1.2: Background
- Page ID
- 211997
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Experiment Background
General References:
SFH: Steinfeld, J. I., Francisco, J. S. and Hase, Chemical Kinetics and Dynamics, 2nd Ed., Prentice Hall, (1999).
- Definition of the Rate of a Chemical Reaction SFH pp. 1-3
- Order and Molecularity of a Reaction SFH pp. 3-6
- Reaction Mechanisms SFH pp. 17-18
- Enzyme-Catalyzed Reactions SFH pp. 159-163
- Measuring Mass and Volume MHS pp. 38-46
- Heating and Cooling Methods MHS pp. 49-58
- Rates of Chemical Reaction SWH pp. 879-884
- Enzyme-Catalyzed Reactions SWH pp. 885-892
Videos: Digital Techniques Manual
1. Volumetric Techniques (pipette, volumetric flask)
2. Balances
Introduction Chemical Kinetics
One of the main goals of chemical kinetics is to understand the steps by which a reaction takes place and how reaction conditions such as temperature and concentration affect the rate at which reactions occur. The series of elementary steps is the reaction mechanism. Understanding the mechanism allows us to find ways to manipulate the reaction. The rate of a chemical reaction depends on the temperature, pressure, pH and concentration of the reactants. From a study of the rate of a reaction, information can be obtained about the steps by which reactants are transformed into products. For a typical reaction:
\[ \rm aA + bB \rightarrow cC + dD\]
the rate of the reaction, v, can be written as:
\[ \rm rate = v = - \dfrac{1}{a} \dfrac{d[A]}{dt} = - \dfrac{1}{b} \dfrac{d[B]}{dt} = \dfrac{1}{c} \dfrac{d[C]}{dt} = \dfrac{1}{d} \dfrac{d[D]}{dt}\]
The rate law, or differential rate law, can also show how the rate of a reaction is a function of the concentrations of the reactants:
\[ v = k \space\space\space [A]^x \space\space\space [B]^y\]
where in this rate equation, k is the rate constant, and x and y represents the order of the reaction with respect to reactants A and B. Please note that x and y are not related directly to the stoichiometric coefficients a and b in the balanced equation but are independent coefficients that must be determined experimentally.
The Decomposition of Hydrogen Peroxide
Hydrogen peroxide is decomposed by the enzyme catalase to water and oxygen:
\[ \rm 2 H_2O_{2 \space\space (aq)} \xrightarrow{catalase} 2 H_2O_{(l)} + O_{2 \space\space (g)}\]
The rate of the decomposition of hydrogen peroxide, the rate at which oxygen is generated and the overall rate of the reaction are related by the stoichiometry of the reaction. The rate law for the above reaction is defined as:
\[ \rm - \dfrac{1}{2} \dfrac{\Delta H_2O_2}{\Delta t} = \dfrac{\Delta O_2}{\Delta t} = \text{rate of reaction}\]
For us, it is much easier to determine the amount of oxygen produced rather than the amount of hydrogen peroxide that remains in solution so for our purposes we will study the kinetics of this reaction by monitoring the rate that oxygen forms. We can do this in one of two ways, by measuring the volume of oxygen that is produced at a constant pressure or by measuring the pressure of oxygen produced under conditions of constant volume. We will use the latter method and follow the rate of the catalase decomposition of hydrogen peroxide by measuring the pressure of O2 produced under conditions of constant volume. One method of measuring the pressure of O2 produced is to attach a pressure sensor to the reaction flask and measure changes in pressure with time. From this data, we can obtain a graph of the pressure of oxygen generated versus time. The rate of the oxygen formation is just the slope of the curve on the plot ΔP/ Δt. The slope will be changing throughout the course of the reaction and is not constant because it depends on the temperature and the amount of hydrogen peroxide remaining in solution at any given time. In the case of O2 formed, the pressure measured can be readily converted back to concentration using the ideal gas equation4 .
\[ \rm PV=nRT\]
that is:
\[ \rm P = \dfrac{n}{V} RT = [O_2]RT\]
In terms of the Kinetics of the reaction and rate of oxygen gas production:
\[ \rm rate = \dfrac{d[O_2]}{dt} = \dfrac{1}{RT} \dfrac{d[P]}{dt}\]
Plotting P v. Time and from the increase in oxygen pressure with time we can relate that back to the concentration of O2 present (M/s).
Enzyme Kinetics5
An Enzyme or high molecular weight protein is synthesized in living cells and is made up mainly of long chains of amino acids connected by peptide bonds. Enzymes are biochemical catalysts that can accelerate reactions by lowering the activation energy, thereby, speeding up the reaction without being used up or changed. Initially, an enzyme combines temporarily with a substrate molecule to form an enzymesubstrate complex, which proceeds to break up and release the product(s) plus the unchanged enzyme. The released enzyme then locates another substrate molecule to continue its catalytic process. A reaction for a typical substrate and enzyme is illustrated below showing the intermediate enzyme-substrate complex, the products and the released enzyme.
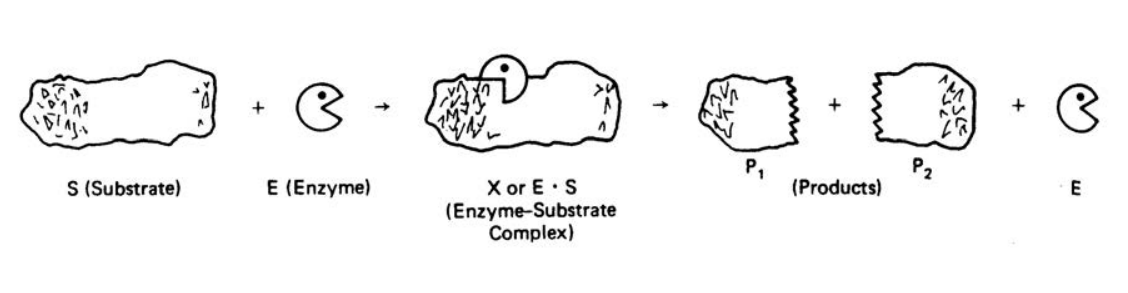
Figure 1. A Schematic Representation of an Enzyme-Catalyzed Mechanism6
Enzyme Kinetics involves the study of enzyme-catalyzed reactions, which may involve the study of the reaction rate and its dependence on substrate concentration, temperature, pH or a host of other variables. The enzyme-substrate complex is an intermediate in the reaction, and generally its presence in low concentrations makes it difficult to isolate.
In 1913 two researchers Michaelis and Menten proposed a theoretical way to study the mechanism of enzyme kinetics. The theory proposes that when an enzyme acts upon a substrate molecule, an enzyme-substrate complex forms and that this complex yields the product(s) in addition to the original unchanged enzyme. If the substrate is present in excess over the enzyme, the enzyme would get used up and form enzyme-substrate complexes. The rate in producing products becomes constant when a condition of steady state is reached.
Many one-substrate reactions follow this simple kinetic scheme, knows as the Michaelis-Menten mechanism.
Catalase
Catalase was chosen for this study because of its presence in the aerobic cells of most living organisms. It’s easy to qualitatively detect its presence in our blood if we get cut. When we apply hydrogen peroxide to the cut, we can see bubbles forming at the interface of the cut and our skin. This is the result of the enzyme catalase present in our blood breaking the hydrogen peroxide down into water and oxygen on the surface of our skin. We all produce a small amount of hydrogen peroxide as a by-product of our metabolism in a cellular membrane called peroxisome. During the normal course of oxidation/reduction reactions that take place in the human body, electrons are given up and some of those electrons promote the reduction of oxygen to hydrogen peroxide. Catalase present in our cells then catalyzes the breakdown of this hydrogen peroxide into oxygen and water in a disproportionation reaction. If the catalase were not present, the hydrogen peroxide produced in this normal metabolic process would build up and cause oxidative damage to our sensitive cellular components.
Catalase is a large quaternary protein, a tetramer, consisting of four large subunits. Each active subunit contains over 500 amino acids with a heme, a prosthetic group consisting of a protoporphyrin ring with a central iron at its center, similar to other quaternary type proteins such as hemoglobin.
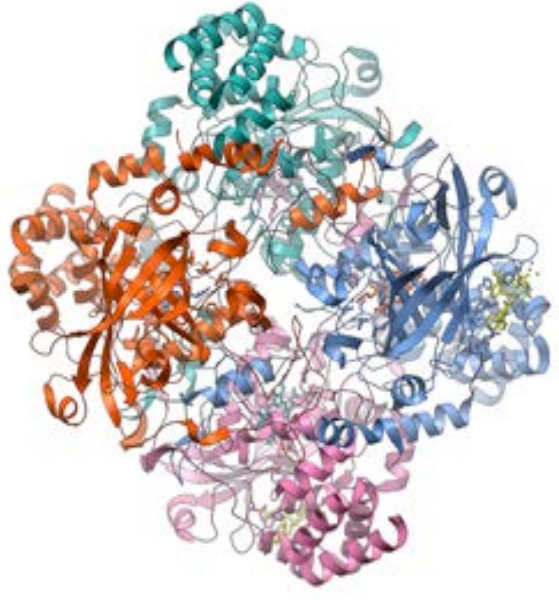
Figure 4. Three-Dimensional Structure of Catalase from PDB7
Footnotes:
4 Raymond Chang, Chemistry, 9th Edition, McGraw Hill, (2007) pp. 550-551.
5 Kinetics discussion adapted and modified from: Massachusetts Institute of Technology, Department of Chemistry, 5.310 Laboratory Chemistry, Kinetics Of An Enzyme – Catalyzed Reaction, Spring, 1993.
6 Steinfeld, J. I., Francisco, J. S. and Hase, Chemical Kinetics and Dynamics, 2nd Ed., Prentice Hall, (1999) pp. 159-163. © Pearson. Steinfeld, J. I., Francisco, J. S. and Hase, Chemical Kinetics and Dynamics, 2nd Ed., Prentice Hall, (1999) pp. 159-163. All rights reserved. This content is excluded from our Creative Commons license. For more information, see http://ocw.mit.edu/help/faq-fair-use/
7 Diagram taken from: Worldwide Protein Data Bank (PDB): www.wwpdb.org/docs.html
Putnam, C.D., A.S. Arvai, et al. (2000) Active and inhibited human catalase structures: ligand and NADPH binding and catalytic mechanism. Journal of Molecular Biology 296: 295-309 doi: 10.1006/jmbi.1999.3458

