechanism of catalysis of the enzyme chymotrypsin.
Figure 6.1.1: mechanism of catalysis of the enzyme chymotrypsin.
The process starts with the binding of the substrate in the S1 pocket. The S1 pocket in chymotrypsin has a hydrophobic hole in which the substrate is bound. Preferred substrates will include amino acid side chains that are hydrophobic, like phenylalanine. If an ionized side chain, like that of glutamic acid binds in the S1 pocket, it will quickly exit, much like water would avoid an oily interior. When the proper substrate binds, it stays and its presence induces an ever so slight shift in the shape of the enzyme. This subtle shape change on the binding of the proper substrate starts the steps of the catalysis and is the reason that the enzyme shows specificity for cutting at specific enzyme positions in the target protein. Only amino acids with the side chains that interact well with the S1 pocket start the catalytic wheels turning.
The slight changes in shape of the enzyme upon binding of the proper substrate cause changes in the positioning of three amino acids (aspartic acid, histidine, and serine) in the active site known as the catalytic triad, during the second step of the catalytic action. The shift of the negatively charged aspartic acid towards the electron rich histidine ring favors the abstraction of a proton by the histidine from the hydroxyl group on the side chain of serine, resulting in production of a very reactive alkoxide ion in the active site. Since the active site at this point also contains the polypeptide chain positioned with the phenylalanine side chain embedded in the S1 pocket, the alkoxide ion performs a nucleophilic attack on the peptide bond on the carboxyl side of phenylalanine sitting in the active site. This reaction, which is the third step of catalysis, breaks the bond and causes two things to happen. First, one end of the original polypeptide is freed and exits the active site. The second is that the end containing the phenylalanine is covalently linked to the oxygen of the serine side chain. At this point we have completed the first (fast) phase of the catalysis.
The second phase of the catalysis by chymotrypsin is slower. It requires that the covalent bond between phenylalanine and serine’s oxygen be broken so the peptide can be released and the enzyme can return to its original state. The process starts with entry of water into the active site. Water is attacked in a fashion similar to that of the serine side chain in the first phase, creating a reactive hydroxyl group that performs a nucleophilic attack on the phenylalanine-serine bond, releasing it and replacing the proton on serine. The second peptide is released in the process and the reaction is complete with the enzyme back in its original state.
Trypsin
Trypsin is a serine protease produced in the pancreas, is found in digestive system of vertebrates to digest food proteins. Trypsin cleaves peptide chains primarily at the carboxyl side of the lysine or arginine residues. Like chymotrypsin, trypsin uses a conserved catalytic triad, Ser, His, Asp, in the active site for catalysis, and the chemical mechanism is the same as chymotrypsin as shown in Figure 6.1.1.


