5.6: Effect of Conjugation
- Page ID
- 227574
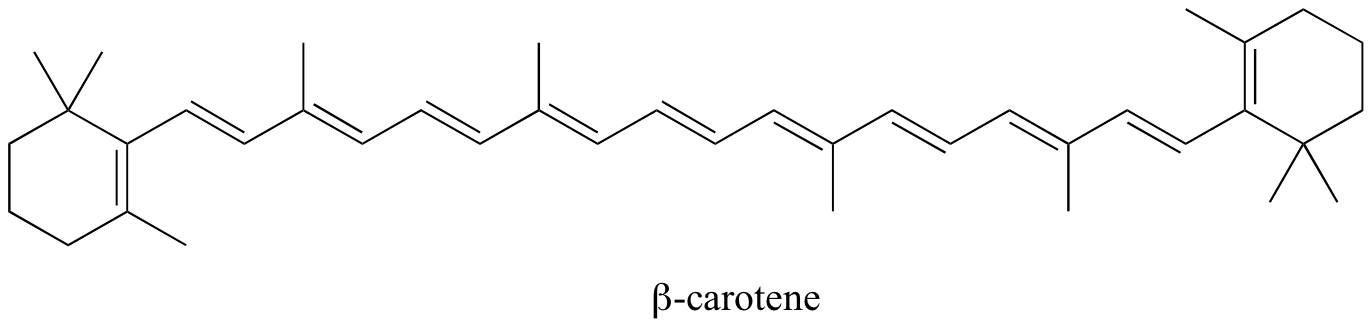
Conjugated systems or conjugated molecules contains an alternation of single and double bonds. For example, beta-carotene represents a conjugate system with 11 conjugated double bonds. Beta-carotene absorbs light with wavelengths in the blue region of the visible spectrum while allowing other visible wavelengths – mainly those in the red-yellow region - to be transmitted. This is why carrots are orange.
Here is the absorbance spectrum of the common food coloring Red #3:
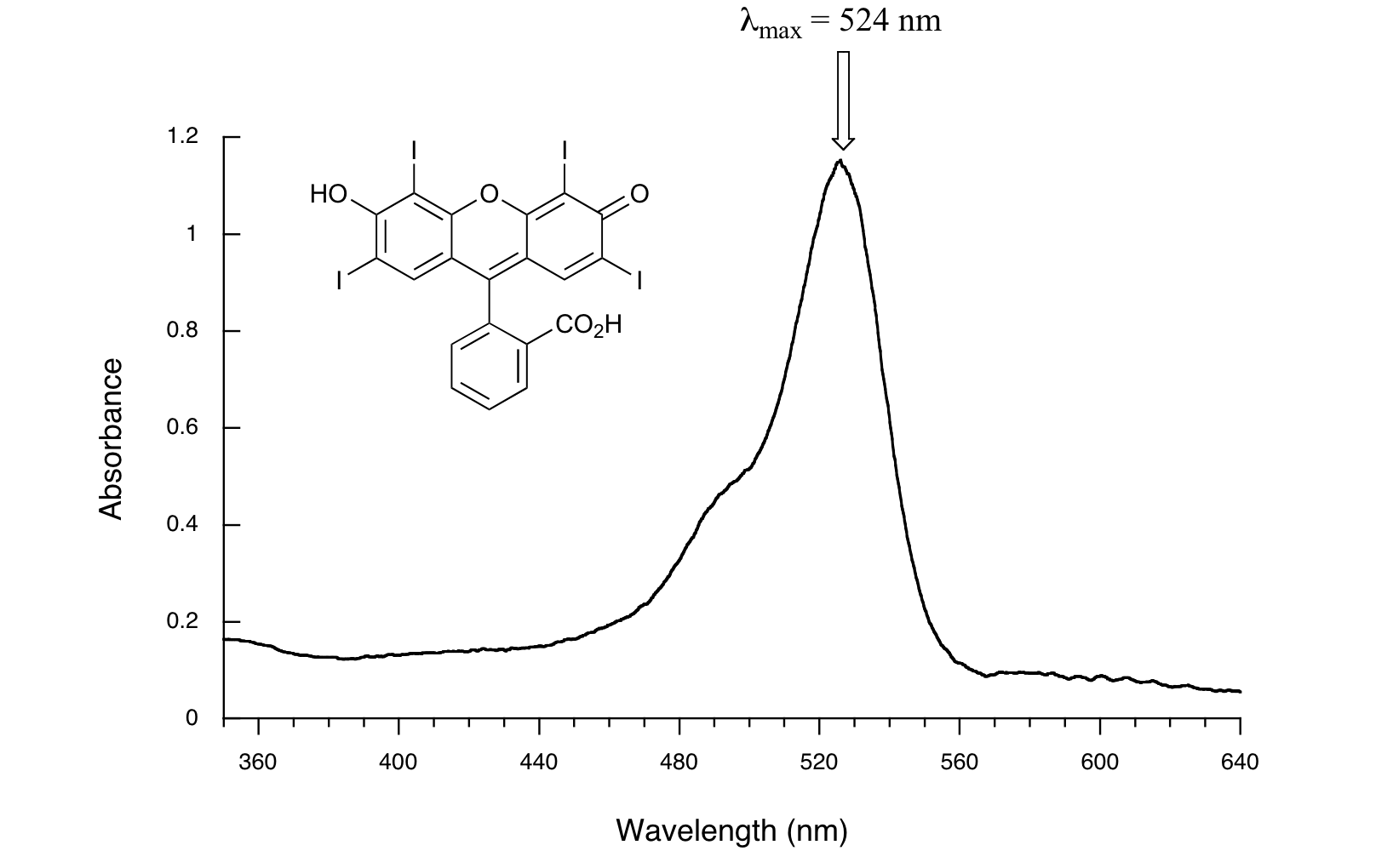
Here, we see that the extended system of conjugated pi bonds causes the molecule to absorb light in the visible range. Because the λmax of 524 nm falls within the green region of the spectrum, the compound appears red to our eyes.
Now, take a look at the spectrum of another food coloring, Blue #1:
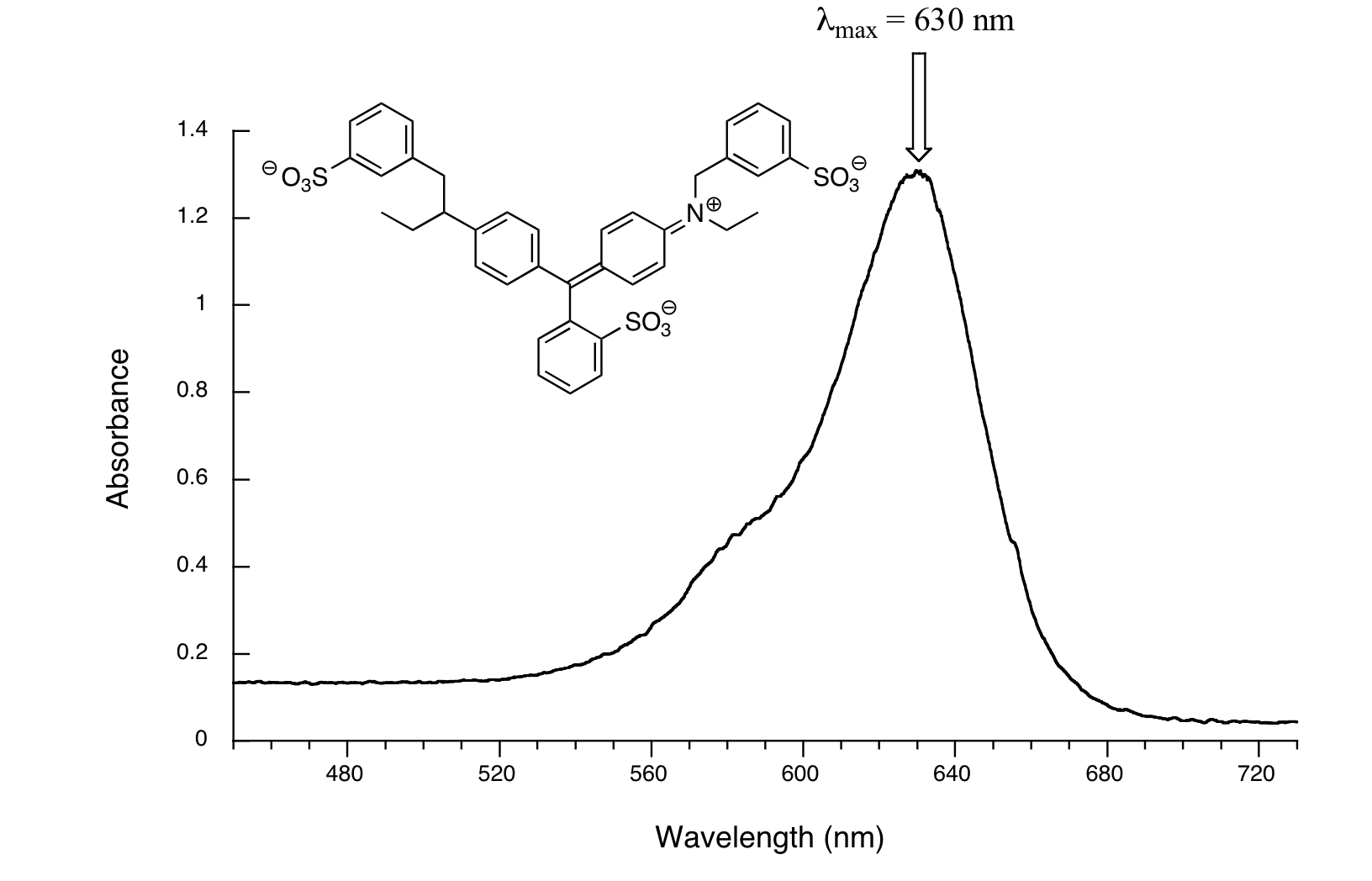
Here, maximum absorbance is at 630 nm, in the orange range of the visible spectrum, and the compound appears blue.
We can observe that the more conjugation present in a molecule, the higher the maximum absorbance ( λmax) values. We say that the λmax shifts towards longer wavelengths (lower energies, red shift). At the same time, the absorbance becomes more intense (higher values on the Y-axis).
Applications of UV spectroscopy in organic and biological chemistry
UV-vis spectroscopy has many different applications in organic and biological chemistry. One of the most basic of these applications is the use of the Beer - Lambert Law to determine the concentration of a chromophore. You most likely have performed a Beer – Lambert experiment in a previous chemistry lab. The law is simply an application of the observation that, within certain ranges, the absorbance of a chromophore at a given wavelength varies in a linear fashion with its concentration: the higher the concentration of the molecule, the greater its absorbance. If we divide the observed value of A at λmax by the concentration of the sample (c, in mol/L), we obtain the molar absorptivity, or extinction coefficient (ε), which is a characteristic value for a given compound.
ε = A/c
The absorbance will also depend, of course, on the path length - in other words, the distance that the beam of light travels though the sample. In most cases, sample holders are designed so that the path length is equal to 1 cm, so the units for molar absorptivity are mol * L-1cm-1. If we look up the value of e for our compound at λmax, and we measure absorbance at this wavelength, we can easily calculate the concentration of our sample. As an example, for NAD+ the literature value of ε at 260 nm is 18,000 mol * L-1cm-1. In our NAD+ spectrum we observed A260 = 1.0, so using equation 4.4 and solving for concentration we find that our sample is 5.6 x 10-5 M.
The literature value of ε for 1,3-pentadiene in hexane is 26,000 mol * L-1cm-1 at its maximum absorbance at 224 nm. You prepare a sample and take a UV spectrum, finding that A224 = 0.850. What is the concentration of your sample?
The bases of DNA and RNA are good chromophores:

Biochemists and molecular biologists often determine the concentration of a DNA sample by assuming an average value of ε = 0.020 ng-1×mL for double-stranded DNA at its λmax of 260 nm (notice that concentration in this application is expressed in mass/volume rather than molarity: ng/mL is often a convenient unit for DNA concentration when doing molecular biology).
50 mL of an aqueous sample of double stranded DNA is dissolved in 950 mL of water. This diluted solution has a maximal absorbance of 0.326 at 260 nm. What is the concentration of the original (more concentrated) DNA sample, expressed in mg/mL?
Because the extinction coefficient of double stranded DNA is slightly lower than that of single stranded DNA, we can use UV spectroscopy to monitor a process known as DNA melting. If a short stretch of double stranded DNA is gradually heated up, it will begin to ‘melt’, or break apart, as the temperature increases (recall that two strands of DNA are held together by a specific pattern of hydrogen bonds formed by ‘base-pairing’).
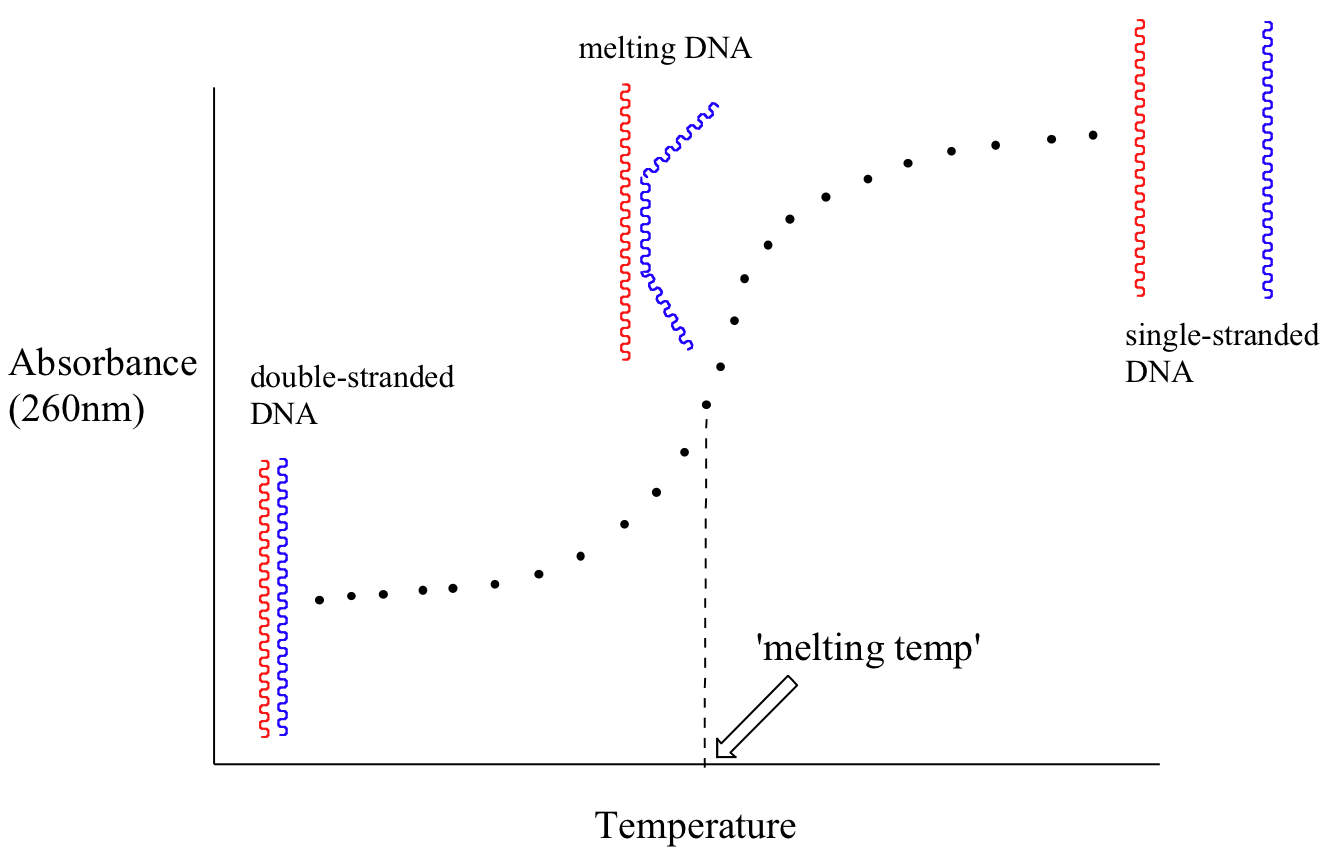
As melting proceeds, the absorbance value for the sample increases, eventually reaching a high plateau as all of the double-stranded DNA breaks apart, or ‘melts’. The mid-point of this process, called the ‘melting temperature’, provides a good indication of how tightly the two strands of DNA are able to bind to each other.
Proteins absorb light in the UV range due to the presence of the aromatic amino acids tryptophan, phenylalanine, and tyrosine, all of which are chromophores.
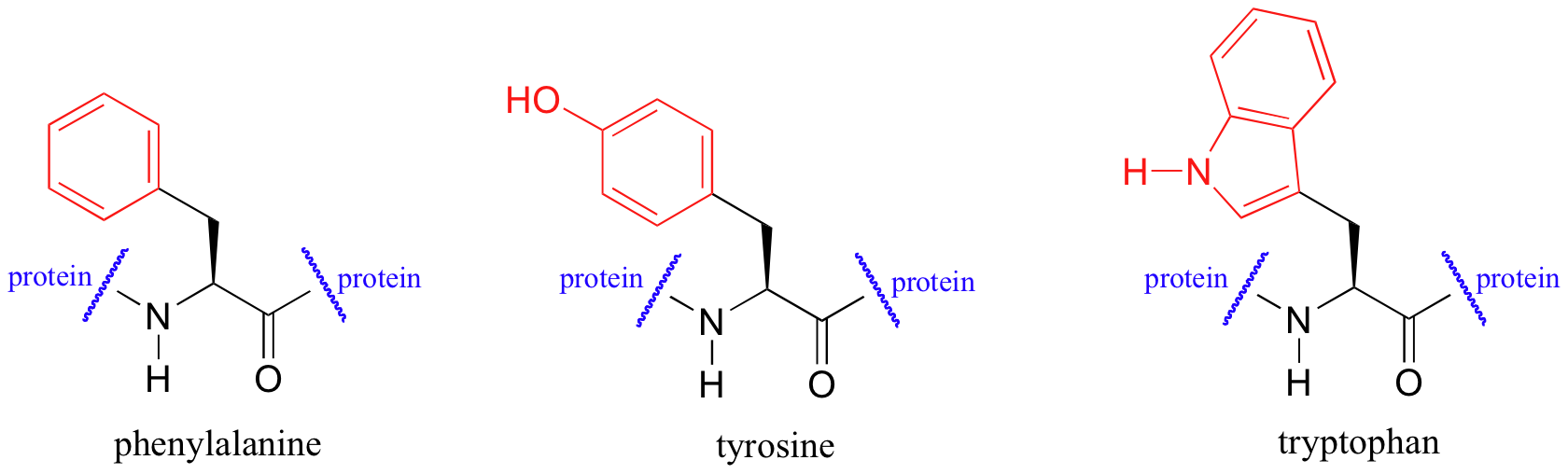
Biochemists frequently use UV spectroscopy to study conformational changes in proteins - how they change shape in response to different conditions. When a protein undergoes a conformational shift (partial unfolding, for example), the resulting change in the environment around an aromatic amino acid chromophore can cause its UV spectrum to be altered.

