9.3: Photosystem II- Harvesting the Photons
- Page ID
- 190060
It all starts with Photosystem II. Photosystem I comes later; the two complexes are named that way because Photosystem I was discovered before Photosystem II.
The picture below is a "cartoon rendering" of an X-ray crystal structure of photosystem II. So, researchers first took a solution of protein and somehow coaxed the protein molecules out of solution in the form of a crystal. In a crystal, the molecules are oriented in an organised array, so they are all lined up in a regular way. The researchers took the crystal and placed it in a beam of X-rays, then measured how the X-rays scattered when they hit the crystal. That scattering pattern was used to construct a three-dimensional map of the atomic positions in the protein. Because that is an awful lot of atoms, biochemists often present the map in different ways that are less complicated.
The cartoon rendering doesn't show where the individual atoms are, but it shows the locations of any alpha helices (pink) and beta sheets (yellow) as well as the chain formed by the rest of the protein that isn't organized in either of those ways (white). The parallel helices are typical of a membrane-bound protein; the top part of the structure can easily fit in among the parallel tails of the lipid bilayer that forms the thylakoid membrane.
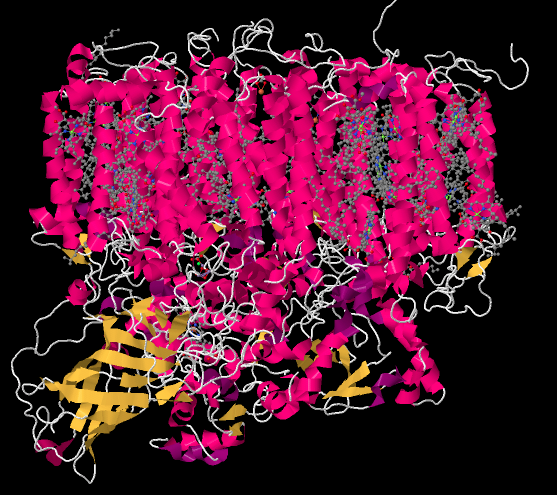
The structure below is shown from the same point of view, but with the protein removed. You can see that the protein is holding lots of smaller molecules in place; it is packed full of them. That's in sharp contrast to the complexes of the oxidative phosphorylation pathway, where there were at most a half-dozen metal clusters, which were directly attached to the protein, and a couple of ubiquinones, which were free.
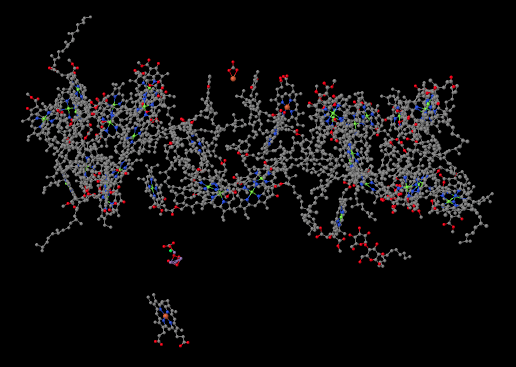

We're going to take just a moment and focus on something interesting in the lower left part of that picture. The whole thing is interesting, of course, but this is something extraordinarily important to us as humans. Halfway between the big, tangled mass of molecules at the top and the single molecule at the bottom (a heme, in case you were wondering) there is a small blob of atoms. Some of the atoms are red (oxygen, in this case) and some are purple (manganese); there is also a grey one (calcium) and even a green one (probably a stray chloride ion that photobombed the structure). This is the oxygen-evolving complex. It's responsible for producing all of the molecular oxygen, O2, on earth. Your life depends on it.
We'll get back to the oxygen-evolving complex in a later unit. Right now let's figure out what all those other molecules are doing in that scrum at the top of the picture, and why they are important. This is the light harvesting complex. Its job is to collect energy from sunlight and convert that energy into forms that the plant can use. Of course, somewhere along the line you learned in biology class that chlorophyll is what makes photosynthesis possible, and there are several chlorophyll molecules in the light harvesting complex. There are many different members of this family of molecules in nature, found in different organisms that can survive on sunlight: plants, algae, some plankton, a few bacteria. The chlorophylls commonly found in plants are chlorophyll a and chlorophyll b, shown below.
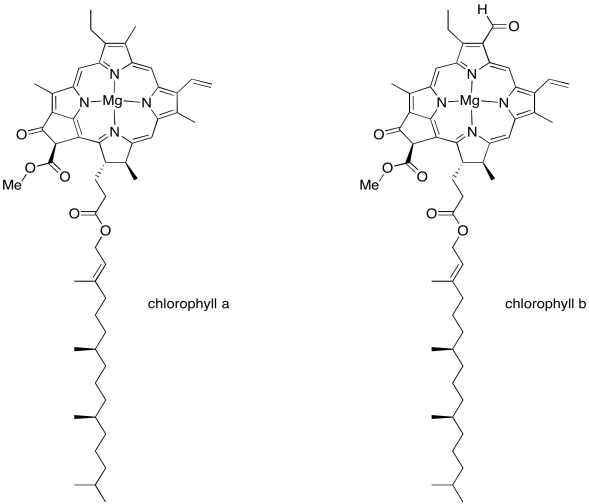
A chlorophyll is an example of a porphyrin, like the one that forms the heme unit in hemoglobin. It has a flat, polycyclic structure composed of four nitrogen-containing pyrole rings. Those nitrogen atoms can bind metal ions such as iron (in hemoglobin, myoglobin, and various cytochromes) or, in this case, magnesium.
Why are there two different chlorophylls, though? There are even more if you count the variations that are common in organisms other than plants. It takes a great deal of energy for a cell to manufacture different molecules; wouldn't it be more efficient to just make one kind of chlorophyll?
The advantage of chlorophyll diversity can be seen in UV-Vis spectra of the different compounds. Chlorophyll a has a couple of absorption maxima around 430 and 660 nm; there are also some smaller absorption features near 410 and 600 nm.
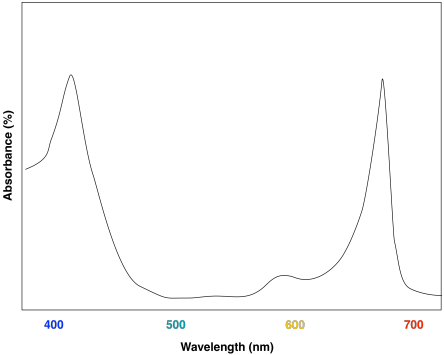
A look at the UV-Vis spectrum of chlorophyll b shows some strong similarities. There are still two major absorption maxima, near those for chlorophyll a, but they have shifted to about 450 and 640 nm.
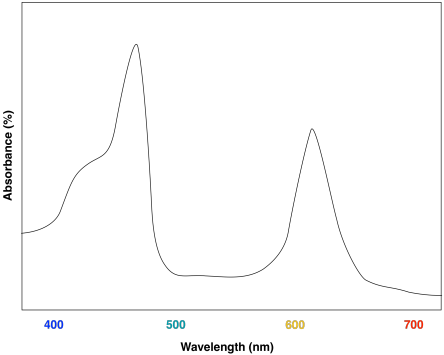
It probably isn't surprising that the two molecules have roughly similar UV-Vis spectra, since they have similar structures. Minor differences in the structures are responsible for subtle variations in energy levels of the donor and acceptor orbitals involved in the electronic transition, resulting in slightly different absorption maxima.
The combination of the two different chlorophylls would lead to a UV-Vis spectrum that looks a little like this one:
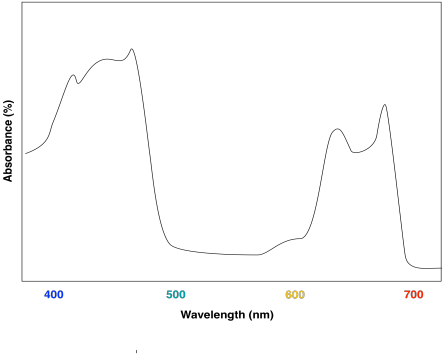
The effect is to harvest light from a slightly broader range of the visible spectrum. That makes for a more efficient process. Because sunlight contains wavelengths that range all across the visible spectrum (and beyond), the more of these wavelengths that can be captured, the more efficient photosynthesis can be.
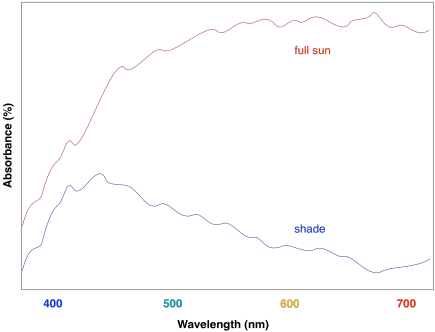
For this reason, the light harvesting complex contains more than one kind of chlorophyll, but it also contains other molecules. The most important class of these non-chlorophyll molecules includes the carotenoids. Compounds such as lycopene, β-carotene, and zeaxanthin are examples of carotenoids.
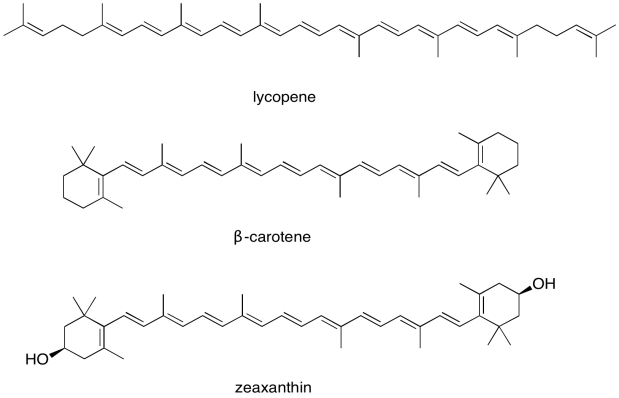
These compounds help to further broaden absorption by the light harvesting complex. We can see how they might start to fill in the gap in the previous spectrum that was based only on chlorophyll content.
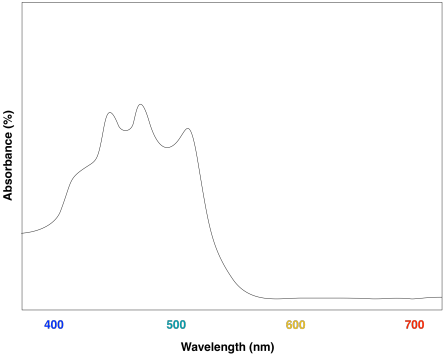
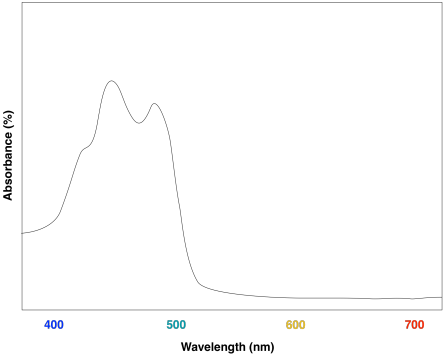
Overall, the result is a collection of molecules that can capture photons from across much of the visible spectrum, although not all. That's important, because solar radiation reaching the surface of the earth covers a broad range of wavelengths. Capturing as much light as possible increases the efficiency of the process. The fact that plants do reflect some visible light results in their green colour.
What next? All of that captured light energy needs to be converted into a different form; that's the whole point of photosynthesis. We aren't quite ready for that, though. The light harvesting complex just collects the light; the nearby reaction center is where the conversion process begins. You can think of the light harvesting complex as an antenna or as a funnel that sends energy to the reaction center, where the process of converting the energy into ATP (and, later, carbohydrates) will begin.
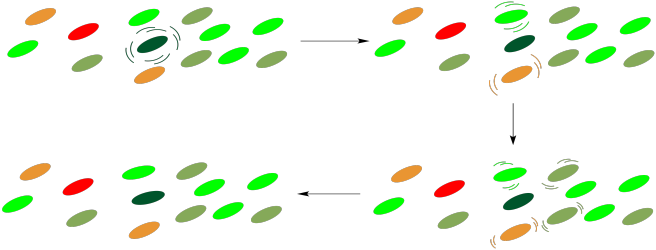
How does the energy get to the reaction center? It happens through a process of radiationless energy transfer. Once it absorbs the right amount of energy, the reaction centre will initiate an electron transport chain, much like the one that operated in oxidative phosphorylation. A photon doesn't need to strike the reaction centre itself, however, because it can strike any of the surrounding chromophores in the light harvesting complex. These molecules are all in close contact with each other, so it is relatively easy for molecules to collide and to transfer that entire amount of energy from one molecule to the next. During that transfer, the first molecule relaxes back to the ground state and the next molecule is excited into the excited state.

This process is very similar to photosensitization. In photosensitization, a molecule containing a chromophore, which can absorb a photon, transfers its energy to a second molecule, which has no chromophore and can't absorb light by itself. In the light harvesting complex, all of the molecules contain chromophores and could potentially absorb a photon. The point is that it doesn't matter which of them absorbs the photon; any of these absorptions will result in a cascade of radiationless transfers until the energy arrives at the right place. Once the energy reaches the reaction center, an electron transport chain will begin, and the energy released along this chain will be captured for the manufacture of ATP.
Conversely, it has been suggested that this array of molecules also affords protection to the cell. Under conditions in which the light intensity is too great, the reaction centre may become excited without an available outlet, as downstream participants in the pathway may not be ready to receive electrons. If that happens, a retro-mechanism may allow the reaction centre to relax once again, with energy dissipated to the surroundings through internal conversion.
In general, if a compound has one main absorption in the visible spectrum, we see the complementary colour. If a compound has two main absorptions in different parts of the spectrum, we generally see the colour of the "trough" or dip in the spectrum between these absorption maxima.
- What colour(s) correspond to the UV-Vis absorption maxima of chlorophyll a? What colour do we see when we look at chlorophyll a?
- What colour(s) correspond to the UV-Vis absorption maxima of lycopene? What colour do we see when we look at lycopene?
- What colour(s) correspond to the UV-Vis absorption maxima of β-carotene? What colour do we see when we look at β-carotene?
- Zeaxanthin is an example of a xanthophyll; xanthophylls get their name from the Greek roots "phyllon" (leaf) and "xanthos" (yellow). What colour probably corresponds to the UV-Vis absorption maximum of zeaxanthin?
- Answer a
-
a) blue and red; green
- Answer b
-
b) green; red
- Answer c
-
c) blue; orange
- Answer d
-
d) violet
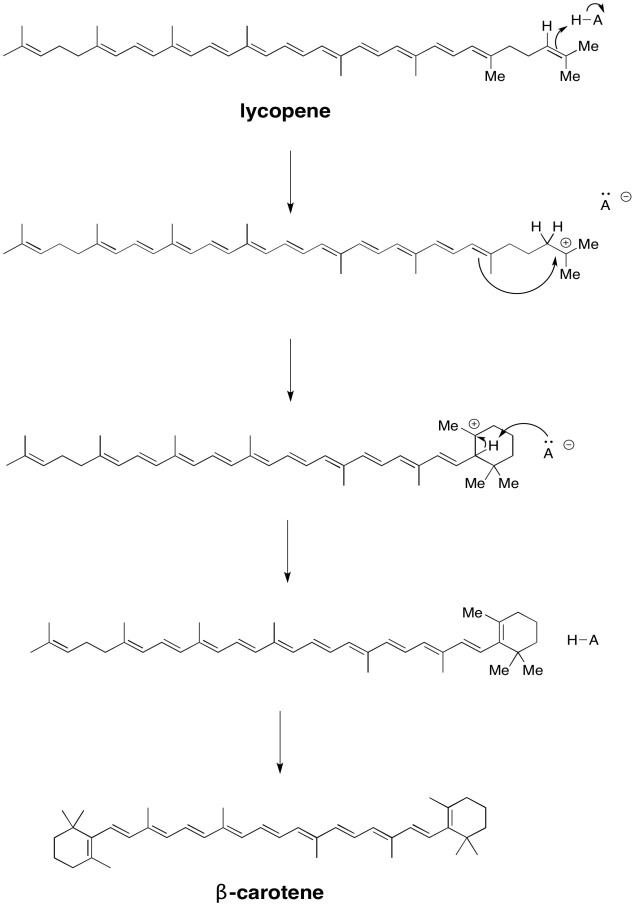
The carotenoids are biosynthetically related to each other. Show how, in principle, lycopene could be converted to β-carotene via simple acid/base catalysis.
- Answer
-
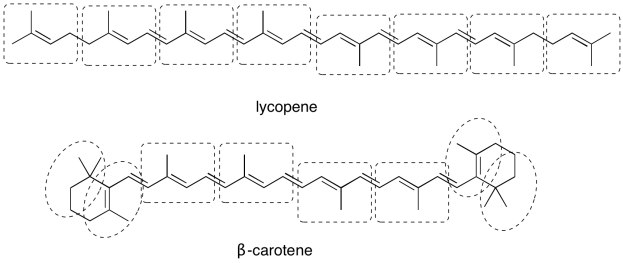
The carotenoids are examples of an enormous class of natural products called terpenes. Terpenes can be thought of as oligomers formed from 5-carbon isoprene units. Show the location of the five-carbon units in lycopene and β-carotene.
- Answer
-
Some marine algae contain phycoerythrin, a protein that is covalently linked to a unique chromophore. The chromophore structure and an artist's rendering of the UV spectrum are shown below.

- What colours are absorbed by the chromophore?
- What colour does the chromophore appear?
- What is the evolutionary advantage that drives the algae to make this chromophore?
1. X-ray crystal structures: Ferreira, K.N.; Iverson, T.M.; Maghlaoui, K.; Barber, J.; Iwata, S. Architecture of the photosynthetic oxygen-evolving center. Science 2004, 303, 1831-1838. Images obtained via RCSB Protein Data Bank (1S5L).


